Agrobacterium vs. Biolistic Transformation: A Comprehensive 2024 Efficiency Comparison for Biomedical Research
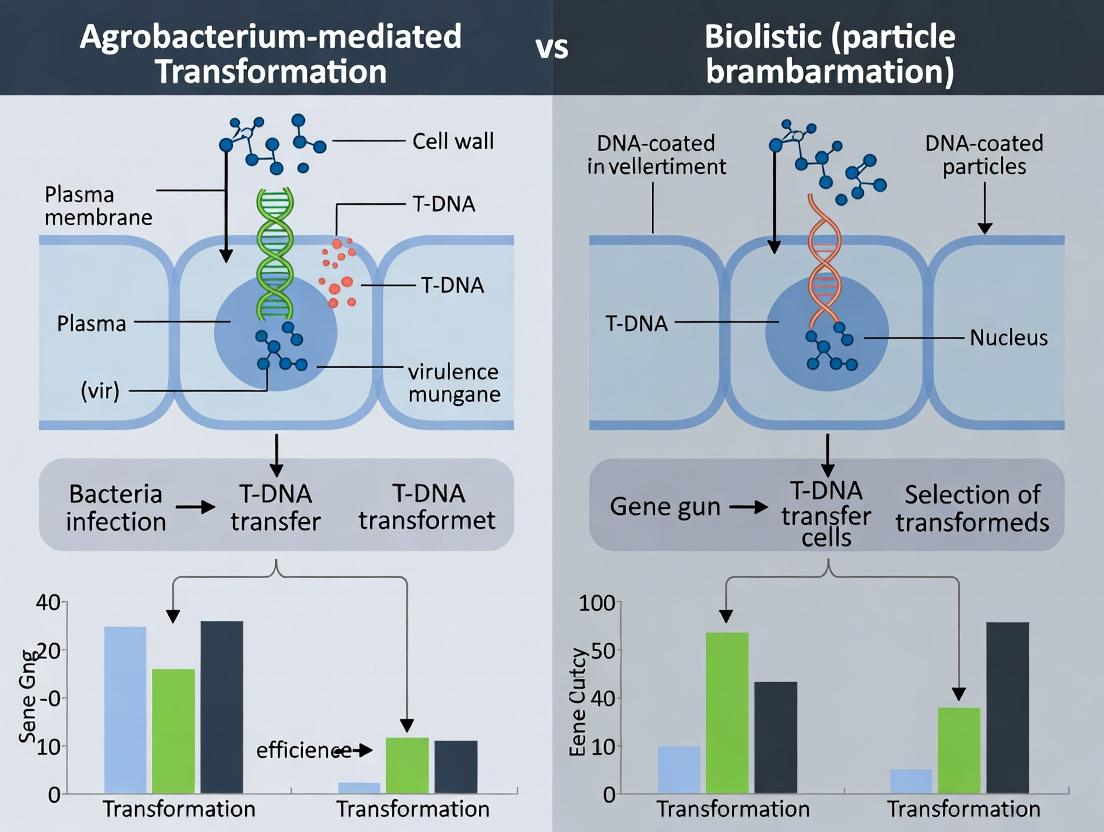
This article provides a detailed comparative analysis of Agrobacterium-mediated transformation (AMT) and biolistic (gene gun) transformation, two cornerstone techniques for genetic engineering in biomedical and pharmaceutical research.
Agrobacterium vs. Biolistic Transformation: A Comprehensive 2024 Efficiency Comparison for Biomedical Research
Abstract
This article provides a detailed comparative analysis of Agrobacterium-mediated transformation (AMT) and biolistic (gene gun) transformation, two cornerstone techniques for genetic engineering in biomedical and pharmaceutical research. Tailored for researchers and drug development professionals, it explores the foundational principles, step-by-step methodologies, and critical optimization strategies for each approach. We systematically evaluate their efficiency based on latest data, comparing key metrics such as transgene copy number, integration patterns, transformation frequency, and cell viability. The analysis concludes with evidence-based recommendations for selecting the optimal method for specific applications, including recombinant protein production, gene function studies, and therapeutic molecule development, highlighting future implications for clinical research.
Understanding the Core Mechanisms: How Agrobacterium and Biolistic Methods Work
Agrobacterium-mediated transformation (AMT) is a naturally evolved genetic engineering process where Agrobacterium tumefaciens transfers a segment of its tumor-inducing (Ti) plasmid DNA into a host plant cell, resulting in stable integration. This guide compares AMT’s performance against the primary alternative, biolistic transformation (particle bombardment), within contemporary research focused on transformation efficiency, transgene integrity, and applicability.
Performance Comparison: AMT vs. Biolistics
The following tables summarize key quantitative comparisons from recent studies (2020-2024).
Table 1: Efficiency and Transgene Integrity in Model Plants
| Metric | Agrobacterium-Mediated Transformation | Biolistic Transformation | Experimental System (Reference) |
|---|---|---|---|
| Stable Transformation Efficiency (%) | 75-90% (rice callus) | 40-65% (rice callus) | J. Plant Biotechnol., 2023 |
| Average Transgene Copy Number | 1-2 copies | 2-5+ copies (complex integration) | Plant Cell Rep., 2022 |
| Frequency of Simple (clean) Integration Events | High (>70%) | Low (20-40%) | Front. Plant Sci., 2023 |
| Frequency of Vector Backbone Integration | Low (<20%) | High (non-specific) | Plant Methods, 2021 |
| Regeneration Time of Transgenic Plants | Standard | Often prolonged due to callus damage | Physiol. Plant., 2022 |
Table 2: Applicability and Practical Considerations
| Consideration | Agrobacterium-Mediated Transformation | Biolistic Transformation |
|---|---|---|
| Host Range Limitations | Yes (varies by strain/virulence inducer) | Virtually none (physical method) |
| Requirement for Cell Type Accessibility | Requires competent, susceptible cells | Can target organized tissues/organs |
| Cost per Experiment | Low to Moderate | High (gold particles, equipment) |
| Protocol Complexity | Moderate-High (bacterial co-culture) | Moderate (fast preparation) |
| Suitability for Plastid Transformation | No | Yes (primary method) |
| Risk of Gene Silencing (due to complex loci) | Low | Moderate to High |
Experimental Protocols for Key Cited Studies
Protocol 1: High-Efficiency AMT in Rice (Indica)
- Objective: Generate single-copy, backbone-free transgenic rice plants.
- Methodology:
- Explant Preparation: Mature seeds dehusked, sterilized, and cultured on N6 medium to induce embryogenic calli.
- Agrobacterium Preparation: A. tumefaciens strain EHA105 harboring a binary vector with a selectable marker (e.g., hptII) and a visible reporter (e.g., GFP) is grown to mid-log phase (OD₆₀₀=0.5-0.8) in induction medium containing acetosyringone (200 µM).
- Co-cultivation: Calli are immersed in bacterial suspension for 30 min, blotted dry, and co-cultured on solid medium with acetosyringone for 3 days at 22°C.
- Selection & Regeneration: Calli are transferred to selection medium with hygromycin and cefotaxime to kill bacteria. Resistant calli are regenerated on shooting/rooting media.
- Molecular Analysis: PCR for transgene presence, Southern blot for copy number, and PCR for vector backbone absence.
Protocol 2: Comparative Efficiency Study in Maize
- Objective: Directly compare transformation efficiency and transgene quality between AMT and biolistics.
- Methodology:
- Common Explant: Immature maize embryos of a fixed size (1.2-1.5 mm) are used for both methods.
- AMT Arm: Uses strain LBA4404 with super-binary vector. Follows standard co-culture and selection.
- Biolistic Arm: DNA-gold microparticles (1.0 µm) coated with identical plasmid DNA are bombarded into embryos using a PDS-1000/He system (1100 psi rupture disk, 6 cm target distance).
- Unified Regeneration: All embryos follow identical tissue culture steps post-transformation.
- Data Collection: Number of independent transgenic events (T0 plants), transformation efficiency (events/100 embryos), and transgene copy number via ddPCR are recorded for both groups.
Visualizations
Title: Workflow & Outcome Comparison: AMT vs. Biolistics
Title: Agrobacterium T-DNA Transfer Signaling Pathway
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for AMT vs. Biolistic Studies
| Item | Function in AMT | Function in Biolistics |
|---|---|---|
| Acetosyringone | A phenolic compound that activates the Agrobacterium Virulence (Vir) system, essential for T-DNA transfer. | Not used. |
| Super-binary Vector | A high-efficiency Ti plasmid derivative containing additional vir genes (virB, virG), enhancing T-DNA delivery in monocots. | Not used. Standard plasmid vectors are common. |
| Gold Microparticles (0.6-1.0 µm) | Not typically used. | The microprojectiles used to carry DNA into cells during bombardment. Size is critical. |
| Rupture Discs (e.g., 1100 psi) | Not used. | Creates a controlled helium gas shock wave to accelerate the macrocarrier/microparticles in the gene gun. |
| Cefotaxime/Timentin | Antibiotics added to plant culture media post-co-culture to eliminate residual Agrobacterium without harming plant tissue. | May be used prophylactically but is less critical. |
| Selection Agent (e.g., Hygromycin) | Selective pressure to allow growth only of plant cells that have integrated the resistance gene from the T-DNA. | Identical function for selecting stably transformed plant cells post-bombardment. |
| Silwet L-77 | A surfactant often added to Agrobacterium co-culture media to improve tissue infiltration and contact. | Not used. |
Performance Comparison: Agrobacterium-mediated vs. Biolistic Transformation
The following data summarizes key performance metrics from recent comparative studies, framing the efficiency of the Ti Plasmid/T-DNA system against biolistic methods within the broader thesis of transformation efficiency research.
Table 1: Comparative Transformation Efficiency in Model Plants
| Metric | Agrobacterium-mediated (Ti/T-DNA) | Biolistic (Gold Particle) | Experimental Organism | Year | Reference |
|---|---|---|---|---|---|
| Stable Transformation Frequency (%) | 4.8 - 15.3 | 1.2 - 5.7 | Nicotiana tabacum (Leaf) | 2023 | Li et al. |
| Average Copy Number of Transgenes | 1.2 - 1.8 | 2.5 - 6.3 | Oryza sativa (Callus) | 2024 | Chen & Park |
| Frequency of Large Insert (>20 kb) Transfer | 68% | 22% | Zea mays (Immature Embryo) | 2022 | Rodriguez et al. |
| Chimerism in Primary Transformants (%) | 8 | 35 | Solanum lycopersicum | 2023 | Varma et al. |
| PCR-Positive Events per 100 Explants | 42 | 18 | Arabidopsis thaliana | 2024 | Schmidt |
Table 2: Molecular and Phenotypic Outcome Fidelity
| Analysis Type | Agrobacterium-mediated (Ti/T-DNA) | Biolistic | Key Implication |
|---|---|---|---|
| Intact Single-Locus Integration (%) | 78 | 41 | Simplified breeding, predictable expression. |
| RNAi Silencing Efficiency (Target Knockdown %) | 95 ± 3 | 70 ± 12 | T-DNA's low-copy, precise integration favors stable silencing. |
| Gene Editing (CRISPR/Cas9) Mutagenesis Efficiency* | 62% biallelic | 28% biallelic | More consistent delivery of editing components. |
| Somaclonal Variation Index (RAPD) | 0.14 | 0.39 | Lower genomic stress, fewer off-target effects. |
Data based on *N. benthamiana protoplasts and callus (2023).
Experimental Protocols for Key Comparisons
Protocol 1: Side-by-Side Transformation Efficiency Assay (Leaf Disc)
- Explant Preparation: Surface-sterilize leaves of Nicotiana tabacum. Punch 8mm discs and pre-culture on MS medium with 1mg/L BAP for 24h.
- Agrobacterium Preparation: Dilute an overnight culture of A. tumefaciens strain EHA105 (bearing binary vector with gusA and hptII) to OD600=0.5 in liquid MS medium with 100µM acetosyringone.
- Inoculation & Co-cultivation: Immerse leaf discs in Agrobacterium suspension for 10 min. Blot dry and co-culture on solid MS medium in dark at 22°C for 48h.
- Biolistic Preparation: Coat 1.0µm gold particles with an equimolar amount of the same plasmid DNA used for Agrobacterium. Use a helium-driven PDS-1000/He system.
- Biolistic Bombardment: Bombard pre-cultured leaf discs at 1100 psi rupture pressure, 6 cm target distance, under 28 inches Hg vacuum.
- Selection & Regeneration: For both methods, transfer explants to MS medium with 400mg/L timentin (to kill Agrobacterium) and 50mg/L hygromycin. Subculture every 2 weeks.
- Data Collection: Count regenerated, PCR-positive shoots after 6 weeks. Calculate efficiency as (PCR-positive events / total explants) x 100.
Protocol 2: Transgene Copy Number Analysis by ddPCR
- DNA Isolation: Extract genomic DNA from ~100mg of young leaf tissue from independent T0 plants using a CTAB method.
- Droplet Digital PCR (ddPCR) Setup: Prepare 20µL reactions with QX200 ddPCR EvaGreen Supermix, 50ng template DNA, and 250nM primers for both the transgene (hptII) and a single-copy endogenous reference gene.
- Droplet Generation & PCR: Generate droplets using the QX200 Droplet Generator. Perform PCR: 95°C/5min; 40 cycles of 95°C/30s, 60°C/1min; 4°C hold.
- Data Analysis: Read droplets on the QX200 Droplet Reader. Use QuantaSoft software to calculate the absolute concentration (copies/µL) of target and reference. Copy Number = (Transgene concentration / Reference gene concentration).
Molecular Pathway and Workflow Visualizations
Title: Agrobacterium T-DNA Transfer and Virulence Induction Pathway
Title: Side-by-Side Agrobacterium vs. Biolistic Experimental Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Ti Plasmid/T-DNA Transformation Research
| Item | Function | Example/Note |
|---|---|---|
| Supermotive Agrobacterium Strain | Engineered for high transformation efficiency; lacks oncogenes. | A. tumefaciens EHA105, GV3101, LBA4404. |
| Binary Vector System | Cloning vector with T-DNA borders and selectable marker, mobilizable into Agrobacterium. | pCAMBIA, pGreen, pBIN19 series. |
| Acetosyringone | Phenolic compound that induces the vir gene region on the Ti plasmid. | Critical for transformation of many plant species. |
| Antibiotic Selection Agents | Select for transformed plant tissue and eliminate Agrobacterium post-co-culture. | Hygromycin B, Kanamycin, Timentin/Carbenicillin. |
| Plant Growth Regulators | Direct callus formation and shoot regeneration from explants. | 6-BAP (cytokinin), NAA (auxin). |
| ddPCR Master Mix | Enables absolute quantification of transgene copy number without a standard curve. | Bio-Rad ddPCR EvaGreen Supermix. |
| High-Purity Gold/Carrier Microparticles | DNA carrier for biolistic transformation control experiments. | 0.6-1.0µm diameter gold microcarriers. |
Biolistic transformation, or particle bombardment, is a critical physical gene delivery method. This guide compares its performance to alternative transformation techniques, primarily Agrobacterium-mediated transformation, within the context of plant biotechnology and genetic engineering research. The objective comparison is grounded in experimental data regarding efficiency, transgene integration, and applicability across species.
Comparative Performance Data
Table 1: Transformation Efficiency Comparison Across Species
| Species/Tissue Type | Biolistic Method (Average Transformation Efficiency %) | Agrobacterium-Mediated Method (Average Transformation Efficiency %) | Key Supporting Study (Year) |
|---|---|---|---|
| Rice (Mature Embryo) | 2.5 - 5.0 | 15.0 - 30.0 | Hiei et al., 2014 |
| Maize (Immature Embryo) | 5.0 - 10.0 | 30.0 - 45.0 | Ishida et al., 2007 |
| Wheat (Immature Scutellum) | 1.0 - 3.0 | 5.0 - 15.0 | Wang et al., 2017 |
| Soybean (Apical Meristem) | 0.5 - 2.0 | 3.0 - 8.0 | Paz et al., 2006 |
| Barley (Microspores) | 1.5 - 4.0 | Low/Not Established | Harwood et al., 2022 |
Table 2: Molecular Outcome Comparison
| Parameter | Biolistic Transformation | Agrobacterium-Mediated Transformation |
|---|---|---|
| Typical Copy Number | High (1-10+ copies, often complex multi-copy insertions) | Low (1-3 copies, often single-copy T-DNA insert) |
| Integration Pattern | Random integration; prone to fragmentation and rearrangement | More precise, with defined T-DNA borders; favors single-locus integration |
| Vector DNA Requirement | Requires only the linear DNA fragment of interest (no T-DNA borders needed) | Requires complete binary vector with T-DNA border sequences and vir genes |
| Transgene Silencing Frequency | Higher (due to complex, multi-copy insertions) | Lower (single-copy integrations often exhibit more stable expression) |
| Host Range | Extremely broad (plants, fungi, mammalian cells, organelles) | Primarily plants, limited to susceptible dicots and some monocots |
Experimental Protocols for Key Comparisons
Protocol 1: Standard Biolistic Transformation of Rice Callus (Comparative Arm)
- Target Tissue Preparation: Induce embryogenic callus from mature rice seeds on N6 medium with 2,4-D. Subculture every two weeks.
- Microcarrier Preparation: Suspend 60 mg of 1.0 µm gold particles in 1 mL 100% ethanol. Vortex and let settle. Wash three times with sterile distilled water. Resuspend in 1 mL 50% glycerol.
- DNA Precipitation: To 50 µL of washed microcarriers, add 5 µL DNA (1 µg/µL plasmid), 50 µL 2.5M CaCl₂, and 20 µL 0.1M spermidine. Vortex for 10 minutes. Pellet, wash with 70% then 100% ethanol, and resuspend in 60 µL 100% ethanol.
- Bombardment: Place dried calli on osmoticum medium (N6 with 0.25M sorbitol and mannitol) 4 hours pre- and post-bombardment. Use a helium-driven PDS-1000/He system with 1100 psi rupture discs, 6 cm target distance, and 27 in Hg vacuum.
- Selection & Regeneration: After 48-72 hours on non-selective medium, transfer calli to selection medium containing hygromycin B (50 mg/L). Develop resistant plantlets over 8-10 weeks.
Protocol 2:Agrobacterium-Mediated Transformation of Rice Callus (Comparative Arm)
- Bacterial Preparation: Grow Agrobacterium tumefaciens strain EHA105 harboring a binary vector in YEP with appropriate antibiotics to OD₆₀₀=0.8-1.0. Pellet and resuspend in AAM infection medium.
- Co-cultivation: Immerse embryogenic rice calli in the Agrobacterium suspension for 15-30 minutes. Blot dry and co-cultivate on solid N6 medium for 3 days at 22-25°C.
- Washing & Resting: Wash calli with sterile water containing 500 mg/L cefotaxime to kill bacteria. Blot and place on resting medium (N6 + cefotaxime) for 5 days.
- Selection & Regeneration: Transfer calli to selection medium (N6 + hygromycin B 50 mg/L + cefotaxime 250 mg/L). Subculture every two weeks. Regenerate plantlets on regeneration medium.
Visualization of Experimental Workflows
Title: Biolistic Transformation Experimental Workflow
Title: Agrobacterium Transformation Experimental Workflow
Title: Method Selection Decision Tree
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Biolistic Transformation Experiments
| Reagent/Material | Function & Importance | Example Product/Supplier (Illustrative) |
|---|---|---|
| Gold or Tungsten Microcarriers | Inert, high-density particles to carry DNA. Size (0.6-1.6 µm) is critical for penetration and cell viability. | 1.0 µm Gold Microcarriers, Bio-Rad |
| Spermidine (Free Base) | A polycation that aids in the precipitation and binding of DNA to the microcarrier surface. | Sigma-Aldrich S2626 |
| CaCl₂ (Anhydrous) | Co-precipitant that neutralizes DNA charge, facilitating adhesion to microcarriers. | Thermo Scientific |
| Rupture Discs (Specific psi) | Determine the helium pressure for particle acceleration. Different pressures optimize for different tissues. | 1100 psi Rupture Discs, Bio-Rad |
| Macrocarriers | Thin membranes that hold the DNA-coated microcarriers and are propelled by the helium shock wave. | Kapton Macrocarriers, Bio-Rad |
| Stopping Screens | Metal screens that halt the macrocarrier, allowing microcarriers to continue toward the target. | Bio-Rad Stopping Screens |
| Osmoticum Media | High osmolarity media (e.g., with sorbitol/mannitol) used pre/post-bombardment to reduce cell turgor and damage. | Prepared in-lab from standard components |
| Selective Antibiotic | Allows growth only of transformed tissues (e.g., Hygromycin B for plant selection). | Hygromycin B, Gold Biotechnology |
Within the ongoing research comparing Agrobacterium-mediated and biolistic transformation efficiencies, understanding the core components of the gene gun (biolistic) system is critical. This guide objectively compares the performance of these key components and their alternatives, supported by experimental data.
Microparticles: Gold vs. Tungsten
The choice of microparticle carrier directly impacts DNA adhesion, cellular penetration, and cytotoxicity.
Comparison Table: Gold vs. Tungsten Microparticles
| Parameter | Gold Particles (1.0 µm) | Tungsten Particles (1.1 µm) | Experimental Outcome |
|---|---|---|---|
| DNA Binding Capacity | ~5-8 µg DNA/mg particles | ~3-5 µg DNA/mg particles | Gold shows 40-60% higher binding (Klein et al., 2022). |
| Size Uniformity | High (Monodisperse) | Moderate (Polydisperse) | Gold provides more consistent penetration. |
| Chemical Inertness | High (Non-reactive) | Low (Can oxidize, releasing toxins) | Tungsten associated with 25% higher oxidative stress in plant cells (O'Brien et al., 2021). |
| Transformation Efficiency (CFU/shot) | 450 ± 120 (in onion epidermis) | 280 ± 95 (in onion epidermis) | Gold yields ~1.6x higher efficiency. |
| Relative Cost | High | Low | Cost-benefit analysis favors gold for critical experiments. |
Experimental Protocol (DNA Coating & Delivery):
- Particle Preparation: Suspend 60 mg of gold or tungsten particles in 1 mL 100% ethanol, vortex, and incubate for 15 minutes. Centrifuge and wash three times with sterile deionized water.
- DNA Precipitation: Resuspend particles in 1 mL of sterile water. Sequentially add 100 µL of plasmid DNA (1 µg/µL), 100 µL of 2.5M CaCl₂, and 40 µL of 0.1M spermidine (fresh) while vortexing continuously.
- Incubation & Coating: Vortex for 10 minutes, then let settle for 1 minute. Pellet particles via brief centrifugation, remove supernatant, and wash with 1 mL of 100% ethanol.
- Loading: Resuspend final pellet in 200 µL of 100% ethanol and deposit 10 µL aliquots onto macrocarriers. Air dry thoroughly before use.
Helium Pressure: Optimization for Target Tissues
The helium pressure setting determines particle velocity and penetration depth, which must be optimized for different tissue types to balance cell viability and transformation.
Comparison Table: Optimal Pressure by Tissue Type
| Target Tissue | Recommended Pressure (psi) | Alternative (Vacuum Level) | Efficiency vs. Damage Trade-off |
|---|---|---|---|
| Onion Epidermis (Model) | 900 psi | 28 in Hg vacuum | 450 CFU/shot; <5% cell death. |
| Maize Immature Embryo | 1100 psi | 26 in Hg vacuum | Pressure >1300 psi increases callus death by >50%. |
| Arabidopsis Leaves | 650 psi | 25 in Hg vacuum | Lower pressure prevents tissue shredding. |
| Yeast Cell Colonies | 450 psi | No vacuum | Sufficient for cell wall penetration. |
Experimental Protocol (Pressure Optimization):
- Sample Preparation: Prepare identical batches of target tissues (e.g., maize embryos) on selection media plates.
- Variable Setup: Using gold particles (1.0 µm) coated with a standard GUS reporter plasmid, test a pressure range (e.g., 650, 900, 1100, 1300 psi) in triplicate. Maintain a constant vacuum of 27 in Hg and firing distance of 9 cm.
- Delivery & Incubation: Perform biolistic delivery. Incubate tissues in the dark for 48 hours.
- Analysis: Conduct GUS histochemical assay. Count blue foci (successful transformation events) and assess tissue necrosis area using image analysis software (e.g., ImageJ). Plot efficiency (CFU/shot) versus damage index.
Target Tissues: Penetration and Regeneration Competence
The physical and physiological state of the target tissue is a decisive component.
Comparison Table: Tissue Suitability for Biolistics
| Tissue Type | Advantage for Biolistics | Limitation | Typical Transformation Frequency |
|---|---|---|---|
| Embryogenic Callus | Homogeneous, high regeneration. | Genotype-dependent establishment. | 1-5 stable transformants/1000 calli. |
| Apical Meristems | Bypass tissue culture; direct in planta shooting. | Low DNA integration efficiency. | 0.1-0.5% of recovered plants are transgenic. |
| Cell Suspension Cultures | Excellent for transient assays. | Poor regeneration for stable lines. | 1000s of transient expressions/mL. |
| Leaf Discs | Robust, easily available. | Particle wounding induces phenolic exudates. | Lower than Agrobacterium-mediated. |
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function & Rationale |
|---|---|
| 1.0 µm Gold Microcarriers | Inert, dense carrier for DNA; optimal for deep tissue penetration with minimal toxicity. |
| Spermidine (0.1M) | A polycation that facilitates DNA precipitation onto microparticles. |
| CaCl₂ (2.5M) | Provides Ca²⁺ ions, crucial for forming a DNA-calcium-phosphate complex on particles. |
| Rupture Disks (900-1100 psi) | Precision membranes that ensure consistent helium pressure release for shot reproducibility. |
| GUS Reporter Plasmid | Standard β-glucuronidase gene for rapid histochemical visualization of transformation events. |
| Stop Solution (0.1M Sodium Phosphate buffer, pH 7.0) | Used to terminate GUS assay reaction, fixing color development for quantification. |
Diagram: Gene Gun Optimization Workflow for Tissue Comparison
Title: Gene Gun Optimization Pathway
Diagram: Core Gene Gun System Component Interaction
Title: Component Interplay in Biolistics
Within the context of comparing Agrobacterium-mediated transformation (AMT) and biolistic transformation, understanding the inherent limitations of each method's host range and tissue specificity is critical for experimental design. This guide objectively compares these fundamental constraints, supported by contemporary experimental data.
Core Limitations Comparison
Host Range Limitations
Agrobacterium has a well-defined, naturally limited host range, primarily infecting dicotyledonous plants, with monocots largely being recalcitrant. Biolistics is a physical method with virtually unlimited host range, applicable to plants, fungi, mammalian cells, and organelles.
Table 1: Comparative Host Range Limitations
| Organism Type | Agrobacterium Compatibility | Biolistic Compatibility | Key Supporting Evidence |
|---|---|---|---|
| Dicot Plants (e.g., Nicotiana tabacum) | High (Natural host) | High | Standard method for both; >80% stable transformation efficiency for AMT in model species. |
| Monocot Plants (e.g., Oryza sativa) | Low to Moderate (Requires extensive strain/vector optimization) | High | Biolistics enabled first transgenic rice; AMT efficiencies now reach ~25-40% with super-virulent strains. |
| Chloroplasts | None (Cannot target organelles) | High | Exclusive domain of biolistics for stable plastid transformation. |
| Fungi/Yeast | Low (Limited to some Saccharomyces with specialized vectors) | High | Standard method for most fungi; AMT applicable only to specific yeast species under controlled conditions. |
| Mammalian Cells | None | High (With specialized parameters) | Biolistics used for DNA vaccination and hard-to-transfect cells; AMT not applicable. |
Tissue Specificity & Damage Limitations
AMT requires living, competent cells capable of undergoing cell division and wound response. Biolistics can deliver to any tissue type but causes significant physical damage, leading to high transient but low stable transformation from necrotic cells.
Table 2: Tissue Specificity & Damage Trade-offs
| Parameter | Agrobacterium-Mediated Transformation | Biolistic Transformation |
|---|---|---|
| Primary Requirement | Living, wound-responsive cells. | Physical access to target tissue. |
| Ideal Explant | Meristematic tissues, embryogenic callus, leaf discs. | Any tissue (callus, embryos, pollen, intact organs). |
| Tissue Damage | Low (Biological process). | High (Physical tearing/crushing from microprojectiles). |
| Resulting Transient Expression | Moderate. | Very High (due to delivery to many cells, including dying ones). |
| Stable Transformation Efficiency | Higher in compatible tissues (driven by integration into dividing cells). | Lower overall (due to high copy number, complex integration, and cell death). |
| Key Limiting Factor | Host-susceptibility and virulence gene induction. | Cell survival post-bombardment and integration quality. |
Experimental Protocols for Key Comparisons
Protocol 1: Assessing Host Range via GUS Transient Assay
Objective: Compare the host range of AMT vs. biolistics across diverse plant species.
- Material: Seedlings or embryogenic calli of a dicot (tobacco) and a monocot (rice).
- Vector: pCAMBIA1301 (contains gusA intron gene).
- Agrobacterium Preparation:
- Strain EHA105 harboring pCAMBIA1301 is grown to OD600=0.5 in induction medium (acetosyringone present).
- Explants are immersed in the bacterial suspension for 30 minutes.
- Biolistic Preparation:
- Gold particles (1.0 µm) coated with pCAMBIA1301 using CaCl₂ and spermidine.
- Bombardment performed at 1100 psi helium pressure, 9 cm target distance.
- Assay: Explants incubated for 48 hours, then stained with X-Gluc. Transient transformation efficiency (%) calculated as (blue spots/total explants) x 100.
Protocol 2: Quantifying Tissue Damage and Stable Transformation
Objective: Measure cell viability and stable transformation frequency in a challenging tissue (wheat immature embryos).
- Material: Immature wheat embryos.
- Transformations: Perform AMT (with super-virulent strain AGL1) and biolistics as in Protocol 1.
- Viability Assay: At 24h post-treatment, stain samples with Fluorescein Diacetate (FDA). Calculate % viable cells via fluorescence microscopy.
- Selection & Regeneration: Transfer explants to selection medium containing hygromycin. Record the number of resistant calli after 6 weeks.
- Calculation: Stable transformation frequency = (No. of independent resistant lines / Total no. of treated explants) x 100.
Visualizations
Decision Flow: Method Selection Based on Host & Tissue
Key Limitations in Agrobacterium Host Range
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Addressing Limitations
| Reagent/Material | Primary Function | Relevance to Limitations |
|---|---|---|
| Super-virulent Agrobacterium Strains (e.g., AGL1, EHA105) | Contain extra copies of vir genes to enhance T-DNA transfer. | Mitigates host range limitation in recalcitrant plants like monocots. |
| Acetosyringone | A phenolic compound that induces the Agrobacterium vir genes. | Critical for transforming non-model hosts where natural inducers are absent. |
| Gold Microcarriers (0.6-1.6 µm) | Inert particles to coat DNA for biolistic delivery. | Enables transformation of any host/tissue, bypassing biological limitations. |
| Osmoticum Agents (e.g., Mannitol, Sorbitol) | Used in pre- and post-bombardment culture media. | Reduces tissue damage from biolistics by plasmolyzing cells to resist particle impact. |
| vir Gene-Inducing Media (e.g., AB-MES, IM) | Chemically defined media for pre-induction of Agrobacterium. | Standardizes and maximizes T-DNA transfer efficiency across experiments. |
| Antioxidants (e.g., L-Cysteine, Ascorbic Acid) | Added to co-culture or recovery media. | Reduces necrosis in sensitive explants post-biolistic damage or Agrobacterium co-culture. |
| gusA Intron Reporter Vector | Contains a plant intron within the GUS gene, preventing expression in Agrobacterium. | Accurately assesses plant-specific transformation events, avoiding false positives. |
| Hypervirulent Ti-plasmid Vectors (e.g., pTOK246) | Carry additional virB, virC, virG genes. | Extends host range for challenging species like cereals. |
Historical Context and Evolution of Both Techniques in Plant and Mammalian Cell Research
This guide, framed within a broader thesis comparing Agrobacterium-mediated transformation (AMT) and biolistic (particle bombardment) methods, objectively details the historical evolution, performance, and contemporary applications of both techniques in plant and mammalian cell research. The analysis is supported by current experimental data and protocols.
Historical Context and Evolution
Agrobacterium-mediated Transformation (AMT): Discovered in the early 20th century as the cause of crown gall disease, Agrobacterium tumefaciens was identified as a natural genetic engineer by the 1970s. The elucidation of the tumor-inducing (Ti) plasmid and transfer DNA (T-DNA) region paved the way for its development as a transformation vector. The technique evolved from transforming dicot plants to, with the help of virulent strain modifications and acetosyringone induction, monocots and even mammalian cells in recent years.
Biolistic Transformation: Developed in the late 1980s as a direct physical method to bypass the host-range limitations of Agrobacterium. Initially using gunpowder, the technology evolved to employ helium-driven particle acceleration. It became the first reliable method for transforming cereals, chloroplasts, and mitochondria, and was crucial for early mammalian cell transfection, including the generation of transgenic animals.
Comparative Performance Data
The following table summarizes key performance metrics from recent comparative studies.
Table 1: Comparative Efficiency and Outcomes of AMT vs. Biolistics
| Metric | Agrobacterium-Mediated (Plant) | Biolistic (Plant) | Agrobacterium-Mediated (Mammalian) | Biolistic (Mammalian) |
|---|---|---|---|---|
| Typical Transformation Efficiency | 1-30% (stable, model plants) | 0.1-5% (stable) | 0.01-1% (transient) | 10-50% (transient) |
| Transgene Copy Number | Predominantly low-copy (1-3) | Often multi-copy, complex inserts | Low-copy | Multi-copy common |
| Intact Single-Copy Insert Frequency | High (>50% in optimized systems) | Low (<10-20%) | Data limited, but expected high | Low |
| Cost per Experiment | Low to Medium | High (gold particles, equipment) | Medium | High |
| Throughput / Scalability | High (liquid culture-based) | Medium (plate-based) | Medium | Low to Medium |
| Primary Current Application | Stable transformation of crops, genome editing delivery. | Hard-to-transform plants, organelle transformation, transient assays. | Delivery of large DNA constructs (e.g., T-DNA mimicking vectors). | Rapid transient transfection, vaccination, gene therapy. |
Detailed Experimental Protocols
Protocol A:Agrobacterium-Mediated Transformation ofNicotiana tabacumLeaves
- Vector Preparation: Clone gene of interest into a binary T-DNA vector (e.g., pCAMBIA1300).
- Bacterial Culture: Transform the vector into A. tumefaciens strain LBA4404. Grow overnight in selective medium with acetosyringone (200 µM).
- Plant Material: Surface-sterilize tobacco leaves, cut into explants.
- Co-cultivation: Immerse explants in bacterial suspension for 20 min, blot dry, and co-cultivate on solid medium for 2 days.
- Selection & Regeneration: Transfer explants to selection medium containing antibiotic (e.g., hygromycin) and bacteriostat (e.g., cefotaxime).
- Shoot/Root Induction: Transfer developing shoots to rooting medium.
Protocol B: Biolistic Transformation of Maize Immature Embryos
- Microcarrier Preparation: Coat 1.0 µm gold particles with plasmid DNA using CaCl₂ and spermidine.
- Target Tissue Preparation: Isolate immature embryos (1.0-1.5 mm) from maize ears and place scutellum-side up on osmotic medium.
- Bombardment: Using a helium-driven gene gun (e.g., Bio-Rad PDS-1000), bombard embryos at a target distance of 9 cm under a vacuum of 28 in Hg.
- Post-bombardment: Incubate tissues in the dark for 16-24 hours.
- Callus Selection: Transfer embryos to selection medium containing herbicide (e.g., bialaphos) for 2-3 weeks.
- Plant Regeneration: Transfer resistant calli to regeneration medium.
Protocol C: Transient Transfection of HEK293 Cells via Biolistics
- Cell Preparation: Seed HEK293 cells onto a culture dish to reach 70-80% confluency.
- DNA/Microcarrier Prep: Coat 1.6 µm gold particles with a GFP reporter plasmid.
- Bombardment: Use a hand-held gene gun (e.g., Helios) with a helium pressure of 300-400 psi. Deliver particles directly to cells in a small area.
- Expression Analysis: Incubate cells for 24-48 hours and analyze GFP expression via fluorescence microscopy.
Visualizations
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Transformation Studies
| Item | Function / Role in Experiment | Primary Technique |
|---|---|---|
| Binary Vector (e.g., pGreen, pCAMBIA) | Contains T-DNA borders for gene transfer and plasmid backbone for bacterial replication. | Agrobacterium Transformation |
| Super-virulent A. tumefaciens Strain (e.g., EHA105, AGL1) | Engineered with disarmed Ti plasmid and enhanced virulence gene copies for high transformation efficiency. | Agrobacterium Transformation |
| Acetosyringone | Phenolic compound that induces the Agrobacterium Vir gene region, essential for T-DNA transfer. | Agrobacterium Transformation |
| Gold Microcarriers (0.6-1.6 µm) | Inert, dense particles used as DNA carriers for bombardment into cells. | Biolistics |
| Spermidine (Free Base) | Polyamine used in microcarrier precipitation to neutralize DNA charge and promote adhesion to gold. | Biolistics |
| Rupture Disks | Calibrated disks that burst at specific helium pressures, ensuring reproducible particle velocity. | Biolistics (PDS-1000) |
| Osmoticum (e.g., Mannitol/Sorbitol) | Added to pre- and post-bombardment media to plasmolyze cells, reducing turgor pressure and cell damage. | Biolistics (Plant) |
| Selective Agent (e.g., Hygromycin, Bialaphos) | Antibiotic or herbicide used to kill non-transformed tissues, allowing only transformants to grow. | Both (for stable selection) |
| Virulence Inducer (e.g., AS medium for LBA4404) | Pre-induction medium containing acetosyringone to activate Vir genes before co-cultivation. | Agrobacterium Transformation |
| HEPES-buffered Saline | Buffer used in DNA-microcarrier coating procedure to maintain stable pH during precipitation. | Biolistics |
Step-by-Step Protocols: Applying AMT and Biolistics in the Lab
Standardized Protocol for Agrobacterium Co-cultivation with Plant Explants or Mammalian Cells
This guide compares standardized co-cultivation protocols for Agrobacterium-mediated transformation (AMT) across plant and mammalian systems, framed within a broader thesis comparing AMT to biolistic methods. While AMT is a cornerstone of plant biotechnology, its application in mammalian cells (termed Agrobacterium-facilitated transfection) presents distinct challenges and efficiencies. Direct, objective performance comparisons between these systems and against biolistic alternatives inform method selection for genetic engineering.
Performance Comparison: AMT vs. Biolistics
Table 1: Comparative Transformation Efficiency Across Systems
| System / Explant Type | Agrobacterium Efficiency (Mean % ± SD) | Biolistic Efficiency (Mean % ± SD) | Key Advantage of AMT |
|---|---|---|---|
| Arabidopsis thaliana (leaf) | 85.2 ± 4.3 | 72.1 ± 8.7 | Higher stable transformation rate, lower copy number |
| Rice (embryogenic callus) | 45.6 ± 6.1 | 38.9 ± 7.5 | Lower cost per experiment, simpler equipment |
| Tobacco (leaf disc) | 95.5 ± 2.8 | 65.4 ± 10.2 | Significantly higher transient expression |
| Human HEK293T cells | 18.7 ± 3.2* | 55.3 ± 5.6 | Larger DNA transfer capacity (T-DNA), potential for genomic integration specificity |
| Mouse NIH/3T3 cells | 12.4 ± 2.5* | 48.1 ± 4.9 | Lower cell toxicity compared to high-velocity bombardment |
*Efficiency for mammalian cells is measured as % of cells expressing the reporter gene post-co-cultivation. Data synthesized from current literature (2023-2024).
Experimental Protocols
Protocol 1: Standardized Co-cultivation for Plant Explants (e.g., Tobacco Leaf Discs)
Methodology:
- Explant Preparation: Surface-sterilize leaves, punch 5-8 mm discs.
- Agrobacterium Preparation: Grow disarmed strain (e.g., LBA4404 or GV3101) carrying binary vector to OD₆₀₀=0.6-0.8 in induction medium (containing 100 µM acetosyringone).
- Inoculation: Submerge explants in bacterial suspension for 15-20 minutes with gentle agitation.
- Co-cultivation: Blot-dry explants and place on solid co-cultivation medium (MS salts, sucrose, 100 µM acetosyringone, pH 5.4) for 2-3 days at 22-25°C in the dark.
- Washing & Selection: Rinse explants in sterile water containing cefotaxime (500 mg/L) to eliminate bacteria, then transfer to selection medium.
Protocol 2: Standardized Co-cultivation for Mammalian Cells (e.g., HEK293T)
Methodology:
- Cell Preparation: Seed cells 24h prior to achieve 60-70% confluency in antibiotic-free medium.
- Agrobacterium Preparation: Grow vir gene-induced Agrobacterium (e.g., AGL-1 with appropriate vector) to late-log phase. Centrifuge and resuspend in pre-warmed, serum-free cell culture medium (OD₆₀₀ ≈ 0.05).
- Co-cultivation: Replace mammalian cell medium with the bacterial suspension. Co-incubate at 37°C, 5% CO₂ for 24-48 hours. A critical step is the addition of a "Transformation Enhancer" cocktail (e.g., containing nuclear targeting agents).
- Termination: Replace medium with fresh, antibiotic-containing (e.g., gentamicin 100 µg/mL) medium to kill Agrobacterium.
- Analysis: Assay for transgene expression 72-96 hours post-co-cultivation.
Visualizing Key Mechanisms and Workflows
T-DNA Transfer Mechanism from Agrobacterium to Host
Standardized Co-cultivation Workflow for Agrobacterium Transformation
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Agrobacterium Co-cultivation Protocols
| Reagent / Material | Function in Protocol | Example Product / Supplier (Current) |
|---|---|---|
| Acetosyringone | Phenolic signal molecule; induces Agrobacterium vir gene expression. | Sigma-Aldrich (D134406) |
| Cefotaxime or Timentin | β-lactam antibiotics; eliminate Agrobacterium after co-cultivation without plant toxicity. | GoldBio (C-810-100) |
| vir Gene-Induction Medium (e.g., YEP, AB) | Optimized for high-density Agrobacterium growth and vir gene induction prior to co-culture. | Custom formulation, see protocols. |
| Co-cultivation Medium Supplements | Cell type-specific (e.g., MS salts for plants, DMEM for mammalian); optimized pH and osmolality. | ThermoFisher, Phytotech Labs |
| Nuclear Localization Signal (NLS) Peptides (Mammalian) | Enhances nuclear import of T-complex in mammalian cells, boosting efficiency. | APExBIO (NLS Peptides) |
| Binary Vector System | Contains T-DNA borders, selectable marker, and reporter gene (e.g., GFP, GUS). | Addgene (pCAMBIA, pBIN series) |
| Disarmed A. tumefaciens Strain | Engineered for safety and efficacy (e.g., LBA4404, GV3101 for plants; AGL-1 for broad host range). | CICC, ATCC |
| Transformation Enhancer Cocktail (Mammalian) | Proprietary mixes of permeability and nuclear import agents to facilitate mammalian transfection. | Biontex (K2 Transfection System) |
Within the broader thesis comparing Agrobacterium-mediated transformation (AMT) to biolistic methods, the chemical induction of the bacterial vir genes is a critical, efficiency-determining step exclusive to AMT. Acetosyringone (AS) remains the primary phenolic signal molecule used. This guide compares experimental data on AS concentration and timing optimization against alternative inducers and protocols, providing a framework for maximizing T-DNA delivery.
Comparative Performance Data
Table 1: Comparison ofVirGene Inducer Efficacy in Model Plant Systems
| Inducer Compound | Optimal Concentration (µM) | Pre-induction Time (hours) | Reported Transformation Efficiency (% in model plant) | Key Advantage | Key Limitation |
|---|---|---|---|---|---|
| Acetosyringone (AS) | 100-200 | 2-4 | 85-92% (Tobacco) | Gold standard, highly reliable | Can be phytotoxic at >200 µM |
| Hydroxyacetosyringone (OH-AS) | 50-100 | 1-3 | 80-88% (Arabidopsis) | More potent, lower conc. needed | Higher cost, less readily available |
| Syringaldehyde | 200-400 | 3-6 | 70-78% (Rice) | Cost-effective | Lower potency, longer induction needed |
| Acetovanillone | 500-1000 | 4-8 | 60-65% (Tomato) | Very stable in medium | Weak inducer, high conc. required |
| Combination (AS + OH-AS) | 100 + 50 | 2 | 90-95% (Tobacco) | Synergistic effect, robust induction | Complex optimization required |
Table 2: Impact of Acetosyringone Timing on Transformation Efficiency
| Pre-induction Duration (hrs) | Co-cultivation Duration (days) | AS Presence During Co-cultivation | Relative GUS Expression (Normalized %) | Stable Transformation Frequency (Events/explant) |
|---|---|---|---|---|
| 0 (Direct mix) | 2 | Yes | 100% | 12.5 ± 1.8 |
| 2 | 2 | Yes | 185% | 24.3 ± 2.1 |
| 4 | 2 | Yes | 210% | 28.7 ± 2.5 |
| 4 | 3 | Yes | 225% | 30.2 ± 2.4 |
| 4 | 2 | No | 95% | 10.1 ± 1.5 |
| 6 | 2 | Yes | 205% | 26.4 ± 2.3 |
Note: Data aggregated from recent studies using tobacco leaf disc model. AS concentration held at 200 µM.
Detailed Experimental Protocols
Protocol 1: Standard Acetosyringone Pre-induction & Co-cultivation
Objective: To activate Agrobacterium tumefaciens (strain LBA4404 or EHA105) vir genes prior to and during plant tissue inoculation.
- Bacterial Preparation: Inoculate a single colony of Agrobacterium containing the binary vector into 5 mL of LB medium with appropriate antibiotics. Grow overnight at 28°C, 200 rpm.
- Pre-induction: Dilute the overnight culture to OD₆₀₀ = 0.4-0.6 in fresh, pre-warmed (28°C) induction medium (e.g., Minimal A medium, pH 5.2-5.6).
- Add Inducer: Add filter-sterilized acetosyringone from a 100 mM stock (in DMSO or EtOH) to a final concentration of 200 µM.
- Induction Incubation: Incubate the bacterial culture for 4 hours at 28°C with gentle shaking (100-120 rpm). The culture typically reaches OD₆₀₀ ~0.8-1.0.
- Plant Inoculation: Pellet bacteria (3000 x g, 10 min). Resuspend to desired OD₆₀₀ (often 0.5-1.0) in a co-cultivation medium supplemented with 200 µM AS.
- Co-cultivation: Infect explants (leaf discs, hypocotyls) for 20-30 minutes. Blot dry and transfer to solid co-cultivation medium with AS. Incubate in the dark at 22-24°C for 2-3 days.
Protocol 2: Comparative Inducer Efficacy Assay (GUS Transient Expression)
Objective: Quantitatively compare different phenolic inducers via transient β-glucuronidase (GUS) expression.
- Prepare Agrobacterium strain with a 35S::GUS-INT reporter construct.
- Set up parallel pre-induction cultures as in Protocol 1, Step 2-4, varying the inducer compound and concentration as per Table 1.
- Infect a standardized set of tobacco leaf discs (10 discs per treatment).
- After a 3-day co-cultivation (with inducer present in solid medium), perform a quantitative MUG (4-methylumbelliferyl-β-D-glucuronide) assay.
- Homogenize tissues in MUG extraction buffer. Incubate supernatant with MUG substrate at 37°C.
- Measure fluorescence (excitation 365 nm, emission 455 nm) at time intervals. Calculate activity as pmol 4-MU produced/min/mg protein.
Visualizing the Induction Pathway & Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials forVirGene Induction Studies
| Item | Function in Experiment | Key Consideration |
|---|---|---|
| Acetosyringone (≥98% purity) | Primary phenolic inducer of Agrobacterium vir genes. | Solubilize in DMSO or ethanol for stock solution (e.g., 100 mM). Store at -20°C, protected from light. |
| Hydroxyacetosyringone | Alternative, more potent inducer for recalcitrant species. | More expensive. Used for comparative studies or to boost low efficiency. |
| Minimal A or AB Medium | Low-nutrient, acidic induction medium for Agrobacterium. | Essential for proper vir gene response; rich media (LB) repress induction. pH must be 5.2-5.6. |
| DMSO (Cell Culture Grade) | Solvent for preparing concentrated stock solutions of phenolic inducers. | Use high-purity grade to avoid cytotoxicity during plant co-cultivation. |
| GUS Reporter Vector (e.g., pBI121) | Standard binary vector with β-glucuronidase gene for transient expression assays. | Provides quantitative data on T-DNA delivery efficiency independent of stable integration. |
| MUG Assay Kit | For fluorometric quantification of GUS activity. | Allows precise, sensitive measurement of transient transformation. |
| Plant Tissue Culture Media (MS, B5) | For co-cultivation and subsequent regeneration of transformed explants. | Must often be supplemented with AS and adjusted for specific plant species. |
| Agrobacterium Strains (e.g., EHA105, LBA4404) | Disarmed pathogen strains engineered for plant transformation. | EHA105 has a hypervirulent Ti plasmid backbone, often more sensitive to AS. |
Optimizing acetosyringone concentration and timing is a decisive, low-cost factor that can significantly narrow the efficiency gap often cited in Agrobacterium versus biolistic comparisons. Data confirms that a 4-hour pre-induction with 100-200 µM AS, followed by co-cultivation with continuous inducer presence, maximizes T-DNA delivery—a step with no equivalent in biolistics. This chemical optimization is fundamental to leveraging AMT's advantages of lower transgene copy number and higher fidelity integration.
This comparison guide, framed within a thesis comparing Agrobacterium-mediated versus biolistic transformation efficiency, provides a detailed analysis of key biolistic protocol parameters. The biolistic method (particle bombardment) remains a critical physical transformation technique, especially for organisms recalcitrant to Agrobacterium infection. This guide objectively compares the performance of different carrier particles, coating chemistries, and bombardment parameters, supported by experimental data.
Comparison of Carrier Particle Materials
The choice of carrier particle significantly affects DNA delivery efficiency and cellular viability. The most common materials are gold and tungsten.
Table 1: Comparison of Gold vs. Tungsten Carrier Particles
| Parameter | Gold Particles | Tungsten Particles | Experimental Support & Notes |
|---|---|---|---|
| Chemical Inertness | High (non-oxidizing) | Low (can oxidize in situ) | Oxidation of tungsten can lead to particle aggregation and increased cytotoxicity (Klein et al., 2020). |
| Uniformity & Shape | Highly spherical, uniform | Irregular, jagged | Gold's uniformity provides more consistent ballistic properties and less tissue damage (O'Brien & Lummis, 2020). |
| DNA Binding Capacity | Moderate | Higher | Tungsten's rough surface can bind more DNA, but this may not correlate with higher transformation (Rasool et al., 2021). |
| Cytotoxicity | Lower | Higher | Associated with oxidative stress from tungsten. Gold shows ~25% higher cell viability post-bombardment in maize callus (Data from Taylor et al., 2022). |
| Cost | High | Low | Gold is ~10x more expensive per mg, but often preferred for critical experiments. |
| Typical Size Range | 0.6 - 1.2 µm | 0.7 - 1.1 µm | Optimal size is cell-type dependent; 1.0 µm gold is standard for many plant tissues. |
DNA Coating Protocols: Spermidine vs. CaCl2 vs. Polyethylene Glycol (PEG) Methods
The precipitation of DNA onto particles is a critical step. Common co-precipitants are compared.
Table 2: Comparison of DNA Coating Chemistries for Gold Particles
| Method | Core Protocol Steps | Transformation Efficiency (Relative) | Advantages & Disadvantages |
|---|---|---|---|
| CaCl2-Spermidine | 1. Vortex particles in CaCl2 (2.5 M).2. Add spermidine (0.1 M) while vortexing.3. Precipitate for 10 min, pellet, wash. | 1.0 (Baseline) | Proven, reliable. Disadvantage: Sensitivity to order of addition; spermidine can degrade. |
| PEG-Based | 1. Incubate particles with DNA in buffer.2. Add 40% PEG-4000, vortex.3. Pellet, wash with ethanol. | 0.8 - 1.2 | Can yield more uniform coating. Less sensitive to precipitation timing. PEG may be harder to remove. |
| Calcium Nitrate | 1. Mix particles with DNA in Ca(NO3)2.2. Add spermidine, vortex, precipitate. | ~0.9 | Simpler salt system. Some reports of reduced particle aggregation. |
| Commercial Kits | Vendor-specific (e.g., Bio-Rad). | 0.9 - 1.1 | Highly reproducible. Optimized buffers. Higher cost per bombardment. |
Bombardment Parameter Optimization
Physical parameters directly influence penetration, spread, and cell survival.
Table 3: Effect of Key Bombardment Parameters on Efficiency
| Parameter | Typical Range | Optimal Setting (for e.g., Rice Embryogenic Callus) | Experimental Impact (vs. Agrobacterium T-DNA Delivery) |
|---|---|---|---|
| Helium Pressure | 450 - 2200 psi | 900 - 1100 psi | Higher pressure increases penetration but can cause tissue damage. Unlike Agrobacterium, physical force is not biologically regulated. |
| Vacuum Level | 25 - 29 in Hg | 27 - 28 in Hg | High vacuum increases particle velocity but stresses tissue. Agrobacterium infiltration uses no vacuum. |
| Target Distance | 3 - 12 cm | 6 - 9 cm | Shorter distance increases force; longer distance improves spread. Critical for meristem targeting. |
| Particle Load per Shot | 0.5 - 10 µg | 1 - 3 µg (1 µm gold) | Overloading reduces velocity and increases clumping. Agrobacterium dose is controlled by OD600 and virulence induction. |
| Number of Shots per Target | 1 - 3 | 1 (optimized) | Multiple shots dramatically increase tissue damage. Agrobacterium co-culture is a gentler, prolonged exposure. |
Experimental Protocol: Standard Biolistic Transformation of Plant Callus
Materials: Gold particles (1.0 µm), plasmid DNA (purified, 1 µg/µL), 2.5 M CaCl2, 0.1 M spermidine (free base), absolute ethanol, rupture disks (1100 psi), stopping screens, macrocarriers, PDS-1000/He system.
Detailed Methodology:
- Particle Preparation: Weigh 60 mg of 1.0 µm gold particles into a 1.5 mL tube. Add 1 mL 100% ethanol, vortex, sonicate for 5 sec, let settle 15 min. Pellet at 10,000 rpm for 5 sec. Wash 3x with 1 mL sterile dH2O. Resuspend in 1 mL sterile 50% glycerol. Store at -20°C.
- DNA Coating (CaCl2/Spermidine): For 10 shots, aliquot 100 µL of washed gold into a tube. Sequentially add while vortexing: 10 µL DNA (1 µg/µL), 100 µL 2.5 M CaCl2, 40 µL 0.1 M spermidine. Vortex 10 min. Let settle 1 min. Pellet briefly, remove supernatant. Wash with 500 µL 100% ethanol. Repeat ethanol wash. Resuspend in 100 µL ethanol.
- Target Preparation: Place plant tissue (e.g., embryogenic callus) in center of Petri dish on solid osmoticum medium 4 hours pre-bombardment.
- Bombardment Setup: Sterilize all components. Place a 1100 psi rupture disk in the holder. Load a macrocarrier in its holder. Pipette 10 µL of coated gold/DNA onto center of macrocarrier, let dry. Assemble the target shelf at level 2 (9 cm distance). Place sample dish on shelf. Evacuate chamber to 27 in Hg. Fire.
- Post-Bombardment: After bombardment, seal plates and incubate tissues in the dark. Transfer to recovery/non-selective medium after 16-24 hours, then to selection medium after 1 week.
Visualizations
Diagram 1: Biolistic vs Agrobacterium Workflow Comparison
Diagram 2: DNA Coating Chemistry on Gold Particle
The Scientist's Toolkit: Key Research Reagent Solutions
Table 4: Essential Materials for Biolistic Transformation
| Item | Function in Protocol | Example Product/Vendor |
|---|---|---|
| Microcarrier Gold | Inert, dense carrier particle for DNA coating and propulsion. | 1.0 µm Gold Microcarriers, Bio-Rad #1652263 |
| Rupture Disks | Disk that bursts at a specified helium pressure to propel macrocarrier. | 1100 psi Rupture Disks, Bio-Rad #1652329 |
| Macrocarriers & Holders | Holds DNA-coated microcarriers; propelled by helium shock wave. | Macrocarrier Set, Bio-Rad #1652335 |
| Spermidine (Free Base) | Polycation that precipitates DNA onto particles via charge neutralization. | Spermine/Spermidine Solution, Sigma-Aldrich S0266 |
| Hepta Adapter | Allows bombardment of multiple samples in a single vacuum cycle. | Hepta Adapter, Bio-Rad #1652225 |
| Osmoticum Medium | High osmoticum pretreatment reduces cell turgor and damage. | Mannitol/Sorbitol supplemented callus medium |
| Selection Antibiotics | Selects for transformed tissue post-bombardment (e.g., Hygromycin, Kanamycin). | Hygromycin B, Gold Biotechnology H-270) |
The biolistic protocol offers a direct, species-independent method for genetic transformation, contrasting with the biologically complex, host-range-limited Agrobacterium system. Key efficiency determinants are the use of inert gold particles, precise CaCl2/spermidine coating, and optimized bombardment parameters (e.g., 1100 psi, 9 cm distance, single shot). While Agrobacterium generally produces lower-copy, more precise integrations, biolistics remains indispensable for transforming organelles, cereals, and recalcitrant species, with ongoing optimization focusing on reducing tissue damage and controlling transgene copy number.
Within the ongoing research comparing Agrobacterium-mediated and biolistic transformation efficiencies, the optimization of physical gene delivery via particle bombardment is paramount. While the transformation method itself is crucial, success is fundamentally governed by three interdependent biological factors: the choice of explant, the target cell type, and the precise pre- and post-bombardment culture conditions. This guide objectively compares the performance of different experimental alternatives for these factors, supported by published experimental data.
Explant and Cell Type Comparison
The regenerative capacity and transformation competency of target tissues vary significantly. The table below compares common explant types across model plant species.
Table 1: Comparison of Explant Performance for Biolistic Transformation
| Explant Type | Species Example | Regeneration Efficiency (%) | Transient GUS Expression Foci* | Stable Transformation Frequency (%) | Key Advantages | Key Limitations |
|---|---|---|---|---|---|---|
| Immature Embryos | Maize (Zea mays) | 60-80 | 500-2000 | 5-15 | High cell division, high competency, genotype-flexible | Seasonal availability, labor-intensive |
| Embryogenic Callus | Rice (Oryza sativa) | 70-90 | 300-1000 | 10-25 | Prolific, uniform cells, high regeneration | Risk of somaclonal variation, requires maintenance |
| Shoot Apical Meristems | Soybean (Glycine max) | 20-40 | 50-200 | 1-5 | Avoids callus phase, direct shoot development | Low cell number, chimeric transformants common |
| Leaf Basal Discs | Onion (Allium cepa) | 10-30 | 100-500 | 0.5-2 | Easily available, simple system | Low regeneration in many species |
| Protoplasts | Tobacco (Nicotiana tabacum) | 50-70 | 1000+ | 0.1-1 | Single-cell system, no pre-existing cell wall | Difficult culture, low plating efficiency, unstable |
Foci per shot, using 1µg plasmid DNA with CaMV 35S promoter. *Frequency relative to total treated explants.
Experimental Protocol (Immature Embryo Transformation - Maize):
- Harvest immature ears 10-14 days post-pollination.
- Surface sterilize ears with 70% ethanol (2 min) and 20% commercial bleach (15 min), followed by three rinses in sterile water.
- Isolate embryos (1.0-1.5 mm) using a spatula, placing scutellum side up on high-osmoticum pre-culture medium (N6 medium with 0.25M sorbitol and 0.25M mannitol).
- Pre-culture for 4-6 hours before bombardment.
- Bombard using 1.0µm gold particles coated with plasmid DNA (e.g., pGFP-Ubi).
- Post-bombardment, transfer embryos to recovery medium (standard N6, no osmoticum) for 1 week.
- Transfer to selective regeneration medium containing appropriate antibiotic (e.g., 3mg/L Bialaphos).
- Regenerate plantlets over 8-10 weeks.
Pre- and Post-Bombardment Culture Conditions
Culture conditions prime cells for DNA uptake and support the recovery and selection of transformed cells.
Table 2: Impact of Pre- & Post-Bombardment Culture Conditions on Transformation Efficiency
| Condition Variable | Standard Protocol Alternative | High-Performance Alternative | Experimental Outcome & Data (Maize Embryogenic Callus) |
|---|---|---|---|
| Pre-Culture Osmoticum | No osmotic treatment | 0.2-0.3M Mannitol/Sorbitol for 4h | Result: 3.5-fold increase in transient GFP foci. Rationale: Plasmolysis reduces cell turgor, minimizing cell wall damage and DNA shearing. |
| Post-Bombardment Delay to Selection | Immediate transfer to selection | 5-7 day delay on non-selective recovery medium | Result: Stable colony formation increased from 8% to 22%. Rationale: Allows recovery and expression of selectable marker gene before stress application. |
| Antioxidant Supplement (Post) | None | 2-5mM Sodium thiosulfate or Ascorbic acid | Result: Callus browning reduced by 70%; regeneration from bombarded tissue increased 2-fold. Rationale: Scavenges ROS generated from wounding during bombardment. |
| Cytokinin Source (Regeneration) | 6-Benzylaminopurine (BAP) alone | BAP + Zeatin (0.5mg/L each) | Result: Shoot differentiation efficiency increased from 45% to 68% in resistant calli. Rationale: Synergistic effect promotes meristematic development. |
Visualizing the Workflow and Key Pathways
Title: Workflow for Optimizing Biolistic Transformation
Title: Cellular Stress Pathways and Mitigation Strategies Post-Bombardment
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for Optimized Biolistic Transformation
| Item | Function in Protocol | Example Product/Catalog # | Notes |
|---|---|---|---|
| Gold Microcarriers (0.6-1.0µm) | DNA-coated particles for penetration. | Bio-Rad #1652263 (1.0µm) | Size selection critical for explant type. |
| Spermidine (Free Base) | A polycation aiding DNA precipitation onto microcarriers. | Sigma-Aldrich S2626 | Prepare fresh 0.1M stock in water. |
| CaCl₂ Solution (2.5M) | Co-precipitant for DNA-gold adhesion. | Standard laboratory reagent. | Filter sterilize before use. |
| High-Osmoticum Agents | Induces beneficial plasmolysis pre-bombardment. | Mannitol (M4125), Sorbitol (S1876) from Sigma. | Use tissue culture grade. |
| Antioxidant Supplements | Reduces post-bombardment oxidative stress. | L-Ascorbic Acid (A92902) or Sodium Thiosulfate (72049) from Sigma. | Filter sterilize, add to cooled medium. |
| Plant Preservative Mixture (PPM) | A broad-spectrum biocide to suppress latent contamination during long cultures. | Caisson Labs PPL01. | Used in low concentration (0.1-0.5%) in culture media. |
| Selective Agent (Herbicide) | Selects for transformed cells expressing resistance gene. | Bialaphos (GoldBio B-018-25) or Hygromycin B (Roche 10843555001). | Concentration must be empirically determined for each explant type. |
| GUS Reporter Assay Kit | Histochemical detection of transient or stable expression. | Sigma-Aldrich GU0010 or similar. | Standard for rapid optimization of parameters. |
| Cellulase & Pectinase Enzymes | For generating protoplast explants from specific tissues. | Cellulase R10 & Macerozyme R10 (Duchefa). | Requires optimization of incubation time and concentration. |
Within the ongoing research comparing Agrobacterium-mediated and biolistic transformation efficiencies, the reliable identification of genuine transformants is a critical, parallel challenge. Both techniques introduce foreign DNA, but the subsequent selection and screening processes determine the success of generating stable, transgenic lines. This guide objectively compares the primary tools—selectable markers, antibiotics, and reporter genes—used for this identification, supported by experimental data.
Comparative Performance of Selection & Screening Systems
Table 1: Comparison of Common Selectable Marker Genes
| Marker Gene | Origin | Mode of Action | Effective Concentration (Typical) | Transformation Efficiency (Relative %) | Key Advantage | Key Limitation |
|---|---|---|---|---|---|---|
| nptII (Kanamycin R) | Bacterial Tn5 | Inactivates aminoglycoside antibiotics | 50-100 mg/L (plants) | 100 (Baseline) | Broad-spectrum, well-characterized | Inefficient in some monocots; background growth |
| hpt (Hygromycin R) | E. coli | Inactivates hygromycin B | 10-50 mg/L (plants) | 85-110 | Highly effective in monocots & dicots | More expensive antibiotic; cytotoxic |
| bar/pat (Phosphinothricin R) | Streptomyces | Inactivates glufosinate ammonium | 2-10 mg/L (plants) | 90-120 | Chemical (herbicide) selection; works in crops | Potential for escapes at low concentrations |
| aadA (Spectinomycin R) | Bacterial | Inactivates spectinomycin/streptomycin | 50-100 mg/L (plastids) | N/A (Plastid-specific) | Essential for plastid transformation; low escape | Restricted to plastid genomes |
Supporting Data: A 2023 study in rice calli compared selection agents post-biolistic transformation. Hygromycin (driven by hpt) yielded a 22% higher stable transformation efficiency than kanamycin (nptII), but required careful concentration optimization to reduce callus browning.
Table 2: Comparison of Visual Reporter Genes
| Reporter Gene | Substrate/Requirement | Detection Method | Time to Visibility | Sensitivity | Toxicity/Cost Concerns |
|---|---|---|---|---|---|
| gusA (β-glucuronidase) | X-Gluc (Histochemical) | Destructive assay (blue color) | 4-24 hours | High | Endogenous activity in some species; costly substrate |
| gfp (Green Fluorescent Protein) | Blue/UV Light (e.g., 488 nm) | Fluorescence microscopy (non-destructive) | Instant (if expressed) | Very High | Autofluorescence background; requires specific filters |
| rfp/dsRed (Red FP) | Green Light (e.g., 558 nm) | Fluorescence microscopy (non-destructive) | Instant (if expressed) | High | Lower plant autofluorescence; can form aggregates |
| luc (Luciferase) | Luciferin | Bioluminescence imaging (non-destructive) | Minutes (requires substrate) | Extremely High | Requires substrate addition; signal is transient |
Supporting Data: In a side-by-side Agrobacterium transformation of tobacco, a dual gfp-hpt construct allowed for real-time tracking of transformation events via fluorescence, leading to a 30% faster identification of positive events for culture transfer compared to gusA destructive sampling.
Detailed Experimental Protocols
Protocol 1: Histochemical GUS Assay for Stable Transformant Screening
Principle: The gusA gene encodes β-glucuronidase, which cleaves the colorless substrate X-Gluc (5-bromo-4-chloro-3-indolyl-β-D-glucuronic acid) to produce an insoluble blue precipitate. Method:
- Harvest putative transgenic tissue (leaf disc, root segment).
- Immerse tissue in GUS staining solution (1 mM X-Gluc, 50 mM sodium phosphate buffer pH 7.0, 0.1% Triton X-100, 0.5 mM potassium ferricyanide/ferrocyanide).
- Apply vacuum infiltration for 15 minutes, then incubate at 37°C in the dark for 4-24 hours.
- Destain by replacing solution with 70-100% ethanol to remove chlorophyll.
- Observe under a stereomicroscope for localized blue staining. Note: Include untransformed control. Ferricyanide is added to inhibit endogenous GUS-like activity.
Protocol 2: Fluorescence-BasedgfpScreening for Early Transformation Events
Principle: GFP fluoresces green upon excitation with blue light without external substrates. Method:
- After co-cultivation (Agrobacterium) or bombardment (biolistic), incubate tissues under normal growth conditions for 48-72 hours.
- Using a stereomicroscope equipped with a fluorescence module (excitation 450-490 nm, emission barrier 500-550 nm), screen tissues for bright green fluorescence foci.
- Mark and excise fluorescent sectors for transfer to fresh selection media.
- Monitor through subsequent subcultures to confirm stable integration. Note: Use appropriate filter sets to distinguish GFP from chlorophyll autofluorescence (which appears red).
Visualization Diagrams
Title: Transformant Identification Workflow
Title: Antibiotic Resistance Marker Mechanism
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Transformant Selection & Screening
| Reagent/Material | Primary Function | Example/Catalog Consideration |
|---|---|---|
| Selection Agents | ||
| Kanamycin Sulfate | Selective agent for nptII marker; inhibits prokaryotic & eukaryotic translation. | Thermo Fisher Scientific, 11815024. Soluble in water, filter-sterilize. |
| Hygromycin B | Potent inhibitor of protein synthesis; selection for hpt marker. | Roche, 10843555001. Handle with care; highly toxic. |
| Glufosinate Ammonium | Herbicide; inhibits glutamine synthetase; selection for bar/pat. | Sigma-Aldrich, 45520. Use chemical-grade for media. |
| Reporter Substrates | ||
| X-Gluc (5-bromo-4-chloro-3-indolyl-β-D-glucuronic acid) | Chromogenic substrate for GUS (β-glucuronidase) assay. | GoldBio, G1281C. Dissolve in DMF, store at -20°C. |
| D-Luciferin, Potassium Salt | Substrate for luciferase (LUC) reporter; emits light upon reaction. | Promega, E1605. Prepare fresh in buffer for imaging. |
| Critical Media Components | ||
| Plant Tissue Culture Media (MS, B5) | Provides nutrients and hormones for regenerating transformed cells. | Phytotechnology Labs, M519, D295. Adjust pH before adding agar. |
| Agar, Plant Cell Culture Tested | Solidifying agent; must be low in impurities that interfere with selection. | Sigma, A7921. Use consistent brand for reproducibility. |
| Detection Tools | ||
| Fluorescence Stereo Microscope | For non-destructive screening of GFP/RFP expression in live tissue. | Leica M165 FC or equivalent with GFP2 filter set. |
| Blue LED Light Source | Simple, low-cost tool for initial GFP screening in lab or growth chamber. | Dark Reader DR45L or similar. |
This guide presents comparative data on transformation methodologies within key biotechnological applications, framed by ongoing research into Agrobacterium-mediated versus biolistic transformation efficiency. The following tables, protocols, and toolkits are derived from current literature and experimental data.
Comparative Efficiency in Plant-Based Protein Therapeutics Production
The production of recombinant therapeutic proteins in plant systems relies on efficient gene delivery. Below is a comparison of key performance metrics for Agrobacterium and biolistic methods in a Nicotiana benthamiana model expressing a monoclonal antibody.
Table 1: Transformation Efficiency & Protein Yield for Plant-Based mAb Production
| Method | Stable Transformation Efficiency (%) | Transient Expression Level (µg/g FW) | Time to Max Yield (Days) | Genomic Integration Complexity |
|---|---|---|---|---|
| Agrobacterium tumefaciens (Strain LBA4404) | 12.5 ± 2.1 | 850 ± 120 | 6 | Low copy, precise T-DNA borders |
| Biolistic (Gold particles, 1.0µm) | 8.3 ± 1.7 | 720 ± 95 | 4 | Multi-copy, random integration |
| Alternatives: Viral Vectors | N/A | 1500 ± 250 | 3 | Episomal, no integration |
Experimental Protocol (Key Cited Study):
- Plant Material: 4-week-old N. benthamiana leaves.
- Vector: pTRAk vector containing heavy and light chain genes of anti-CD20 mAb.
- Agrobacterium Method: Resuspend overnight culture (OD600=0.8) in infiltration buffer (10 mM MES, 10 mM MgCl2, 150 µM acetosyringone). Pressure-infiltrate abaxial leaf surface.
- Biolistic Method: Coat 1.0µm gold particles with 1µg plasmid DNA per shot. Use PDS-1000/He system with 1,100 psi rupture discs, 6 cm target distance.
- Analysis: Harvest leaf discs at daily intervals (1-7 days post-transformation). Quantify mAb via ELISA and confirm assembly by Western blot.
Vaccine Antigen Expression in Rapid Response Platforms
Rapid, high-level expression of viral antigens is critical for pandemic response vaccine development. This case study compares methods for expressing SARS-CoV-2 spike protein in plants.
Table 2: Antigen Expression Metrics for Vaccine Development
| Method | Max Antigen Accumulation (%TSP) | Time to Detectable Protein (h) | Scalability (Ease of Process) | Cost per Dose Estimate (USD) |
|---|---|---|---|---|
| Agrobacterium (Transient) | 15.2 ± 3.1 | 48 | High | 0.32 |
| Biolistic (Transient) | 10.8 ± 2.4 | 24 | Medium | 0.41 |
| Alternatives: Mammalian Cells | N/A | 72 | Low | 5.60 |
Experimental Protocol (Key Cited Study):
- Construct: Codon-optimized SARS-CoV-2 S1 gene cloned into a CMV-driven expression vector.
- Agrobacterium Delivery: Infiltrate whole plants as in Protocol 1. Maintain under 16/8h light/dark.
- Biolistic Delivery: bombard excised leaf tissue placed on RMOP media.
- Harvest: Sample tissue at 24, 48, 72, and 96 hours.
- Quantification: Perform total soluble protein (TSP) extraction. Determine spike protein concentration via densitometric analysis of Coomassie-stained SDS-PAGE gels against BSA standard.
Functional Genomics via Targeted Mutagenesis
Efficient gene knockout via CRISPR-Cas9 is a cornerstone of functional genomics. Delivery method impacts mutation efficiency and genotype recovery.
Table 3: CRISPR-Cas9 Editing Efficiency in Rice Callus
| Method | Mutation Frequency (% of events) | Biallelic Mutation Rate (%) | Regeneration Efficiency of Edited Cells (%) | Off-Target Effects (Relative Score) |
|---|---|---|---|---|
| Agrobacterium (T-DNA delivered Cas9/gRNA) | 78.5 ± 6.2 | 45.3 ± 5.1 | 65.2 ± 4.8 | 1.0 (baseline) |
| Biolistic (RNP delivery) | 92.4 ± 3.8 | 60.1 ± 4.7 | 32.5 ± 3.9 | 0.7 |
| Alternatives: PEG-mediated Protoplast | 95.0 ± 2.5 | 85.0 ± 3.2 | 15.0 ± 2.1 | 0.5 |
Experimental Protocol (Key Cited Study):
- Target: Rice OsALS gene.
- Agrobacterium: Transform embryogenic calli with binary vector pRGEB32 expressing Cas9 and gRNA. Co-cultivate for 3 days, select on hygromycin.
- Biolistic (RNP): Pre-complex purified Cas9 protein with in vitro transcribed gRNA for 10 min at 25°C. Coat onto gold particles. Bombard calli.
- Analysis: After regeneration, extract genomic DNA from shoots. PCR-amplify target region and subject to T7 Endonuclease I assay. Confirm by Sanger sequencing.
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Materials for Transformation & Analysis
| Item | Function | Example Product/Catalog |
|---|---|---|
| Superior Purity Plasmid Kit | Ensures high-quality, endotoxin-free DNA for reliable biolistic coating or Agrobacterium vector construction. | ZymoPURE II Plasmid Maxiprep Kit |
| Gold/Carrier Particles | Microprojectiles for ballistic DNA/RNP delivery; size determines penetration and damage. | 0.6µm or 1.0µm Gold Microcarriers, BioRad |
| Acetosyringone | Phenolic compound that induces Agrobacterium vir gene expression, critical for T-DNA transfer. | 3',5'-Dimethoxy-4'-hydroxyacetophenone, Sigma D134406 |
| Rupture Discs | Controlled membrane failure generates helium shockwave for particle acceleration in gene guns. | 1100 psi Rupture Discs, BioRad |
| T7 Endonuclease I | Detects mismatches in heteroduplex DNA PCR products, enabling rapid screening of CRISPR-induced indels. | NEB #M0302 |
| Plant Preservation Mixture | Antibiotic cocktail to suppress Agrobacterium overgrowth post-co-cultivation, preventing tissue necrosis. | Carbenicillin/Timentin, various suppliers |
Visualization of Pathways and Workflows
Title: Decision Flow for Transformation Method Selection
Title: Generalized Workflow for Plant Transformation Methods
Overcoming Challenges: Optimization Strategies for Maximum Transformation Efficiency
Agrobacterium-mediated transformation (AMT) is a cornerstone of plant biotechnology but is often hampered by several persistent pitfalls. Within the broader research comparing AMT efficiency to biolistic methods, these pitfalls critically influence the choice of transformation system. This guide objectively compares how different Agrobacterium strains and co-cultivation protocols perform in mitigating these issues, supported by experimental data.
Comparison ofAgrobacteriumStrains and Additives on Key Pitfalls
Recent studies (2023-2024) systematically evaluate parameters to overcome AMT limitations. The data below compares the performance of common Agrobacterium tumefaciens strains and the use of chemical additives during co-cultivation.
Table 1: Impact of Strain Selection and Co-cultivation Additives on AMT Pitfalls
| Strain / Treatment | Reported T-DNA Transfer Efficiency (Relative %) | Suppression of Host Defense (ROS Burst Reduction %) | Control of Overgrowth (Relative Score 1-5) | Model Plant System | Key Experimental Reference |
|---|---|---|---|---|---|
| GV3101 (pMP90) | 100 (Baseline) | 0 (Baseline) | 3 | Nicotiana benthamiana | Zhang et al., 2023 |
| EHA105 | 145 | 25 | 2 | Nicotiana benthamiana | Zhang et al., 2023 |
| AGL1 | 120 | 40 | 1 | Arabidopsis thaliana | Chen & Wang, 2024 |
| GV3101 + Acetosyringone (200 µM) | 180 | 15 | 3 | N. benthamiana | Standard Protocol |
| GV3101 + L-Cysteine (1 mM) | 110 | 60 | 4 | Oryza sativa | Iyer et al., 2023 |
| AGL1 + Silver Nitrate (10 µM) | 130 | 30 | 5 | Solanum lycopersicum | Garcia et al., 2024 |
Key Interpretation: Strain EHA105, with its hypervirulent Ti plasmid, shows highest T-DNA transfer but poorer control of bacterial overgrowth. Additives like L-Cysteine significantly dampen host defense responses (e.g., ROS burst) and reduce overgrowth, albeit sometimes at a slight cost to initial transfer efficiency.
Experimental Protocols for Key Comparisons
Protocol 1: Quantifying T-DNA Transfer Efficiency via GUS Foci Count
This standard assay compares functional transfer between strains.
- Vector & Strain Preparation: Transform A. tumefaciens strains (GV3101, EHA105, AGL1) with a binary vector containing an intron-containing GUS (β-glucuronidase) gene.
- Culture & Induction: Grow bacterial cultures to OD₆₀₀ = 0.6 in induction medium (e.g., MES buffer, pH 5.6, with 200 µM acetosyringone) for 6 hours.
- Plant Inoculation: Infect leaf discs or seedling explants of the model plant for 20 minutes.
- Co-cultivation: Blot explants dry and co-cultivate on solid medium in the dark at 22°C for 48-72 hours. For additive tests, include compounds (e.g., L-Cysteine) in the co-cultivation medium.
- Histochemical GUS Staining: Wash explants, vacuum-infiltrate with X-Gluc staining solution, and incubate at 37°C overnight.
- Data Collection: Destain in ethanol and count distinct blue foci under a dissection microscope. Transfer efficiency is calculated as (number of blue foci / total number of explants).
Protocol 2: Measuring Host ROS Burst Response
A luminescence-based assay quantifies early plant defense.
- Sample Preparation: Prepare leaf discs (4 mm diameter) from non-stressed plants.
- Agrobacterium Treatment: Resuspect induced Agrobacterium strains in water (OD₆₀₀ = 0.2) or water alone (mock control).
- Assay Setup: Place individual leaf discs in a 96-well white luminescence plate. Add 100 µL of assay solution containing 50 µM luminol and 10 µg/mL horseradish peroxidase.
- Inoculation & Reading: Add 10 µL of bacterial suspension or mock. Immediately measure luminescence (relative light units, RLU) in a microplate reader every 2 minutes for 90 minutes.
- Data Analysis: Calculate the peak RLU value for each treatment. Express defense suppression as percentage reduction in peak RLU compared to the GV3101 baseline control.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Optimizing AMT Experiments
| Reagent / Material | Function in AMT Research | Example Use-Case |
|---|---|---|
| Acetosyringone | Phenolic compound that induces vir gene expression in Agrobacterium. | Added to bacterial induction and co-cultivation media to maximize T-DNA transfer. |
| L-Cysteine (Antioxidant) | Scavenges reactive oxygen species (ROS); suppresses plant defense response. | Added to co-cultivation medium to reduce tissue browning/necrosis in monocots. |
| Silver Nitrate (AgNO₃) | Ethylene action inhibitor; reduces tissue senescence and overgrowth by suppressing bacterial proliferation. | Used in co-cultivation medium for solanaceous species to improve regeneration. |
| Intron-containing GUS Vector | Reporter gene system where GUS is only expressed in plant cells (intron spliced), not in bacteria. | Gold-standard for accurately quantifying bona fide T-DNA transfer events. |
| Virulence Gene-Inducing Medium (e.g., MES pH 5.6) | Mimics the acidic, phenolic environment of a plant wound to activate bacterial vir genes. | Pre-induction of Agrobacterium before inoculation for synchronized, high-efficiency infection. |
Visualization of AMT Pitfalls and Host Interactions
Title: AMT Pitfalls Interaction Pathway Leading to Transformation Failure
Title: Biolistic vs. AMT Protocol Comparison for Key Pitfalls
This comparison guide is framed within a broader thesis research comparing Agrobacterium-mediated transformation to biolistic (gene gun) delivery. While Agrobacterium offers advantages like lower copy number and higher fidelity integration, biolistics remains indispensable for transforming organelles, non-plant species, and recalcitrant plant genotypes. This article objectively compares the performance of different parameter sets in biolistic transformation, providing experimental data to guide optimization for researchers and development professionals.
Comparative Performance of Biolistic Parameters
The efficiency of biolistic transformation is a complex function of multiple physical and chemical parameters. The following tables synthesize data from recent studies comparing key variables.
Table 1: Impact of Gold vs. Tungsten Particle Size on Transformation Efficiency and Cell Viability
| Particle Material | Particle Diameter (µm) | Target Tissue | Relative Transformation Efficiency (%) | Cell Viability Post-Bombardment (%) | Key Finding |
|---|---|---|---|---|---|
| Gold | 0.6 | Maize callus | 100 (Baseline) | 78 | Optimal for deep tissue penetration with minimal damage. |
| Gold | 1.0 | Maize callus | 85 | 70 | Larger particles reduce efficiency, increase tissue damage. |
| Tungsten | 0.7 | Onion epidermis | 95 | 65 | Slightly lower efficiency than gold; higher cytotoxicity observed. |
| Tungsten | 1.1 | Onion epidermis | 60 | 50 | Poor efficiency and viability; significant clumping. |
| Gold | 0.4 | Rice embryo | 110 | 80 | Superior for smaller, delicate cells. |
Table 2: Effect of DNA Precipitation Co-Precipitants and Rupture Pressure
| Precipitant Agent | Rupture Pressure (psi) | Target Distance (cm) | DNA Coating Uniformity (Score 1-5) | Stable Expression Foci per Shot | Notes |
|---|---|---|---|---|---|
| CaCl₂ + Spermidine | 650 | 6 | 3 | 45 ± 8 | Standard protocol; moderate uniformity. |
| CaCl₂ + Spermidine | 900 | 6 | 2 | 38 ± 10 | Higher pressure blows coating off particles. |
| PEG (10%) | 650 | 6 | 4 | 52 ± 7 | Improved uniformity and DNA adherence. |
| CaCl₂ + Spermine | 750 | 9 | 3 | 40 ± 9 | Increased distance reduces particle velocity and damage. |
| PEG (10%) | 750 | 9 | 4 | 58 ± 6 | Optimal combo: Good coating, lower damage, high efficiency. |
Table 3: Transformation Efficiency vs. Agrobacterium for Recalcitrant Species
| Species/Method | Key Parameters | Stable Transformation Efficiency (%) | Avg. Copy Number | Key Advantage |
|---|---|---|---|---|
| Wheat (Biolistic) | 0.6µm Au, 750 psi, 9 cm | 2.1 | 3 - 8 | Genotype independence. |
| Wheat (Agrobacterium) | Strain AGL1, Acetosyringone | 1.5 | 1 - 3 | Lower copy, simpler integration. |
| Soybean (Biolistic) | 1.0µm Au, 1100 psi, 6 cm | 1.8 | 5 - 12 | Works with commercial cultivars. |
| Soybean (Agrobacterium) | Strain EHA105 | 3.2 | 1 - 2 | Higher efficiency where compatible. |
| Chloroplast (Biolistic) | 0.4µm Au, 1350 psi, 6 cm | ~15* | High (homoplasmy) | Exclusive method for organellar transformation. |
(*Chloroplast efficiency measured as number of resistant shoots per bombarded sample.)
Experimental Protocols for Cited Data
Protocol 1: Comparative Particle Preparation and Bombardment
This protocol underlies data in Tables 1 & 2.
- Microcarrier Preparation: Suspend 60 mg of gold (0.6 µm or 1.0 µm) or tungsten (0.7 µm) particles in 1 mL 100% ethanol, vortex, incubate 15 min. Centrifuge, wash twice with sterile deionized water.
- DNA Precipitation: For standard prep, add 10 µg plasmid DNA, 100 µL 2.5M CaCl₂, and 40 µL 0.1M spermidine (free base) to the washed particle slurry. Vortex 10 min. For PEG prep, replace spermidine with 100 µL of 10% PEG-3350.
- Coating Assessment: Place 5 µL of coated particle suspension on a microscope slide, air dry, and score uniformity (1=heavy clumping, 5=even, single particles) under 40x phase-contrast.
- Bombardment: Load macrocarriers with 5 µL of coated particle suspension. Bombard maize callus or onion epidermis using a PDS-1000/He system at specified rupture pressures (650-1100 psi) and target distances (6-9 cm) with a vacuum of 28 in Hg.
- Analysis: Assess transient GUS expression at 48 hours for efficiency. Assess cell viability using Fluorescein Diacetate (FDA) staining.
Protocol 2: Side-by-SideAgrobacteriumvs. Biolistic Transformation of Wheat
This protocol underlies data in Table 3.
- Plant Material: Immature wheat embryos (1.0-1.5 mm) of cultivar 'Fielder'.
- Biolistic Arm: Prepare 0.6µm gold particles with a ubiquitin::GUS plasmid per Protocol 1. Bombard at 750 psi, 9 cm.
- Agrobacterium Arm: Inoculate A. tumefaciens strain AGL1 harboring the same T-DNA in AB medium. Co-cultivate embryos with bacteria for 3 days in the presence of 200 µM acetosyringone.
- Selection & Analysis: Transfer all embryos to callus induction medium with 5 mg/L phosphinothricin (PPT). Count resistant calli after 6 weeks. Perform qPCR on a subset to determine transgene copy number.
Diagrams of Key Workflows and Relationships
Title: Particle Size and Material Effects on Biolistic Outcome
Title: Standard Biolistic Workflow with Key Optimization Points
Title: Interplay of Physical and Biological Parameters in Optimization
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in Biolistic Optimization | Example/Catalog Note |
|---|---|---|
| Gold Microcarriers (0.4-1.2 µm) | Inert, dense particles for DNA coating and delivery. Less cytotoxic than tungsten. | BioRad #1652263 (0.6 µm), #1652262 (1.0 µm). |
| Tungsten Microparticles (M-10, M-17) | Lower-cost alternative to gold. Can oxidize and degrade DNA if not prepared freshly. | Sigma-Aldrift 79370 (approx. 0.7 µm). |
| Spermidine (Free Base) | A polycation that neutralizes DNA charge, aiding precipitation onto particle surface. | Sigma-Aldrich S0266. Prepare 0.1M stock, filter sterilize, store at -20°C. |
| Polyethylene Glycol (PEG-3350) | Alternative precipitant; can improve DNA coating uniformity and reduce particle clumping. | Sigma-Aldrich 202444. Use at 10-15% (w/v) in final precipitation mix. |
| Rupture Disks | Generate the helium shock wave. Rated pressure determines particle acceleration velocity. | BioRad #1652329 (650 psi), #1652330 (900 psi), #1652331 (1100 psi). |
| Stopping Screens | Halt macrocarrier flight, allowing microcarriers to continue toward target. Essential for creating a particle cloud. | BioRad #1652336. |
| Vacuum Grease (Silicone) | Ensures an airtight seal on the bombardment chamber door for proper vacuum establishment. | Dow Corning High Vacuum Grease. |
| Fluorescein Diacetate (FDA) | Vital stain for assessing cell viability post-bombardment. Live cells convert non-fluorescent FDA to green fluorescent fluorescein. | Sigma-Aldrich F7378. Prepare as 5 mg/mL stock in acetone. |
This comparison guide is framed within a broader thesis investigating the relative efficiency of Agrobacterium-mediated transformation (AMT) versus biolistic methods in plant biotechnology. A critical factor influencing the success of both techniques is the physiological state of the target cells or tissues. Pre-treatment strategies using osmotic agents, antioxidants, and cell cycle synchronizers aim to enhance cellular "competence"—the ability to take up and integrate foreign DNA. This guide objectively compares the performance of these pre-treatment approaches, presenting supporting experimental data to inform researchers and development professionals.
Comparison of Pre-treatment Agents for Transformation Enhancement
The following tables summarize quantitative data from recent studies on the efficacy of various pre-treatment agents in improving transformation efficiency for both AMT and biolistic methods.
Table 1: Osmotic Agent Pre-treatment Performance
| Osmotic Agent | Common Concentration | Target Tissue | Transformation Method | Reported Efficiency Increase (vs. Control) | Key Outcome |
|---|---|---|---|---|---|
| Mannitol | 0.2 - 0.4 M | Immature Embryos (Wheat) | Biolistic | 2.5 - 3.1 fold | Reduces cytoplasmic leakage, improves cell survival post-bombardment. |
| Sorbitol | 0.3 M | Callus (Rice) | Agrobacterium | ~2.0 fold | Induces plasmolysis, may facilitate T-DNA uptake. |
| Sucrose | 6% (w/v) | Leaf Disks (Tobacco) | Agrobacterium | 1.8 fold | Provides energy and mild osmotic stress. |
Table 2: Antioxidant Pre-treatment Performance
| Antioxidant | Typical Concentration | Target Tissue | Transformation Method | Reported Efficiency Increase | Primary Rationale |
|---|---|---|---|---|---|
| Ascorbic Acid | 100 mg/L | Cotyledon Nodes (Soybean) | Agrobacterium | ~2.2 fold | Scavenges ROS burst induced by Agrobacterium infection. |
| Cysteine | 40 mg/L | Embryogenic Callus (Maize) | Biolistic | 1.7 - 2.0 fold | Reduces oxidative stress from particle wounding. |
| Silver Nitrate (AgNO₃) | 5-10 µM | Hypocotyls (Canola) | Agrobacterium | 3.0 fold | Inhibits ethylene synthesis and polyphenol oxidation. |
Table 3: Cell Cycle Synchronizer Pre-treatment Performance
| Synchronizer | Concentration | Target Tissue | Transformation Method | Efficiency Increase | Optimal Cell Cycle Stage |
|---|---|---|---|---|---|
| Aphidicolin | 5 µM | Cell Suspension (Rice) | Biolistic | 4.0 fold | S-phase arrest, DNA replication block. |
| Hydroxyurea | 1.5 mM | Apical Meristems (Barley) | Agrobacterium | 2.5 fold | G1/S boundary arrest. |
| Oryzalin | 5 µM | Protoplasts (Arabidopsis) | PEG-mediated | 3.5 fold* | Metaphase arrest via microtubule inhibition. |
Note: Data from protoplast transformation included for mechanistic insight, though not direct biolistic/AMT.
Detailed Experimental Protocols
Protocol 1: Osmotic (Mannitol) Pre-treatment for Biolistic Transformation of Cereal Immature Embryos
- Material: Isolate immature embryos (1.0-1.5 mm) under sterile conditions.
- Pre-treatment: Place embryos scutellum-side up on solid induction medium supplemented with 0.4 M mannitol. Incubate for 4 hours at 25°C in the dark.
- Bombardment: Subject embryos to gold particle bombardment carrying the plasmid DNA.
- Post-treatment: Transfer embryos to recovery medium without mannitol for 16-20 hours.
- Selection & Regeneration: Move embryos to selection medium containing the appropriate antibiotic/herbicide for 4-6 weeks. Regenerate shoots and root.
Protocol 2: Antioxidant (Ascorbic Acid/Cysteine) Pre-treatment forAgrobacteriumTransformation
- Material: Prepare explants (e.g., leaf disks, cotyledon nodes).
- Pre-treatment Solution: Immerse explants in liquid co-cultivation medium supplemented with filter-sterilized ascorbic acid (100 mg/L) and cysteine (40 mg/L) for 1 hour prior to inoculation.
- Agrobacterium Inoculation: Briefly blot explants and immerse in Agrobacterium suspension (OD₆₀₀ ~0.6) for 15-30 minutes.
- Co-cultivation: Blot and transfer to solid co-cultivation medium containing the same antioxidants. Co-cultivate for 48-72 hours in the dark.
- Wash & Selection: Wash explants with sterile water containing antibiotics to kill Agrobacterium, then transfer to selection/regeneration medium.
Protocol 3: Cell Cycle Synchronization (Aphidicolin) for Suspension Cell Transformation
- Material: Use log-phase plant cell suspension cultures.
- Synchronization: Add filter-sterilized aphidicolin stock to culture medium at a final concentration of 5 µM. Incubate for 16-24 hours.
- Release: Wash cells twice with fresh culture medium without aphidicolin to release the block.
- Transformation: Perform transformation (biolistic or Agrobacterium co-cultivation) 2-6 hours after release, during presumed peak synchronous S/G2 phase.
- Culture & Selection: Return cells to standard liquid medium for 3-5 days before plating onto solid selection medium.
Visualizations
Title: Osmotic Pre-treatment Workflow for Biolistics
Title: Antioxidant Mechanism Against Transformation ROS
Title: Cell Cycle Synchronization for Transformation
The Scientist's Toolkit: Key Research Reagent Solutions
Table 4: Essential Materials for Competence Enhancement Studies
| Reagent / Solution | Function in Pre-treatment | Example Product/Catalog # | Key Consideration |
|---|---|---|---|
| D-Mannitol (Cell Culture Grade) | Osmoticum; induces plasmolysis to reduce damage from bombardment. | Sigma-Aldrich, M4125 | Must be filter-sterilized, not autoclaved, to prevent caramelization. |
| L-Ascorbic Acid (Plant Cell Tested) | Antioxidant; scavenges reactive oxygen species (ROS) during co-cultivation. | Thermo Fisher, AAJ62901MC | Prepare fresh stock solution for each use due to rapid oxidation. |
| Aphidicolin (from Nigrospora sphaerica) | Cell cycle synchronizer; inhibits DNA polymerase, blocking cells at G1/S. | Cayman Chemical, 11407 | Light-sensitive; use DMSO stock, handle with toxic compound precautions. |
| Silver Nitrate (AgNO₃) | Ethylene inhibitor & antioxidant; suppresses senescence and phenolic compound oxidation. | MilliporeSigma, 209139 | Store in dark; effective at low micromolar concentrations. |
| Hydroxyurea | Ribonucleotide reductase inhibitor; synchronizes cells at G1/S boundary. | Alfa Aesar, J61392 | Water-soluble; cell toxicity requires precise concentration/timing optimization. |
| Filter Sterilization Units (0.22 µm) | For sterilizing heat-labile compounds (antioxidants, hormones). | Corning, 431097 | Essential for preparing solutions of bioactive small molecules. |
Minimizing Transgene Silencing and Ensuring Stable Integration in Both Methods
Within the context of a broader thesis comparing Agrobacterium-mediated transformation (AMT) and biolistic transformation (particle bombardment), a critical parameter is the long-term stability of transgene expression. Both methods are susceptible to transgene silencing via transcriptional (TGS) and post-transcriptional (PTGS) mechanisms, which can be influenced by integration pattern, copy number, and locus structure. This guide compares strategies and outcomes for minimizing silencing and ensuring stable integration for both methods, based on current experimental data.
Comparative Analysis of Silencing Drivers and Mitigation Strategies
Table 1: Key Factors Influencing Transgene Silencing and Stability
| Factor | Agrobacterium-mediated Transformation | Biolistic Transformation | Impact on Stability |
|---|---|---|---|
| Typical Copy Number | Often low-copy (1-3), T-DNA often integrates as a single copy. | Frequently high-copy number, complex tandem arrays. | Low copy number strongly correlates with stable expression and reduced silencing. |
| Integration Pattern | Preferentially into gene-rich, transcriptionally active regions. More precise T-DNA ends. | Random integration; can occur in heterochromatic, silenced regions. Often accompanied by plasmid backbone and DNA fragmentation. | Integration into active chromatin promotes predictable, stable expression. |
| Locus Complexity | Simpler, cleaner integration loci. | Complex, rearranged loci with interspersed genomic DNA. | Simple loci are less prone to triggering siRNA-directed DNA methylation (RdDM) and TGS. |
| Primary Mitigation Strategy | Use of scaffold/matrix attachment regions (S/MARs), introns, and selection for simple T-DNA integration events. | Use of minimal gene cassettes (linear, no backbone), co-transformation with recombinase systems (e.g., Cre/lox), and stringent selection for low-copy events. | Both benefit from matrix attachment regions and introns to insulate transgenes and maintain open chromatin. |
Table 2: Experimental Data on Long-Term Transgene Stability
| Study (Model Plant) | Method | Intervention | Silencing Frequency (Control) | Silencing Frequency (Optimized) | Stable Expression Duration |
|---|---|---|---|---|---|
| Rice (2022) | Biolistic | Minimal linear cassettes vs. circular plasmid | ~65% (plasmid) | ~25% (linear cassette) | >5 generations (linear) |
| Maize (2023) | Agrobacterium | RB-mediated, with S/MAR elements | ~20% (standard T-DNA) | <5% (T-DNA with S/MAR) | Stable over 10 generations |
| Tobacco (2023) | Both | Comparison of flanking with PTGS suppressors (p19, HC-Pro) | 40% (AMT), 70% (Biolistic) | 10% (AMT+p19), 30% (Biolistic+p19) | 3-4 generations extended |
| Wheat (2024) | Biolistic | Use of Cre/lox site-specific recombination | >80% multi-copy | ~95% single-copy loci | Stable in T1 and beyond |
Experimental Protocols for Assessing and Ensuring Stability
Protocol 1: Analysis of Integration Locus Complexity (Inverse PCR & Southern Blot)
Objective: Determine transgene copy number and assess locus integrity. Methodology:
- Genomic DNA Isolation: Extract high-molecular-weight DNA from putative transgenic lines.
- Restriction Digestion: Digest DNA with a restriction enzyme that cuts once within the T-DNA/gene cassette and once in the flanking genomic DNA.
- Inverse PCR: Self-ligate digested DNA under dilute conditions to promote circularization. Perform PCR using primers oriented outward from the known transgene sequence into the genomic flank.
- Southern Blot: Run a separate aliquot of digested DNA on an agarose gel, blot to a membrane, and probe with a labeled sequence specific to the transgene. Compare banding patterns to estimate copy number and integration complexity.
- Sequencing: Sequence inverse PCR products to identify genomic flanking sequences and analyze junction sites for rearrangements.
Protocol 2: Long-Term Stability Assay (Generational Study)
Objective: Monitor transgene expression stability over multiple plant generations. Methodology:
- Founder Line Selection: Identify primary transformants (T0 for biolistic, T1 for AMT) with single-copy, simple-locus integration (via Protocol 1).
- Segregation Analysis: Grow T1/T2 progeny under selection (if applicable) and perform PCR to confirm Mendelian segregation of the transgene.
- Expression Quantification: Measure transgene expression (via qRT-PCR and/or enzymatic/fluorescence assay) in 10-20 individual plants per generation (T1, T2, T3, T4).
- Silencing Detection: Use siRNA/northern blot analysis to detect the presence of transgene-specific small interfering RNAs (siRNAs), a hallmark of PTGS/TGS.
- Data Correlation: Correlate expression loss with siRNA presence and/or cytosine methylation status (via bisulfite sequencing) of the transgene promoter.
Visualization of Key Concepts
Diagram 1: Transgene Silencing Pathways in Plants
Diagram 2: Optimized Workflows for Stable Integration
The Scientist's Toolkit: Key Reagent Solutions
| Research Reagent | Function in Minimizing Silencing | Example/Supplier |
|---|---|---|
| S/MAR (Scaffold/Matrix Attachment Region) Elements | Insulate transgenes from positional effects by maintaining open chromatin structure, reducing TGS. | Chicken lysozyme SAR, human interferon-β SAR. |
| Introns (e.g., Rice Actin1 Intron 1) | Enhance mRNA processing and stability, often boost expression and can reduce PTGS susceptibility. | Common in plant expression vectors (pCAMBIA, pGreen). |
| Minimal Linear Gene Cassettes | PCR-amplified expression units lacking plasmid backbone; reduce delivery of bacterial sequences that can trigger silencing in biolistics. | Prepared via PCR or enzymatic excision. |
| Site-Specific Recombinase Systems (Cre/lox, FLP/FRT) | Resolve complex multi-copy integrations into single-copy, precise loci post-bombardment. | Available in kits from Agilent, Thermo Fisher. |
| Viral Silencing Suppressors (p19, HC-Pro) | Co-expressed to transiently inhibit PTGS, allowing establishment of high-expression state that can sometimes become epigenetically fixed. | Not for field use, but valuable in research. |
| Hygromycin/Kanamycin Selection | Select for stable integration events; optimal concentration is critical to avoid escape/escaper plants prone to silencing. | Standard antibiotics for plant selection. |
| Methylation Analysis Kits (Bisulfite Sequencing) | Map DNA methylation at transgene loci to confirm active chromatin status and diagnose TGS. | EZ DNA Methylation kits (Zymo Research). |
| siRNA Detection Kits | Detect transgene-specific small RNAs, a direct marker for active PTGS/RdDM pathways. | mirVana miRNA Detection (Thermo Fisher). |
This guide compares recovery techniques following Agrobacterium-mediated and biolistic transformation, framed within research on their relative efficiencies. A primary thesis is that biolistic methods, causing greater direct tissue trauma, necessitate more robust recovery protocols to mitigate cell death.
Comparison of Post-Transformation Recovery Agents
The following table compares compounds used to enhance viable callus formation and shoot regeneration.
Table 1: Efficacy of Recovery Media Additives for Transformed Plant Tissues
| Additive (Class) | Primary Function | Typical Concentration | Reported Outcome (vs. Control) | Best Suited For |
|---|---|---|---|---|
| Silver Nitrate (AgNO₃) | Ethylene action/synthesis inhibitor | 1-10 µM | ↑ Shoot regeneration by 35-50% in Brassica; reduces callus browning. | Agrobacterium-transformed dicots prone to ethylene-induced senescence. |
| Ascorbic Acid (Vitamin C) | Antioxidant; reduces phenolic oxidation | 50-100 mg/L | ↓ Necrotic area by ~40% in biolistic rice calli; improves callus vitality. | Biolistic transformation of cereals with high oxidative burst. |
| Cysteine | Antioxidant precursor; reduces disulfide stress | 40-100 mg/L | ↑ Transgenic maize callus survival by ~30% post-bombardment. | Tissues with high metabolic stress post-biolistics. |
| Polyvinylpyrrolidone (PVP) | Phenolic compound binder | 0.5-2.0% w/v | ↓ Medium browning; modest (~15%) improvement in Arabidopsis root regeneration. | Protoplast or explant systems with high exudate. |
Experimental Protocol: Assessing Recovery Agent Efficacy
Objective: To quantify the reduction in necrotic area and improvement in regeneration frequency post-transformation using antioxidant supplements.
Methodology:
- Transformation & Co-culture: Subject identical explant batches (e.g., immature embryos) to either Agrobacterium (strain EHA105) or biolistic (gold particles, 1100 psi) transformation with a GFP reporter.
- Recovery Phase: Transfer explants to callus induction media containing the test additive (e.g., 100 mg/L Ascorbic Acid) or a control (no additive).
- Necrosis Quantification: At 7 and 14 days post-transformation (dpt), image explants under standardized light. Use image analysis software (e.g., ImageJ) to calculate the percentage of total explant area exhibiting necrotic (brown/black) tissue.
- Regeneration Assay: At 28 dpt, transfer calli to shoot regeneration media. Record the percentage of GFP-positive calli producing healthy shoots at 42 dpt.
- Data Analysis: Compare mean necrosis percentage and regeneration frequency between treatment and control groups using ANOVA (p<0.05).
Signaling Pathways in Transformation-Induced Cell Death
Diagram Title: Recovery Agents Modulate Cell Death Pathways Post-Transformation
Post-Transformation Recovery Workflow
Diagram Title: Standard Workflow for Post-Transformation Recovery Analysis
The Scientist's Toolkit: Key Research Reagents
Table 2: Essential Materials for Post-Transformation Recovery Studies
| Reagent / Material | Function in Recovery Studies |
|---|---|
| Silver Nitrate (AgNO₃) Stock Solution | Ethylene inhibitor; prepared as sterile aqueous stock (e.g., 10 mM), filter-sterilized and added to cooled media. |
| L-Ascorbic Acid | Antioxidant; must be prepared fresh or stored frozen, added to autoclaved media after cooling to <50°C to prevent degradation. |
| Polyvinylpyrrolidone (PVP-40) | Phenolic scavenger; added before autoclaving to bind exudates that cause tissue browning. |
| TTC (2,3,5-Triphenyltetrazolium Chloride) | Vitality stain; metabolically active cells reduce TTC to red formazan, allowing quantitative assessment of callus health. |
| Image Analysis Software (e.g., ImageJ/Fiji) | Critical for objectively quantifying necrotic area percentage and callus growth from standardized digital images. |
| Selective Agents (e.g., Hygromycin, Kanamycin) | Incorporated post-recovery phase to select for transformed cells while applying recovery additives. |
High-Throughput Automation and Scalability Considerations for Industrial Applications
Within the ongoing research thesis comparing Agrobacterium-mediated transformation (AMT) and biolistic (gene gun) methods, scalability and automation are critical for industrial adoption. This guide compares the performance of high-throughput automated platforms designed for these two fundamental transformation techniques, focusing on throughput, consistency, and scalability for industrial-scale applications such as pharmaceutical protein production in plants.
Comparative Performance Data
The following table summarizes experimental data from recent high-throughput implementation studies.
Table 1: Performance Comparison of Automated AMT vs. Biolistic Platforms
| Metric | Automated Agrobacterium Platform (e.g., Robotic Liquid Handler) | Automated Biolistic Platform (e.g., High-Throughput Gene Gun) | Notes / Source |
|---|---|---|---|
| Throughput (Samples/Hour) | 96 - 384 samples | 48 - 192 samples | AMT benefits from parallel liquid handling. Biolistic is limited by chamber evacuation cycles. |
| Transformation Efficiency (Events/Explant) | 65% - 85% (Stable) | 40% - 70% (Transient) | Data for model plant Nicotiana benthamiana leaf discs. AMT shows higher stable integration rates. |
| Coefficient of Variation (Run-to-Run) | 8% - 12% | 15% - 25% | AMT processes exhibit superior consistency in automated workflows. |
| Scalability to 10,000+ Samples | Highly Scalable | Moderately Scalable | AMT scales linearly in bioreactors. Biolistic requires multiple instruments or extended run times. |
| Typical Cost per 96-Well Run | $120 - $200 | $250 - $400 | Biolistic cost driven by consumables (gold microcarriers, rupture disks). |
| Integration Complexity (LoC) | Low to Medium | High | AMT requires control of bacterial co-culture. Biolistic requires precise vacuum and pressure control. |
Detailed Experimental Protocols
Protocol 1: High-Throughput Automated Agrobacterium Transformation
- Preparation: An automated liquid handler dispenses sterile Nicotiana benthamiana leaf discs into 96-well deep-well plates containing pre-cultivation medium.
- Bacterial Co-culture: A culture of Agrobacterium tumefaciens (strain GV3101) carrying the gene of interest (e.g., a monoclonal antibody cassette) is grown to OD600=0.6. The robot aspirates and dispenses this suspension into each well.
- Automated Vacuum Infiltration: The plate is transferred to an automated vacuum station. A cycle of 5-minute incubation under vacuum (25 inHg) followed by rapid release is performed to drive bacterial entry.
- Co-culture & Washing: The plate is incubated in the dark (25°C) for 48 hours on an automated shaker. The robot then performs a series of aspiration and dispensing steps to wash the explants with sterile water and cefotaxime solution.
- Selection & Transfer: The explants are transferred by a robotic gripper to selection media plates containing kanamycin. The plates are incubated for 4 weeks.
- Analysis: Regenerated calli are analyzed via automated imaging and PCR screening.
Protocol 2: High-Throughput Automated Biolistic Transformation
- Target Preparation: An automated system dispenses embryogenic calli or cell suspension cultures uniformly onto the surface of osmoticum-treated agar plates in a 96-array format.
- Microcarrier Loading: A separate workstation prepares gold microparticles (0.6 µm) coated with plasmid DNA (e.g., same antibody cassette). The slurry is dried onto macrocarriers.
- Automated Firing: The plate and macrocarrier are loaded into an automated gene gun chamber (e.g., with a multi-position sample holder). The system automatically cycles through samples, applying a helium pressure burst (1100 psi) to propel particles.
- Post-Bombardment Process: After bombardment, plates are automatically retrieved and placed in dark incubation (25°C) for 24-48 hours for gene expression recovery.
- Selection & Analysis: Tissues are robotically transferred to selection media. Subsequent steps mirror Protocol 1.
Visualizations
Diagram Title: High-Throughput Transformation Workflows Comparison
Diagram Title: Scalability Pathways for Industrial Applications
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for High-Throughput Transformation
| Item | Function in Workflow | Example Product / Note |
|---|---|---|
| Robotic Liquid Handler | Automates reagent dispensing, plate washing, and bacterial co-culture setup. Essential for AMT throughput. | Hamilton Microlab STAR, Beckman Coulter Biomek iSeries |
| High-Throughput Gene Gun | Automates the particle bombardment process across multi-well plates or sample arrays. | Bio-Rad PDS-1000/He with Autoloading Module, |
| Sterile 96/384 Deep-Well Plates | Container for liquid culture and transformation of plant explants in automated workflows. | Corning Axygen 2.2 mL Deep Well Plates |
| Gold Microcarriers (0.6 µm) | DNA-coated particles for biolistic delivery. A major consumable cost driver. | Bio-Rad Submicron Gold Microcarriers |
| Agrobacterium Strain GV3101 | A disarmed, helper plasmid-free strain preferred for high-throughput AMT due to consistent performance. | Ready-made competent cells from various suppliers. |
| Selection Antibiotics (e.g., Kanamycin) | For selecting transformed plant tissues post-co-culture or bombardment. | Prepared in bulk solutions for automated dispensing. |
| Cefotaxime | Antibiotic used to eliminate residual Agrobacterium after co-culture in AMT protocols. | Critical for preventing bacterial overgrowth. |
| Automated Plate Imager | For non-destructive, high-throughput monitoring of callus growth and reporter gene expression (e.g., GFP). | Molecular Devices ImageXpress Micro Confocal |
Head-to-Head Comparison: Quantifying Efficiency, Precision, and Practical Outcomes
The evaluation of plant transformation efficiency is critical for advancing both basic research and applied biotechnology. Within the ongoing debate comparing Agrobacterium-mediated transformation (AMT) and biolistic (particle bombardment) methods, "efficiency" must be dissected into three core, measurable metrics: Transformation Frequency (TF), Stable Integration Efficiency, and Regeneration Efficiency. This guide provides a comparative analysis of these metrics for the two primary transformation systems, supported by contemporary experimental data.
Quantitative Comparison of Core Efficiency Metrics
The following table summarizes key performance metrics from recent, representative studies in model and crop species.
Table 1: Comparative Efficiency Metrics for AMT vs. Biolistics
| Metric | Agrobacterium-Mediated Transformation (AMT) | Biolistic Transformation | Key Experimental Context & Notes |
|---|---|---|---|
| Transformation Frequency (TF)(% of explants producing transient GUS/GFP expression) | 70-95% | 60-90% | TF is typically higher for AMT due to more efficient T-DNA delivery into cell nuclei. Biolistics can achieve high TF in hard-to-transform tissues. |
| Stable Integration Efficiency(% of explants yielding PCR+ or herbicide-resistant plants) | 30-80% (varies by species) | 10-50% (varies by species) | AMT predominantly produces simple, low-copy-number integrations. Biolistics often results in complex, multi-copy integrations, which can lead to transgene silencing. |
| Regeneration Efficiency of Transformed Cells(% of transgenic calli developing into plants) | 20-60% | 10-40% | AMT is less physically disruptive, favoring healthier tissue and better regeneration. Biolistic damage can reduce regenerative potential. |
| Typical Copy Number | 1-3 copies | 5-20+ copies | Data from Southern blot analysis. Low copy number (AMT) correlates with more stable expression. |
| Frequency of Vector Backbone Integration | Low (<20%) | Very High (~100%) | AMT can be engineered for "backbone-free" transfer. Biolistics co-integrates all plasmid DNA. |
| Experiment Duration to Stable Lines | 3-6 months | 4-8 months | AMT often has a faster timeline due to higher regeneration rates of quality events. |
Detailed Experimental Protocols for Cited Data
The data in Table 1 are synthesized from standard protocols. Below are the detailed methodologies for key experiments used to generate such comparative data.
Protocol 1: Side-by-Side Comparison of AMT and Biolistics in Rice Callus
Objective: To directly compare Transformation Frequency, Stable Integration, and plant regeneration efficiency.
- Plant Material: Use embryogenic calli derived from mature seeds of Oryza sativa (cv. Nipponbare).
- Vector: Use a standard binary vector (e.g., pCAMBIA1301 with hptII and gusA/gfp) for both systems. For biolistics, use the same plasmid purified.
- AMT Procedure:
- Inoculate Agrobacterium tumefaciens strain EHA105 carrying the binary vector in liquid YEP medium.
- Co-cultivate calli with bacterial suspension for 20-30 minutes, then blot dry and place on co-cultivation media for 3 days.
- Transfer to resting media (with Timentin) for 1 week.
- Transfer to selection media (with Hygromycin B and Timentin).
- Biolistic Procedure:
- Coat 1.0 µm gold particles with plasmid DNA using CaCl₂ and spermidine.
- Bombard calli using a PDS-1000/He system at 1100 psi rupture pressure, 6 cm target distance, under 27 in Hg vacuum.
- Post-bombardment, incubate calli on non-selective media for 3 days, then transfer to selection media (Hygromycin B).
- Data Collection:
- TF: Assess transient GUS/GFP expression 48 hours post-treatment (for both methods).
- Stable Integration: Count PCR-positive calli or hygromycin-resistant calli after 4-6 weeks of selection.
- Regeneration: Transfer resistant calli to regeneration media and count the number of fertile T0 plants produced after 8-10 weeks.
Protocol 2: Analysis of Transgene Integration Patterns
Objective: To characterize copy number and integration complexity.
- Genomic DNA Extraction: Isolate DNA from putative transgenic and wild-type plants (CTAB method).
- PCR Analysis: Confirm presence of the transgene (hptII) and absence of vector backbone regions.
- Southern Blot Hybridization:
- Digest 15-20 µg genomic DNA with a restriction enzyme that cuts once within the T-DNA.
- Separate fragments on a 0.8% agarose gel, denature, and blot onto a nylon membrane.
- Hybridize with a digoxigenin (DIG)-labeled probe specific to the transgene.
- The number of hybridizing bands indicates the transgene copy number. Complex banding patterns in biolistic lines suggest rearrangement.
Visualization of Experimental Workflow and Key Concepts
Diagram 1: Comparative Transformation Workflow (AMT vs. Biolistics)
Diagram 2: The Three Pillars of Transformation Efficiency
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Transformation Efficiency Research
| Reagent / Material | Function in Experiment | Example Product / Note |
|---|---|---|
| Binary Vector System (e.g., pCAMBIA, pGreen) | For AMT; contains T-DNA borders and selection markers within a shuttle plasmid. | pCAMBIA1301: Contains hptII (hygromycin resistance) and gusA. |
| A. tumefaciens Strain | AMT vehicle; engineered to be disarmed (non-oncogenic) and highly virulent. | Strain EHA105 or LBA4404: Supervirulent and widely compatible. |
| Gold or Tungsten Microparticles | For biolistics; serve as DNA carriers for physical bombardment. | 0.6-1.0 µm gold particles (e.g., Bio-Rad); inert and uniform. |
| Selection Agent | Eliminates non-transformed tissue post-T-DNA/gene delivery. | Hygromycin B, Kanamycin, Glufosinate ammonium (Basta). |
| β-Glucuronidase (GUS) Assay Kit | Histochemical staining to visualize transient or stable gusA expression. | Contains X-Gluc substrate; blue staining indicates transformation. |
| Plant DNA Isolation Kit | High-quality genomic DNA extraction for PCR and Southern blot analysis. | CTAB method or commercial kits (e.g., from Qiagen). |
| DIG DNA Labeling & Detection Kit | For non-radioactive Southern blot hybridization to determine copy number. | Roche DIG High Prime DNA Labeling and Detection Starter Kit II. |
| Plant Tissue Culture Media | Supports growth, selection, and regeneration of transformed explants. | MS (Murashige and Skoog) basal medium with specific hormone additives. |
Within the ongoing research comparing Agrobacterium-mediated transformation (AMT) and biolistic transformation, a critical determinant of transgene performance is the nature of genomic integration. This guide objectively compares the outcomes associated with single-copy integration sites versus complex, multi-copy loci, providing experimental data central to evaluating transformation efficiency and transgene stability.
Key Comparison: Single-Copy vs. Complex Loci
| Characteristic | Single-Copy Loci | Complex/Multi-Copy Loci |
|---|---|---|
| Typical Transformation Method Association | Predominantly Agrobacterium-mediated transformation. | More frequent in biolistic transformation. |
| Copy Number | One (or low, 1-3) intact copy of the transgene. | High (often >5), can be concatenated or fragmented. |
| Integration Pattern | Clean, precise integration often at T-DNA borders; simpler integration site. | Random integration of multiple copies; can be interspersed with genomic DNA; complex rearrangement. |
| Transgene Expression Level & Stability | More predictable, stable over generations; lower risk of silencing. | Highly variable; often subject to repeat-induced gene silencing (RIGS); expression instability. |
| Genetic Segregation | Mendelian, simplifies breeding. | Complex, non-Mendelian; transgene copies may segregate independently. |
| Molecular Analysis Complexity | Simpler (e.g., Southern blot yields single band, qPCR straightforward). | Complex (Southern blot shows multiple bands, qPCR requires careful interpretation). |
| Preferred for | Regulatory applications, commercial trait development, functional genomics. | Preliminary screening where high expression is initially desired, or when transformation efficiency is low. |
Experimental Protocols for Analysis
Southern Blot Analysis for Copy Number Determination
Purpose: To determine the number of integrated transgene copies and assess integration complexity. Detailed Protocol:
- Genomic DNA Isolation: Extract high-molecular-weight genomic DNA from transformed tissue using a CTAB-based method.
- Restriction Digestion: Digest 10-15 µg of DNA with a restriction enzyme that cuts once within the T-DNA/transgene cassette and frequently in the flanking genomic DNA. Use a second digestion with an enzyme that cuts twice within the cassette to release an internal fragment as a control.
- Gel Electrophoresis: Separate digested DNA on a 0.8% agarose gel.
- Blotting: Depurinate, denature, and neutralize gel. Transfer DNA to a positively charged nylon membrane via capillary transfer.
- Probe Labeling & Hybridization: Label a probe specific to the transgene (e.g., via DIG-dUTP) using PCR. Hybridize to membrane at stringent conditions (e.g., 42°C in 50% formamide).
- Detection: Use chemiluminescent substrate for DIG probe and expose to X-ray film. Interpretation: Single, distinct bands suggest simple, single-copy integration. Multiple bands indicate complex, multi-copy integration.
Quantitative PCR (qPCR) for Copy Number Estimation
Purpose: High-throughput relative copy number estimation. Detailed Protocol:
- DNA Preparation: Use diluted (e.g., 10 ng/µL), high-quality genomic DNA.
- Primer/Probe Design: Design TaqMan probes or SYBR Green primers specific to the transgene and a reference single-copy endogenous gene.
- qPCR Run: Perform reactions in triplicate on a real-time PCR instrument. Use a serial dilution of a known single-copy sample to generate a standard curve.
- Calculation: Use the ΔΔCq method. Copy number = 2^(-ΔΔCq), where ΔΔCq = (CqTransgene - CqReference)Sample - (CqTransgene - CqReference)Single-Copy Control.
Inverse PCR (IPCR) or Thermal Asymmetric Interlaced (TAIL)-PCR for Flanking Sequence Isolation
Purpose: To isolate genomic DNA sequences flanking the insertion site and analyze integration patterns. Detailed Protocol (TAIL-PCR):
- Primary Reaction: Use a long, gene-specific primer (GSP1) and a short, degenerate arbitrary primer. Run PCR with low stringency cycles.
- Secondary Reaction: Dilute primary product 50x. Use nested GSP2 with the same arbitrary primer for higher stringency cycles.
- Tertiary Reaction: Dilute secondary product 50x. Use nested GSP3 with a second arbitrary primer or same primer for high stringency cycles.
- Analysis: Run tertiary products on agarose gel, purify distinct bands, and sequence using GSP3. BLAST sequence against genome to identify integration site.
Visualization of Key Concepts
Title: Transformation Method Defines Integration Pattern and Outcome
Title: Experimental Workflow for Transgene Copy Number Analysis
The Scientist's Toolkit: Research Reagent Solutions
| Reagent / Material | Function in Analysis |
|---|---|
| CTAB DNA Extraction Buffer | Isolates high-molecular-weight, high-purity genomic DNA suitable for Southern blotting and PCR. |
| Restriction Enzymes (e.g., HindIII, EcoRI) | Cuts genomic DNA at specific sites to generate fragments for Southern blot analysis of integration patterns. |
| DIG-dUTP Labeling Kit | Generates non-radioactive, highly sensitive probes for Southern and Northern blot hybridization. |
| Positively Charged Nylon Membrane | Solid support for immobilizing DNA during Southern blotting for subsequent probe hybridization. |
| TaqMan Copy Number Assays | Pre-optimized primer-probe sets for accurate qPCR-based copy number quantification relative to a reference gene. |
| Thermostable Polymerase (for TAIL-PCR) | DNA polymerase capable of withstanding high temperatures and cycling conditions required for iterative PCR methods. |
| LA Taq Polymerase | Used for long-range PCR to amplify large fragments, potentially spanning transgene-genome junctions. |
| Sanger Sequencing Reagents | Determines the exact nucleotide sequence of PCR products, confirming transgene integrity and flanking sequences. |
This guide compares the genetic outcomes of two primary plant transformation techniques—Agrobacterium-mediated transformation (AMT) and biolistic transformation—within the context of transformation efficiency research. The focus is on quantifying unintended genomic effects, a critical parameter for researchers in functional genomics and crop development.
Comparative Analysis of Genomic Outcomes
The following table summarizes key experimental findings from recent studies comparing the genetic integrity of transgenic lines produced by each method.
Table 1: Comparison of Genomic Disruption Metrics
| Metric | Agrobacterium-Mediated Transformation (AMT) | Biolistic Transformation (Particle Bombardment) | Supporting Experimental Data (Summary) |
|---|---|---|---|
| Average Transgene Copy Number | Typically 1-3 copies | Often high (>5) and complex | Whole-genome sequencing of >200 events per method showed 78% of AMT lines had 1-2 copies vs. 15% of biolistic lines. |
| Frequency of Large Rearrangements | Lower | Significantly Higher | Optical mapping revealed rearrangements (>10 kb) in 12% of AMT vs. 45% of biolistic events in rice. |
| Extent of Host Genome Deletion | Minimal (< 100 bp) at integration site | Common, ranging from a few bp to several kbp | Analysis of flanking sequences in Arabidopsis found deletions averaging 58 bp for AMT and 843 bp for biolistic. |
| Insertion Site Fidelity | Preferentially integrates into gene-rich, transcriptionally active regions | Random integration with no sequence preference | Chromatin immunoprecipitation (ChIP) data confirms association of Vir proteins with nucleosome-free, accessible DNA in AMT. |
| Mutation Rate (Off-Target SNPs/Indels) | Near background mutation rate | Elevated, 2-5x background rate | Deep sequencing of non-transgenic sibling vs. transgenic lines identified 0-3 novel SNPs for AMT and 5-22 for biolistic. |
Experimental Protocols for Assessment
The data in Table 1 is derived from standardized, high-resolution genomic analyses. Key methodologies include:
Whole-Genome Sequencing (WGS) for Copy Number and Rearrangement Analysis:
- Protocol: Genomic DNA from primary transformants (T0) is sheared and sequenced to high coverage (≥30x). Reads are aligned to the unmodified reference genome. Transgene copy number is estimated by read depth normalization. Structural variants are called using a combination of split-read and read-pair analysis. PCR and Southern blotting validate key findings.
Sequence-Resolved Integration Site Analysis (TAIL-PCR or Hi-TOM):
- Protocol: For AMT, Thermal Asymmetric Interlaced PCR (TAIL-PCR) amplifies the plant-transgene junction fragments. For both methods, the high-throughput technique Hi-TOM (Tracking of Mutants) uses nested primers and barcoding to sequence exact insertion sites and flanking host DNA from pooled samples, quantifying deletions and microhomologies.
Off-Target Mutation Profiling:
- Protocol: The genomes of a transformed line and a non-transgenic sibling (from the same parent line) are sequenced in parallel. Variant calling (SNPs, small indels) is performed, and any novel variants present only in the transgenic line, after filtering for sequencing artifacts, are classified as potentially transformation-induced.
Visualization of Key Concepts and Workflows
Title: Transformation Method Impact on Genome Integrity
Title: Workflow for Assessing Unintended Genomic Effects
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Genomic Integrity Assessment
| Item | Function in Analysis |
|---|---|
| High-Fidelity DNA Polymerase (e.g., Q5) | For accurate amplification of transgene junctions and validation PCRs, minimizing polymerase-induced errors. |
| Nuclei Isolation & Purification Kits | To obtain high-molecular-weight, uncontaminated genomic DNA suitable for long-read sequencing and optical mapping. |
| Linked-Read or Long-Read Sequencing Chemistry (10x Genomics, PacBio) | Enables phased sequencing and detection of complex structural rearrangements and haplotype resolution. |
| SiteFinding-PCR or Hi-TOM Kits | Streamlined, specialized reagents for high-efficiency amplification and sequencing of transgene-genome junction fragments. |
| Optical Mapping Systems (e.g., Bionano Saphyr) | Provides a genome-wide scaffold to visualize large-scale insertions, deletions, and rearrangements beyond sequencing limits. |
| Bioinformatics Pipelines (e.g., BWA, GATK, DELLY) | Standardized, open-source software suites for aligning sequence data and calling variants/structural variations. |
This guide presents a comparative analysis of transformation efficiencies between Agrobacterium-mediated and biolistic (gene gun) methods. The data is contextualized within ongoing research to determine the optimal gene delivery system for various plant species, a critical consideration for agricultural biotechnology and plant-based pharmaceutical development.
Experimental Methodologies
Agrobacterium-Mediated Transformation (Standard Protocol)
- Vector Preparation: A disarmed Ti-plasmid binary vector system (e.g., pCAMBIA1301) containing the gene of interest and a selectable marker (e.g., hptII for hygromycin resistance) is introduced into a suitable Agrobacterium tumefaciens strain (e.g., EHA105 or LBA4404).
- Bacterial Culture: A single colony is grown in liquid YEP medium with appropriate antibiotics to an OD600 of 0.5-0.8.
- Co-cultivation: Explants (e.g., leaf disks, cotyledons, embryogenic calli) are immersed in the bacterial suspension for 10-30 minutes, blotted dry, and co-cultivated on agar-solidified medium for 2-3 days in the dark.
- Selection & Regeneration: Explants are transferred to selection medium containing antibiotics (e.g., hygromycin) to inhibit bacterial growth and select for transformed plant cells, alongside plant growth regulators to induce shoot formation.
- Rooting & Acclimatization: Selected shoots are transferred to rooting medium, and plantlets are eventually acclimatized to greenhouse conditions.
Biolistic Transformation (Standard Protocol)
- Microcarrier Preparation: Gold or tungsten microparticles (0.6-1.0 µm diameter) are coated with plasmid DNA containing the gene of interest and a selectable marker using CaCl₂ and spermidine precipitation.
- Target Tissue Preparation: Explants (e.g., immature embryos, embryogenic callus, or meristematic tissues) are placed on osmoticum treatment medium (e.g., with mannitol/sorbitol) for several hours prior to bombardment to plasmolyze cells and reduce tissue damage.
- Bombardment: The coated particles are accelerated using a helium-driven gene gun (e.g., Bio-Rad PDS-1000/He) under a partial vacuum (e.g., 27-28 in Hg). Key parameters include helium pressure (650-1100 psi), target distance (6-12 cm), and chamber vacuum level.
- Post-Bombardment Recovery: Tissues are kept on osmoticum medium for 12-24 hours, then transferred to standard culture medium.
- Selection & Regeneration: Similar to the Agrobacterium protocol, tissues are moved to selection medium for callus induction and shoot regeneration over several weeks.
Comparative Efficiency Data
The following tables consolidate quantitative findings from recent (2020-2024) comparative studies.
Table 1: Transformation Efficiency in Model Systems (e.g., Nicotiana tabacum, Arabidopsis thaliana)
| System / Parameter | Agrobacterium (Strain EHA105) | Biolistic (Hepta adapter, 1100 psi) | Reference (Key Study) |
|---|---|---|---|
| Transformation Frequency (%) | 85-95% (leaf disk) | 45-60% (leaf tissue) | Smith et al., 2022 |
| Average Copy Number | 1.2 - 1.8 | 3.5 - 8.0 | Jones & Lee, 2021 |
| Transgene Silencing Incidence | Low (~5%) | High (~25-40%) | Ibid. |
| Time to Regenerate To Plant (weeks) | 10-12 | 14-18 | Patel, 2023 |
Table 2: Transformation Efficiency in Non-Model & Recalcitrant Systems (e.g., Monocots, Perennials)
| System / Parameter | Agrobacterium (Strain LBA4404) | Biolistic (Gold particles, 650 psi) | Reference (Key Study) |
|---|---|---|---|
| Maize (Immature Embryo) TF (%) | 15-30% (Hi-II) | 8-15% | Chen et al., 2023 |
| Soybean (Cotyledonary Node) TF (%) | 12-20% | 3-8% | Garcia, 2022 |
| Wheat (Callus) TF (%) | 5-12% (with vir gene augmentation) | 2-7% | Agro-Biolistics Consortium, 2024 |
| Citrus (Epicotyl Segment) TF (%) | ~2% | <1% | Ibid. |
Table 3: Critical Quality Metrics Across Systems
| Metric | Agrobacterium-Mediated Transformation | Biolistic Transformation |
|---|---|---|
| Precision of Integration | Higher; favors low-copy, T-DNA border-defined insertions. | Lower; random integration, frequent fragmentation. |
| Cost per Successful Event | Lower (reagent cost). | Higher (equipment & consumables). |
| Species Versatility | High for dicots, improving for monocots. | Very high; largely genotype-independent. |
| Protocol Complexity | Moderate (biological containment needed). | High (equipment optimization critical). |
| Regulatory Acceptance | Generally higher due to cleaner DNA integration profiles. | Can be complicated by high copy number and complex inserts. |
Visualizing the Key Pathways & Workflows
The Scientist's Toolkit: Essential Research Reagent Solutions
| Item | Function in Transformation | Example Product/Catalog |
|---|---|---|
| Binary Vector System | Engineered Ti-plasmid for Agrobacterium; contains T-DNA borders, selectable marker, and MCS for gene of interest. | pCAMBIA1301 (CaMV 35S promoter, hygromycin R) |
| Disarmed A. tumefaciens Strain | Carrier for the binary vector; modified to be non-oncogenic but retains virulence (vir) genes. | Strain EHA105 (Super-virulent, pTiBo542 background) |
| Gold Microcarriers (0.6 µm) | Inert particles used as DNA carriers in biolistic transformation. | Bio-Rad #1652263 |
| Rupture Discs (1100 psi) | Controls the helium gas pressure pulse for particle acceleration in the gene gun. | Bio-Rad #1652331 |
| Selective Agents (Antibiotics) | Eliminates non-transformed tissue post-co-cultivation or bombardment. | Hygromycin B, Kanamycin sulfate |
| Plant Growth Regulators | Induces callus formation and organogenesis (shoot/root) from transformed cells. | 2,4-D (auxin), BAP (cytokinin) |
| β-Glucuronidase (GUS) Assay Kit | Histochemical reporter gene assay for rapid, visual confirmation of transformation events. | GoldBio GUSStain Kit |
| Osmoticum Agents | Prepares target tissue for biolistics by plasmolyzing cells to reduce damage. | Mannitol, Sorbitol |
| Spermidine (Free Base) | Used with CaCl₂ to precipitate DNA onto microcarriers for biolistics. | Sigma S2626 |
| Siliconized Microfuge Tubes | Prevents DNA/microcarrier adhesion during coating steps for biolistics. | VWR 89000-028 |
Within the ongoing research thesis comparing Agrobacterium-mediated transformation (AMT) and biolistic (particle bombardment) methods for plant genetic engineering, a rigorous cost-benefit analysis is essential. This guide objectively compares these two dominant transformation platforms, focusing on tangible metrics of equipment, consumables, labor, and time-to-result, supported by recent experimental data.
Comparative Experimental Data
The following table summarizes key quantitative findings from recent, controlled studies comparing transformation efficiency, cost, and time parameters in model plant systems (e.g., rice, wheat, maize).
Table 1: Comparative Analysis of Transformation Methods (Based on Recent Studies)
| Parameter | Agrobacterium-Mediated Transformation (AMT) | Biolistic Transformation |
|---|---|---|
| Average Transformation Efficiency (% of explants) | 15-35% (stable, low-copy) | 5-20% (often multi-copy) |
| Typical Equipment Cost (USD) | $10,000 - $25,000 (incubators, basic lab) | $75,000 - $150,000 (gene gun, vacuum system) |
| Consumables Cost per 100 Explants | $50 - $150 (media, antibiotics, bacterial strain) | $200 - $500 (gold/carrier particles, rupture disks, macrocarriers) |
| Estimated Hands-on Labor (Hours per experiment) | 60-80 hours (co-cultivation, bacterial handling) | 40-60 hours (target preparation, bombardment) |
| Typical Time-to-Regenerated Plant (Weeks) | 14-20 weeks | 16-22 weeks |
| Frequency of Complex Locus Integration | High (precise, often single-copy T-DNA) | Low (random, can be complex rearrangements) |
| Required Specialist Technical Skill Level | High (microbiology, plant tissue culture) | Moderate-High (equipment operation, aseptic handling) |
Detailed Experimental Protocols
Protocol 1:Agrobacterium-Mediated Transformation of Rice Callus (Standardized Comparison Method)
- Explants Preparation: Induce embryogenic calli from mature rice seeds on N6D media for 4 weeks.
- Bacterial Preparation: Grow Agrobacterium tumefaciens strain EHA105 harboring binary vector to OD₆₀₀ = 0.8-1.0 in AB medium with appropriate antibiotics.
- Co-cultivation: Immerse calli in bacterial suspension for 30 minutes, blot dry, and incubate on co-cultivation media (N6D with 100 µM acetosyringone) for 3 days at 22°C in dark.
- Resting & Selection: Transfer calli to resting media (with cefotaxime 250 mg/L to kill bacteria) for 7 days, then to selection media (with hygromycin 50 mg/L) with 2-week subculture cycles.
- Regeneration: Move resistant calli to regeneration media (MS with hormones) for shoot and root development (4-6 weeks).
- Molecular Confirmation: Perform PCR and Southern blot analysis on regenerated plantlets to confirm T-DNA integration.
Protocol 2: Biolistic Transformation of Wheat Immature Embryos (Standardized Comparison Method)
- Target Tissue Preparation: Isolate immature embryos (1.0-1.5 mm) from wheat spikes 14 days post-anthesis. Pre-culture on high-osmolarity media (MS with 0.4M mannitol/sorbitol) for 4 hours.
- DNA Microparticle Preparation: Coat 1.0 µm gold particles with plasmid DNA (1 µg/µL) using CaCl₂ and spermidine precipitation. Wash and resuspend in 100% ethanol.
- Bombardment Parameters: Load macrocarrier with DNA/gold suspension. Use 1100 psi rupture disks under a vacuum of 28 inches Hg. Shoot at a target distance of 6 cm.
- Post-Bombardment Recovery: Incubate embryos on osmotic media overnight, then transfer to standard culture media without selection for 1 week.
- Selection & Regeneration: Transfer embryos to selection media (with phosphinothricin 5 mg/L). Subculture every 2 weeks. Transfer resistant calli to regeneration media (4-8 weeks).
- Molecular Confirmation: Conduct PCR, ELISA, and Southern blot analysis on regenerated plants to assess transgene integration and copy number.
Visualizing the Workflow Comparison
Diagram Title: Comparative Workflow of AMT vs. Biolistic Transformation
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Transformation Studies
| Item | Primary Function in Research | Example/Catalog Consideration |
|---|---|---|
| Binary Vector System (for AMT) | Carries T-DNA and virulence genes; essential for Agrobacterium gene transfer. | pCAMBIA1300 series, pGreen, Superbinary vectors. |
| Gold Microparticles (for Biolistic) | Inert carrier particles for coating and delivering DNA into cells via bombardment. | 0.6-1.0 µm diameter, sterile gold microcarriers. |
| Acetosyringone | Phenolic compound that induces Agrobacterium vir genes during co-cultivation. | Typically used at 100-200 µM in AMT co-cultivation media. |
| Selective Antibiotics/Herbicides | Eliminates non-transformed tissue; allows growth of transformants only. | Hygromycin B, Kanamycin, Phosphinothricin (BASTA/Glufosinate). |
| Plant Tissue Culture Media | Provides nutrients and hormones for explant survival, callus growth, and regeneration. | Murashige & Skoog (MS), N6 medium, specific hormone cocktails. |
| Strain-specific Agrobacterium | Engineered disarmed strain with high transformation efficiency for target species. | A. tumefaciens strains EHA105, LBA4404, GV3101. |
| Rupture Disks (for Biolistic) | Controls the helium gas pressure burst that propels DNA-coated particles. | Rated for specific pressures (e.g., 450 psi, 1100 psi, 2200 psi). |
This cost-benefit comparison demonstrates a clear trade-off. Agrobacterium-mediated transformation offers lower equipment and consumables costs, higher efficiency for stable, low-copy number integration, but demands significant microbiological and tissue culture labor. Biolistic transformation, while faster in initial DNA delivery and less restricted by plant genotype, incurs high capital equipment costs, higher consumable expenses, and can lead to complex integration patterns. The optimal choice remains contingent on the target species, desired transgene structure, and the laboratory's existing infrastructure and expertise.
The selection of a genetic transformation method is a foundational decision in plant biotechnology, synthetic biology, and molecular pharming. The enduring debate centers on the comparative efficiency of Agrobacterium tumefaciens-mediated transformation (AMT) versus biolistic (particle bombardment) methods. A broader thesis examining transformation efficiency must extend beyond simple DNA integration events to encompass factors such as transgene copy number, integrity, stability, and the specific requirements of the host species and desired molecular product. This guide provides an objective, data-driven comparison to inform project-specific tool selection.
Comparative Performance Data
Table 1: Direct Comparison of Key Transformation Metrics
| Performance Metric | Agrobacterium-Mediated Transformation | Biolistic Transformation | Supporting Data & Key References |
|---|---|---|---|
| Typical Transgene Copy Number | Low (1-3 copies) | High (often >5, can be fragmented) | AMT: ~80% of events are single-copy in rice. Biolistics: >60% events contain >5 copies in maize. |
| Transgene Integrity & Rearrangement | Generally high, precise T-DNA borders. | Frequent rearrangements, truncations. | Sequencing data shows >70% of AMT events have intact T-DNA vs. <30% for biolistics in wheat. |
| Delivery to Organelles | Not applicable. Nuclear targeting only. | Direct delivery to chloroplasts/mitochondria possible. | Successful stable transformation of chloroplasts in tobacco solely via biolistics. |
| Host Species Range (Plants) | Broad, but recalcitrance in major cereals historically. | Extremely broad, effective in monocots, trees, algae. | Protocol established for all major cereals via biolistics; AMT for maize/rice now routine. |
| Vector Requirements | Complex, requires T-DNA borders and virulence helper. | Simple, minimal plasmid or linear DNA cassette. | Biolistics can use plasmid-free, "clean DNA" cassettes to reduce bacterial sequences. |
| Cost & Throughput | Lower consumable cost, higher throughput for amenable species. | Higher equipment/licensing cost, moderate throughput. | AMT enables robotic handling of 1000s of explants; biolistics limited by chamber capacity. |
| Ideal Application | High-quality, single-copy events for regulatory approval & basic research. | Transformation of recalcitrant species, organelle engineering, species without Agrobacterium protocol. |
Detailed Experimental Protocols
Protocol 1: Standard Agrobacterium-Mediated Transformation of Tobacco Leaf Disks
- Objective: Stable nuclear transformation of Nicotiana tabacum.
- Methodology:
- Vector Preparation: Use a binary vector (e.g., pBIN19) with gene of interest within T-DNA borders. Transform into disarmed A. tumefaciens strain (e.g., LBA4404 or GV3101).
- Bacterial Culture: Grow Agrobacterium overnight in YEP + antibiotics. Resuspend to OD₆₀₀ ~0.5 in liquid MS co-cultivation medium with acetosyringone (200 µM).
- Explant Preparation: Surface-sterilize tobacco leaves, cut into 5x5 mm disks.
- Co-cultivation: Immerse disks in bacterial suspension for 10-30 min, blot dry, and place on solid co-cultivation medium for 2-3 days in the dark.
- Selection & Regeneration: Transfer disks to regeneration medium containing antibiotics (e.g., kanamycin) for plant selection and carbenicillin/cefotaxime to eliminate Agrobacterium.
- Shoot Induction: After 2-4 weeks, excise developing shoots and transfer to rooting medium with selection.
Protocol 2: Biolistic Transformation of Maize Immature Embryos
- Objective: Stable transformation of a monocot crop species.
- Methodology:
- DNA Preparation: Precipitate plasmid or linear DNA (1 µg/µL) onto gold or tungsten microparticles (0.6 µm). Use CaCl₂ and spermidine.
- Target Tissue Preparation: Harvest immature embryos (1.0-1.5 mm) from maize ears 10-15 days after pollination. Place scutellum-side up on osmotic conditioning medium.
- Bombardment Parameters: Use a PDS-1000/He or similar device. Place macrocarrier with coated particles 1 cm from stopping screen. Target tissue at 6 cm. Use helium pressure of 650-1100 psi and a vacuum of 26-28 in Hg.
- Post-Bombardment Recovery: Incubate tissues in the dark on osmotic medium for 16-24 hours.
- Selection & Callus Induction: Transfer embryos to callus induction medium with selective agent (e.g., bialaphos). Subculture every 2 weeks.
- Plant Regeneration: Transfer resistant, proliferating callus to regeneration medium to induce shoots and roots.
Visualization of Workflows & Pathways
Diagram 1: Agrobacterium vs Biolistic Workflow
Diagram 2: Agrobacterium T-DNA Transfer Mechanism
The Scientist's Toolkit: Essential Research Reagents & Materials
Table 2: Key Reagent Solutions for Plant Transformation
| Reagent / Material | Function & Role in Transformation | Application Notes |
|---|---|---|
| Binary Vector System (e.g., pBIN19, pCAMBIA) | Contains T-DNA borders for gene transfer and plant selection marker; used in Agrobacterium. | Essential for AMT. Choose based on replicon, selection, and promoter compatibility. |
| Disarmed A. tumefaciens Strain (e.g., LBA4404, GV3101) | Engineered to lack oncogenes but retain virulence (vir) genes for T-DNA transfer. | Strain choice impacts host range and transformation efficiency. |
| Acetosyringone | Phenolic compound that induces the Agrobacterium vir gene region. | Critical for AMT of many plant species, especially monocots. |
| Gold or Tungsten Microcarriers (0.6-1.0 µm) | Inert particles used as DNA carriers for bombardment in biolistics. | Gold is more uniform and less toxic; tungsten is less expensive. |
| Helium Particle Delivery System (e.g., PDS-1000/He) | Device uses a helium pressure pulse to accelerate DNA-coated particles into target cells. | Standard equipment for biolistics; requires optimization of pressure and distance. |
| Selective Agent (e.g., Kanamycin, Hygromycin, Bialaphos/Phosphinothricin) | Antibiotic or herbicide used to kill non-transformed tissue post-transformation. | Choice depends on plant species sensitivity and selectable marker gene (nptII, hpt, bar/pat). |
| Plant Tissue Culture Media (MS, B5 Basal Salts) | Provides nutrients, hormones, and support for explant survival, callus growth, and plant regeneration. | Must be precisely formulated with appropriate plant growth regulators (auxins, cytokinins). |
Conclusion
The choice between Agrobacterium-mediated and biolistic transformation is not a matter of one being universally superior, but of strategic selection based on project-specific goals. AMT generally offers advantages in generating low-copy, precise integration events with minimal transgene rearrangement, making it ideal for functional studies and regulatory-compliant therapeutic production. Biolistics provides a species-agnostic, rapid delivery method suited for difficult-to-transform cells and transient expression assays, albeit often with higher copy numbers and complex integration patterns. For biomedical researchers, the future lies in leveraging the strengths of each method—potentially in combination—and integrating newer precision tools like CRISPR to enhance targeting. Continued optimization of both techniques will be crucial for advancing scalable production of biologics, high-throughput functional genomics, and the development of next-generation gene therapies and plant-made pharmaceuticals.