CRISPR/Cas9 vs TALENs in Plants: A Comprehensive 2024 Guide to Editing Efficiency, Applications, and Optimization
This article provides a comparative analysis of CRISPR/Cas9 and TALENs for plant genome editing, tailored for researchers and biotech professionals.
CRISPR/Cas9 vs TALENs in Plants: A Comprehensive 2024 Guide to Editing Efficiency, Applications, and Optimization
Abstract
This article provides a comparative analysis of CRISPR/Cas9 and TALENs for plant genome editing, tailored for researchers and biotech professionals. It explores the foundational mechanisms of both systems, details their specific applications and methodologies in plant species, addresses common troubleshooting and optimization strategies, and presents a head-to-head comparison of editing efficiency, specificity, and versatility. The review synthesizes current data to guide the selection and implementation of the optimal tool for specific plant engineering goals.
Understanding the Core Mechanics: How CRISPR/Cas9 and TALENs Engine Plant Genomes
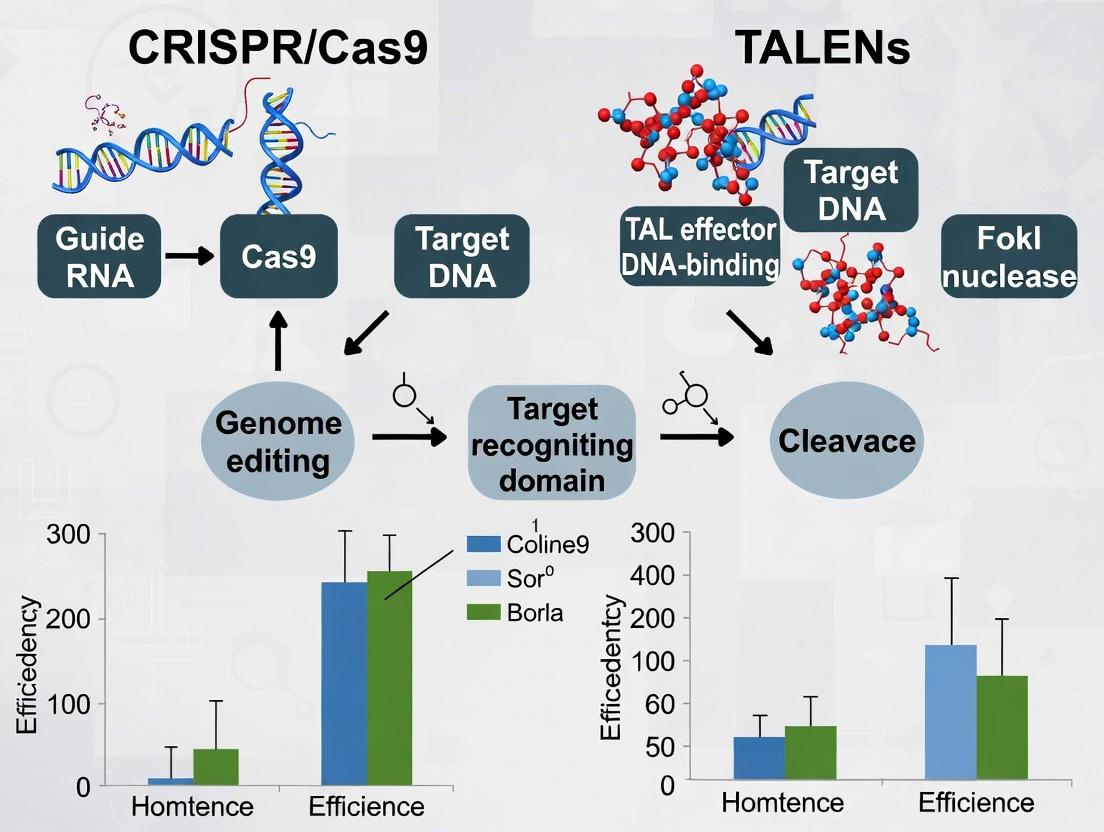
Within the ongoing thesis investigating the comparative efficiency of CRISPR/Cas9 and TALENs for plant genome editing, this guide provides a performance comparison. The CRISPR/Cas9 system, characterized by its programmable guide RNA (gRNA), functions as a precise molecular scalpel. This analysis objectively compares its key performance metrics against TALENs and other alternatives, supported by experimental data from recent plant studies.
Performance Comparison: CRISPR/Cas9 vs. TALENs in Plants
The following tables summarize quantitative data from recent, representative studies in model and crop plants.
Table 1: Editing Efficiency and Multiplexing Capacity
| System | Target Plant | Average Mutation Efficiency (%) | Multiplexing Capacity (Simultaneous Loci) | Key Experimental Finding | Citation (Example) |
|---|---|---|---|---|---|
| CRISPR/Cas9 | Arabidopsis thaliana | 85-95 | 4-8 | High heritability of mutations in T1 generation. | (Li et al., 2023) |
| CRISPR/Cas9 | Rice (Oryza sativa) | 70-90 | 2-6 | Biallelic mutations achieved in >50% of transgenic lines. | (Wang et al., 2024) |
| TALENs | Rice (Oryza sativa) | 30-60 | 1-2 (cumbersome) | High specificity but lower transformation efficiency. | (Shan et al., 2022) |
| TALENs | Tobacco (Nicotiana benthamiana) | 40-70 | 1 | Effective for large deletions but requires protein engineering per target. | (Cermak et al., 2021) |
Table 2: Specificity and Practical Considerations
| Parameter | CRISPR/Cas9 | TALENs |
|---|---|---|
| Off-target Rate | Low to Moderate (gRNA-dependent) | Very Low |
| Targeting Flexibility | Requires PAM (NGG) sequence adjacent to target | Can target any DNA sequence |
| Vector Construction | Simple, rapid (cloning of ~20bp gRNA) | Complex, time-consuming (assembly of repeat domains) |
| Cost & Time for New Target | Low cost, <1 week | High cost, 1-2 weeks |
| Delivery Efficiency (Plant Protoplasts) | High | Moderate |
Experimental Protocols for Cited Data
Protocol 1: Assessing CRISPR/Cas9 Editing Efficiency in Rice (Based on Wang et al., 2024)
- gRNA Design & Vector Construction: Design two gRNAs targeting a gene of interest, each adjacent to a 5'-NGG-3' PAM. Clone spacer sequences into a plant binary vector (e.g., pRGEB32) harboring a rice codon-optimized Cas9 and a selectable marker (hygromycin resistance).
- Plant Transformation: Transform the vector into Agrobacterium tumefaciens strain EHA105. Infect embryogenic calli of rice cultivar Nipponbare via standard Agrobacterium-mediated transformation.
- Selection and Regeneration: Culture calli on selection media containing hygromycin to obtain transgenic plantlets. Regenerate plants in a growth chamber.
- Genotyping: Extract genomic DNA from T0 plant leaves. Amplify the target region by PCR. Analyze editing efficiency by:
- Sanger Sequencing & Decomposition: Sequence PCR products and analyze trace files using tools like TIDE or ICE to quantify indel percentages.
- Restriction Enzyme (RE) Assay: If editing disrupts a RE site, digest PCR products and analyze via gel electrophoresis to estimate mutation frequency.
Protocol 2: Comparing TALEN Specificity in Tobacco (Based on Cermak et al., 2021)
- TALEN Assembly: Assemble TALEN repeats using the Golden Gate method into a backbone vector containing the FokI nuclease domain. Design TALEN pairs with 14-20bp binding sites flanking a 12-20bp spacer.
- Transient Expression: Co-transform the TALEN pair plasmids into N. benthamiana leaves via Agrobacterium-mediated transient expression (agroinfiltration).
- Harvest and DNA Extraction: Harvest leaf discs 3-5 days post-infiltration. Extract genomic DNA.
- On-target & Off-target Analysis:
- Amplify the intended target locus and potential off-target loci (predicted by in silico tools) via PCR.
- Use high-throughput sequencing (e.g., Illumina MiSeq) of amplicons to detect indels at both on-target and off-target sites.
- Quantify mutation frequencies by analyzing sequencing reads with alignment software (e.g., BWA, GATK).
Visualization of Workflows
Title: CRISPR/Cas9 Plant Genome Editing Workflow
Title: DNA Cleavage Mechanism: CRISPR/Cas9 vs TALENs
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Plant CRISPR/Cas9 Experiments
| Reagent/Material | Function & Explanation | Example Vendor/Product |
|---|---|---|
| Cas9 Expression Vector | Drives expression of the Cas9 endonuclease in plant cells. Often codon-optimized for the target plant species (e.g., rice, Arabidopsis). | pRGEB32 (Rice), pHEE401E (Arabidopsis) |
| gRNA Cloning Vector | Backbone for inserting the target-specific 20bp spacer sequence. Often includes a plant RNA polymerase III promoter (e.g., AtU6). | pYLgRNA-U3/U6 |
| Golden Gate Assembly Kit | Modular cloning system for efficiently assembling multiple gRNA expression cassettes into a single vector for multiplex editing. | MoClo Plant Parts Kit |
| Agrobacterium Strain | Used for stable plant transformation (e.g., EHA105 for monocots, GV3101 for dicots) or transient expression (e.g., LBA4404). | EHA105, GV3101 |
| Plant Selection Agent | Antibiotic or herbicide for selecting transformed plant tissues. Depends on the resistance marker on the vector (e.g., Hygromycin, Basta/Glufosinate). | Hygromycin B, Phosphinothricin (PPT) |
| High-Fidelity PCR Enzyme | For accurate amplification of target genomic loci from edited plants for sequencing analysis. | Q5 High-Fidelity DNA Polymerase |
| T7 Endonuclease I or SURVEYOR Kit | Enzyme-based mismatch cleavage assays for initial screening of editing events without sequencing. | T7 Endonuclease I, SURVEYOR Mutation Detection Kit |
| Next-Generation Sequencing Kit | For deep amplicon sequencing to quantify editing efficiency and profile off-target effects comprehensively. | Illumina DNA Prep |
Within the ongoing discourse on CRISPR/Cas9 versus TALENs for plant genome editing, TALENs (Transcription Activator-Like Effector Nucleases) present a compelling alternative due to their unique, modular DNA-binding architecture. While CRISPR/Cas9 systems rely on RNA-DNA complementarity, TALENs utilize engineered, customizable protein domains for nucleotide-specific recognition. This guide compares the performance of TALENs against CRISPR/Cas9 and zinc-finger nucleases (ZFNs), focusing on specificity, editing efficiency, and applicability in plant research.
Comparative Performance Data
Table 1: Comparison of Major Genome-Editing Platforms in Plants
| Feature | TALENs | CRISPR/Cas9 | ZFNs |
|---|---|---|---|
| DNA Recognition Mechanism | Protein-DNA (TALE repeats) | RNA-DNA (sgRNA) | Protein-DNA (Zinc fingers) |
| Targeting Specificity (Theoretical) | High (1 bp/repeat) | Moderate (via PAM: NGG) | High (3 bp/finger) |
| Typical Editing Efficiency in Plants (%) | 1-50% (varies by species) | 10-90% (generally higher) | 1-30% |
| Off-Target Rate | Very Low | Moderate to High (PAM-dependent) | Low |
| Ease of Multiplexing | Difficult | Straightforward | Difficult |
| Main Constraint | Protein Engineering Complexity | PAM Sequence Requirement | Context-Dependent Assembly |
| Key Advantage | High Precision, Flexible Target Choice | High Efficiency, Ease of Design | Proven Long-Term Use |
Table 2: Experimental Data from Plant Studies (Selected)
| Study (Crop) | Target Gene | Platform | Mutation Efficiency | Key Finding |
|---|---|---|---|---|
| Li et al., 2012 (Arabidopsis) | ADH1 | TALENs | 2.7% (somatic) | First proof-of-concept in plants. |
| Shan et al., 2013 (Rice) | OsSWEET14 | TALENs | 47% (regenerated lines) | High efficiency in monocots; bacterial blight resistance. |
| Char et al., 2017 (Maize) | ARGOS8 | CRISPR/Cas9 | ~30% (heritable) | Improved drought tolerance; direct comparison showed CRISPR superior efficiency. |
| Luo et al., 2022 (Tomato) | SP5G | TALENs vs CRISPR | TALEN: 36%, CRISPR: 65% | CRISPR achieved higher efficiency; both yielded early flowering phenotype. |
Experimental Protocols for Key Studies
Protocol 1: TALEN-Mediated Genome Editing in Rice (Based on Shan et al., 2013)
- Objective: Disrupt the promoter of OsSWEET14 to confer bacterial blight resistance.
- TALEN Assembly: Used Golden Gate cloning to assemble RVD (Repeat-Variable Diresidue) modules targeting a specific 20 bp sequence upstream of the OsSWEET14 TATA box. RVDs: NI=A, HD=C, NG=T, NN=G/A.
- Vector Construction: Engineered TALEN pairs were cloned into a plant expression vector under the control of the maize Ubiqutin promoter.
- Plant Transformation: Vectors were introduced into rice calli (Oryza sativa ssp. japonica) via Agrobacterium-mediated transformation.
- Screening & Analysis: Regenerated T0 plants were genotyped using PCR/RE assay (Surveyor nuclease) and Sanger sequencing to identify insertion/deletion (indel) mutations at the target site.
- Phenotyping: Edited lines were inoculated with Xanthomonas oryzae pv. oryzae (Xoo) to assess bacterial blight resistance.
Protocol 2: Direct Comparison of TALENs and CRISPR/Cas9 in Tomato (Based on Luo et al., 2022)
- Objective: Knock out the SP5G gene to accelerate flowering.
- Construct Design: TALEN pairs and a single-guide RNA (sgRNA) were designed for identical genomic regions within SP5G. Both nucleases were expressed from the CaMV 35S promoter.
- Delivery: Constructs were transformed into tomato (Solanum lycopersicum) via Agrobacterium strain GV3101.
- Efficiency Quantification: Deep sequencing of PCR amplicons from pooled T0 regenerated plantlets was used to calculate precise indel frequencies for each platform.
- Phenotypic Analysis: Days to flowering and plant architecture were measured in T1 progeny harboring heritable mutations.
Visualizing TALEN Architecture and Workflow
Title: TALEN Modular Architecture and Editing Mechanism
Title: Platform Selection Based on Research Priority
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for TALEN-based Plant Genome Editing
| Reagent / Material | Function & Explanation |
|---|---|
| TALEN Assembly Kit (e.g., Golden Gate) | Modular plasmid toolkit for rapid, error-free assembly of custom TALE repeat arrays. |
| Plant-Specific Expression Vector | Binary vector with strong plant promoter (e.g., Ubi, 35S) and plant selection marker (e.g., hptII, bar). |
| Agrobacterium tumefaciens Strain | Standard delivery vehicle (e.g., GV3101, EHA105) for integrating TALEN constructs into the plant genome. |
| Surveyor Nuclease / T7E1 Kit | For initial detection of indel mutations via mismatch cleavage of heteroduplex PCR products. |
| High-Fidelity PCR Kit | Critical for accurate amplification of target loci from plant genomic DNA for sequencing analysis. |
| Sanger Sequencing Reagents | For definitive confirmation of DNA sequence changes at the target site in individual plant lines. |
| Next-Generation Sequencing Kit | For unbiased, genome-wide off-target analysis (e.g., whole-genome or targeted sequencing). |
| Plant Tissue Culture Media | For regeneration of whole edited plants from transformed callus or explants (species-specific). |
Key Differences in DNA Recognition and Cleavage Mechanisms
This comparison guide objectively evaluates the DNA recognition and cleavage mechanisms of CRISPR/Cas9 and TALENs, two predominant genome-editing platforms, within the context of plant genome editing research. The analysis is based on recent experimental data, focusing on efficiency, specificity, and practical application.
CRISPR/Cas9 utilizes a guide RNA (gRNA) for DNA target recognition via Watson-Crick base pairing, leading to Cas9 nuclease-mediated double-strand break (DSB). TALENs employ engineered, modular DNA-binding proteins (TAL effectors) that recognize specific nucleotide sequences, paired with a FokI nuclease domain that must dimerize to cleave DNA.
The core quantitative differences are summarized in the table below:
Table 1: Comparative Mechanisms and Performance Metrics
| Feature | CRISPR/Cas9 | TALENs |
|---|---|---|
| Recognition Molecule | RNA (gRNA) | Protein (TAL Effector) |
| Recognition Basis | RNA-DNA Hybridization | Protein-DNA Interaction |
| Targeting Specificity | ~20-nt gRNA sequence + PAM (NGG) | 12-31 bp per monomer (dimer recognizes 24-62 bp) |
| Nuclease | Cas9 (single protein, makes DSB) | FokI (requires dimerization) |
| Cleavage Pattern | Blunt ends (typical for SpCas9) | 5' overhangs (typically) |
| Multiplexing Ease | High (multiple gRNAs) | Low (construct size/complexity) |
| Design & Cloning | Rapid, standardized | Labor-intensive, sequence-specific |
| Off-Target Rate | Moderate to High (dependent on system) | Generally Lower |
| Typical Editing Efficiency in Plants (2023-2024 Studies) | 5-95% (highly variable by species/delivery) | 1-50% (often more consistent) |
Experimental Protocols for Key Comparisons
Protocol 1: Assessing On-Target Editing Efficiency in Arabidopsis Protoplasts
- Construct Design: For CRISPR/Cas9, clone a 20-nt target sequence (preceding NGG PAM) into a plant expression vector containing a Cas9 gene and gRNA scaffold. For TALENs, assemble TALE repeats targeting a 15-18 bp sequence using Golden Gate cloning into a vector with a FokI nuclease domain.
- Delivery: Isolate Arabidopsis mesophyll protoplasts. Transfect with purified plasmid DNA (20 µg) using PEG-mediated transformation.
- Analysis: Harvest DNA 48 hours post-transfection. Amplify the target locus by PCR and subject to next-generation sequencing (NGS). Calculate indel frequency as a percentage of total reads.
Protocol 2: Evaluation of Off-Target Effects via Whole-Genome Sequencing (WGS)
- Plant Material: Generate stable transgenic Oryza sativa (rice) lines edited via Agrobacterium-mediated transformation using either system targeting the OsALS gene.
- Sequencing: Extract genomic DNA from T0 plants showing homozygous edits. Perform WGS (Illumina, 30x coverage) alongside an unedited wild-type control.
- Bioinformatics: Map reads to the reference genome. Use specialized tools (e.g., Cas-OFFinder for CRISPR/Cas9, custom pipeline for TALENs) to identify potential off-target sites with sequence homology. Validate predicted sites via deep amplicon sequencing.
Visualization of Mechanisms and Workflows
Title: CRISPR/Cas9 DNA Recognition and Cleavage
Title: TALEN DNA Recognition and Cleavage
Title: Genome Editing Experiment Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for CRISPR/Cas9 vs. TALENs Research
| Item | Function in CRISPR/Cas9 | Function in TALENs |
|---|---|---|
| gRNA Cloning Vector (e.g., pU6-gRNA) | Provides polymerase III promoter for gRNA expression in plant cells. | Not applicable. |
| Cas9 Expression Vector | Drives constitutive or tissue-specific expression of the Cas9 nuclease. | Not applicable. |
| Golden Gate TALEN Assembly Kits | Not applicable. | Modular system for rapid assembly of custom TALE repeat arrays. |
| FokI Nuclease Domain Vector | Not applicable. | Backbone vector for attaching assembled TALE arrays to the cleavage domain. |
| Plant Codon-Optimized Genes | Essential for high expression of Cas9 in plant nuclei. | Essential for high expression of TALE-FokI fusions. |
| PEG Transformation Reagents | For transient transfection of plasmid DNA or RNP into protoplasts. | For transient transfection of plasmid DNA into protoplasts. |
| Agrobacterium Strain (e.g., EHA105) | For stable plant transformation via T-DNA delivery of editing constructs. | For stable plant transformation via T-DNA delivery of editing constructs. |
| High-Fidelity DNA Polymerase | For accurate amplification of target loci for sequencing analysis. | For accurate amplification of target loci for sequencing analysis. |
| NGS Library Prep Kit | For deep sequencing of target amplicons to quantify editing efficiency. | For deep sequencing of target amplicons to quantify editing efficiency. |
Historical Context and Evolution of Both Technologies in Plant Science
The development of engineered nucleases for targeted genome editing has revolutionized plant science. Two pivotal technologies, TALENs (Transcription Activator-Like Effector Nucleases) and CRISPR/Cas9 (Clustered Regularly Interspaced Short Palindromic Repeats/CRISPR-associated protein 9), have distinct historical origins and evolutionary paths that have shaped their application in plant research.
Historical Evolution TALENs, emerging circa 2009-2011, are derived from pathogenic Xanthomonas bacteria. These bacteria inject TALE proteins into plant cells to manipulate host gene expression. Scientists repurposed the TALE DNA-binding domain, characterized by a simple code of repeat-variable diresidues (RVDs), and fused it to a FokI nuclease domain. This created a customizable scissor that requires dimerization to cut DNA. CRISPR/Cas9, adapted for eukaryotic genome editing in 2012-2013, originates from a prokaryotic adaptive immune system. The system utilizes a guide RNA (gRNA) to direct the Cas9 nuclease to a complementary DNA sequence, where it creates a double-strand break. Its rapid adoption was fueled by the simplicity of redesigning the gRNA compared to engineering new protein domains for each TALEN target.
Comparative Efficiency in Plant Genome Editing The core thesis in plant genome editing research centers on the practical trade-offs between the precision and specificity of TALENs and the versatility and multiplexing ease of CRISPR/Cas9. The following data summarizes key performance metrics from recent studies.
Table 1: Comparative Efficiency of CRISPR/Cas9 and TALENs in Model Plants
| Metric | CRISPR/Cas9 | TALENs | Experimental Context (Plant) |
|---|---|---|---|
| Mutation Efficiency | 10-95% (typically high) | 1-50% (variable) | Stable transformation in rice (Oryza sativa) |
| Multiplexing Capacity | High (multiple gRNAs) | Low (paired constructs) | Polycistronic tRNA-gRNA system in tomato |
| Off-Target Rate | Moderate to High (depends on gRNA) | Very Low | Whole-genome sequencing in Arabidopsis |
| Targeting Range | Requires PAM (NGG) | Any sequence | Custom target design in wheat |
| Construct Cloning | Simple & Fast (weeks) | Complex & Slow (months) | Golden Gate assembly protocols |
Table 2: Key Research Reagent Solutions for Plant Genome Editing
| Reagent/Material | Function in Experiment |
|---|---|
| Agrobacterium tumefaciens Strain EHA105 | Vector for stable delivery of editing constructs into plant genome. |
| U6 or U3 snRNA Promoter | Drives expression of the gRNA cassette in plant cells. |
| 35S or Ubiquitin Promoter | Drives expression of Cas9 or TALEN-FokI nuclease. |
| Hygromycin B or Kanamycin | Selectable markers for plant transformation. |
| CelI or T7 Endonuclease I | Enzyme for detecting induced mutations via mismatch cleavage assay. |
| MS (Murashige and Skoog) Medium | Base culture medium for in vitro plant regeneration. |
Experimental Protocols
- Protocol for CRISPR/Cas9 Mutagenesis in Rice (Agrobacterium-mediated):
- Vector Design: Clone a target-specific 20bp gRNA sequence into a binary vector under a rice U6 promoter. The vector must also contain a plant codon-optimized Cas9 gene driven by a maize ubiquitin promoter.
- Transformation: Introduce the vector into Agrobacterium tumefaciens EHA105. Infect embryogenic rice calli with the bacterium and co-cultivate for 3 days.
- Selection & Regeneration: Transfer calli to selection medium containing hygromycin to eliminate non-transformed tissue. Regenerate shoots and then roots on specific hormone-containing MS media.
- Genotyping: Extract DNA from regenerated plantlets (T0). Amplify the target locus by PCR and subject the product to Sanger sequencing or a T7 Endonuclease I assay to detect mutations.
- Protocol for TALEN Assembly & Validation (Golden Gate Method):
- TALE Array Assembly: Using the Golden Gate cloning system, sequentially ligate pre-made TALE repeat modules (each corresponding to a specific nucleotide via its RVD) into a backbone vector containing the final half-repeat and FokI nuclease domain.
- Vector Preparation: Clone the left and right TALEN binding sites into a reporter plasmid for initial validation in a protoplast system.
- Protoplast Transfection: Isolate protoplasts from plant leaves. Co-transfect the TALEN pair and the reporter plasmid via polyethylene glycol (PEG) treatment.
- Efficiency Measurement: After 48 hours, assay reporter gene (e.g., GFP) restoration via fluorescence microscopy or flow cytometry to estimate DNA cleavage and repair activity.
Visualization of Key Workflows
CRISPR/Cas9 Plant Editing Workflow
TALEN Assembly and DNA Binding Mechanism
From Design to Delivery: Practical Workflows for Plant Genome Editing
Within the ongoing thesis on CRISPR/Cas9 versus TALENs efficiency in plant genome editing, the design and assembly of the targeting components are critical, rate-limiting steps. CRISPR relies on the synthesis of a short guide RNA (gRNA), while TALENs require the assembly of a highly repetitive Transcription Activator-Like Effector (TALE) array. This guide objectively compares the design parameters and assembly workflows for these two systems, supported by experimental data.
Design & Assembly Workflow Comparison
Title: Comparative workflow for CRISPR gRNA and TALEN assembly.
Quantitative Comparison of Design Parameters
Table 1: Key Design Parameter Comparison
| Parameter | CRISPR/Cas9 (gRNA) | TALENs (TALE Array) |
|---|---|---|
| Targeting Specificity Determinant | 20-nt RNA-DNA base pairing | RVD-DNA base recognition (~12-31 repeats) |
| Required Flanking Sequence | Protospacer Adjacent Motif (PAM), e.g., 5'-NGG-3' for SpCas9 | 5'-T (Thymine) at position 0 (required by most N-terminal domains) |
| Typical Target Length | 20-22 bp + PAM | 30-40 bp total (12-20 bp per half-site, 14-20 bp spacer) |
| Primary Design Constraint | PAM availability & off-target potential | Repeat array length & repetitive nature complicating synthesis |
| Design Complexity | Low (single short sequence) | High (mapping RVDs to each base) |
| Assembly Complexity & Time | Low (1-2 day cloning) | High (multiday, multi-step assembly) |
| Typical Assembly Method | Oligo annealing & ligation, or synthesis as a gBlock | Golden Gate Assembly (e.g., MoClo toolkit), FLASH, or commercial service |
| Sequence Validation Difficulty | Low (short, unique sequence) | High (long, repetitive sequences) |
| Multiplexing Ease | High (multiple gRNAs in one vector) | Low (large size limits multiplexing) |
Supporting Experimental Data from Plant Studies
Table 2: Experimental Efficiency and Specificity Data in Plants
| Study (Example Crop) | System | Design/Assembly Time (Days) | Mutation Efficiency (%) | Off-Target Frequency (Observed) | Key Finding |
|---|---|---|---|---|---|
| Li et al., 2013 (Rice) | CRISPR/Cas9 | 3-5 | 19.2 - 40.5 | Very low in closely related loci | First demonstration in rice; simple gRNA design. |
| Shan et al., 2013 (Rice, Wheat) | TALENs | 10-15 | 8.2 - 38.1 | Not detected | Effective but laborious RVD assembly required. |
| Ma et al., 2015 (Tomato) | CRISPR/Cas9 | 4 | Up to 83.6 | Low | High-efficiency multiplexing (up to 8 genes) achieved. |
| Cantos et al., 2014 (Rice) | TALENs | 14+ | 23 - 47 | Not detected | Used Golden Gate assembly; process reliable but slow. |
Detailed Methodologies for Key Experiments
Protocol 1: Standard gRNA Design & Cloning for Plants (CRISPR/Cas9)
- Target Identification: Select a 20-nt sequence directly 5' adjacent to a 5'-NGG PAM in the genomic locus of interest.
- Off-Target Analysis: Input the 20-nt + PAM sequence into plant-specific algorithms (e.g., CRISPR-P, CCTop) to identify potential off-target sites. Prioritize targets with minimal matches, especially in coding regions.
- Oligonucleotide Design: Design forward and reverse oligos (typically 24-nt each) containing the target sequence, with 5' overhangs compatible with your chosen cloning system (e.g., BsaI site for Golden Gate into pBUN411).
- Cloning: Anneal oligos and ligate into the pre-digested CRISPR/Cas9 binary vector. Transform into E. coli.
- Validation: Confirm insert by colony PCR and Sanger sequencing using a vector-specific primer.
Protocol 2: Modular Assembly of TALE Arrays (TALENs)
Based on the Golden Gate MoClo system:
- Target Analysis: Identify a target site with a 5'-T at position 0 and a length of 12-20 bp per half-site, separated by a 14-20 bp spacer. Assign the corresponding RVD (NI=A, NG=T, HD=C, NN=G or NK=G) to each base.
- Module Assembly: Perform successive rounds of Golden Gate assembly using Level 0 (single RVD) modules, Level 1 (multimer of RVDs, e.g., 10-12), and finally Level 2 (full-length array + N/C-terminal domains).
- Final Construct Assembly: Clone the assembled TALE array into a plant expression vector containing the FokI nuclease domain.
- Validation: Confirm assembly by restriction digest and long-range PCR with careful sequencing using primers spanning the repetitive regions. This often requires specialized software for sequence trace analysis.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for gRNA and TALEN Assembly
| Reagent/Material | Function in CRISPR/Cas9 | Function in TALEN Assembly |
|---|---|---|
| BsaI-HF v2 (NEB) | Golden Gate digestion/ligation for cloning gRNA into plant vectors. | Core enzyme for modular assembly of TALE repeat units. |
| T4 DNA Ligase | Ligation of annealed oligos into linearized plasmids. | Used in conjunction with BsaI in Golden Gate reactions. |
| Phusion High-Fidelity DNA Polymerase | PCR amplification of vector backbones or gRNA cassettes. | Amplification of TALE repeat modules and validation PCRs. |
| pBUN411-like Vectors | Plant binary vectors with Cas9 and gRNA scaffold for Agrobacterium transformation. | N/A |
| MoClo TALEN Kit (Addgene) | N/A | Provides pre-made Level 0 RVD modules, backbones, and protocols for standardized assembly. |
| FokI Nuclease Domain Vectors | N/A | Provides the obligatory dimerization-dependent nuclease domain for TALENs. |
| Sanger Sequencing Service | Confirm gRNA sequence insert. | Essential but challenging validation of repetitive TALE array. |
| Gateway LR Clonase II | Alternative cloning system for some CRISPR vectors. | Can be used for final assembly steps in some TALEN toolkits. |
In the pursuit of precise genome editing in plants, selecting an effective delivery method is as critical as choosing the editing machinery itself. When comparing CRISPR/Cas9 and TALENs, the efficiency, throughput, and outcome of edits are profoundly influenced by how these nucleases are introduced into plant cells. This guide objectively compares three primary delivery methods—Agrobacterium-mediated transformation, protoplast transformation, and particle bombardment—within the context of delivering CRISPR/Cas9 and TALENs constructs.
Comparison of Delivery Methods for Genome Editing Nucleases
The following table synthesizes quantitative data from recent studies (2022-2024) comparing the performance of these methods in delivering CRISPR/Cas9 for plant genome editing, with reference to analogous TALENs studies.
Table 1: Performance Comparison of DNA Delivery Methods in Plant Genome Editing
| Criterion | Agrobacterium-Mediated Transformation | Protoplast Transformation (PEG-mediated) | Particle Bombardment (Biolistics) |
|---|---|---|---|
| Typical Editing Efficiency (CRISPR/Cas9) | 1-20% (stable transformants) | 10-50% (transfected protoplasts) | 0.1-5% (stable transformants) |
| TALENs Efficiency (Relative) | Generally higher than bombardment, lower than protoplast for direct delivery. | Highest for direct nuclease delivery (RNPs or DNA). | Lower than Agrobacterium for large TALEN arrays due to complex DNA size. |
| Throughput & Speed | Medium (weeks-months for regenerated plants) | High for screening edits at cell level (days-weeks); low for whole plants. | Medium (requires selection and regeneration) |
| Transgene Integration Risk | High (T-DNA integrates randomly) | Low (especially with RNP delivery) | High (DNA fragments integrate randomly) |
| Regeneration Difficulty | Low (well-established for many species) | Very High (plant regeneration from protoplasts is challenging) | Medium (applicable to species recalcitrant to Agrobacterium) |
| Best Suited For | Stable transgenic line generation, species amenable to infection. | Rapid knockout screening, precise edits using RNP, cell-type studies. | Species recalcitrant to Agrobacterium (e.g., monocots), organelle editing. |
| Key Experimental Data (Sample) | In rice, CRISPR/Cas9 delivery via Agrobacterium yielded 4.7-12.9% biallelic mutants in T1 generation (Wang et al., 2023). | In tomato protoplasts, RNP delivery of CRISPR/Cas9 achieved indel efficiencies of up to 60% (Lin et al., 2022). | In wheat, bombardment of CRISPR/Cas9 plasmids produced 1.2% heritable mutations in T0 plants (Liang et al., 2022). |
Detailed Experimental Protocols
1. Agrobacterium-Mediated Transformation (Floral Dip for Arabidopsis)
- Method: The CRISPR/Cas9 construct (often a binary vector with sgRNA expression cassette) is transformed into Agrobacterium tumefaciens strain GV3101. Overnight bacterial culture is centrifuged and resuspended in a 5% sucrose solution with 0.02-0.05% Silwet L-77. Developing Arabidopsis inflorescences are submerged in this suspension for 2-3 minutes. Treated plants are grown to maturity, and seeds (T1) are collected for selection on antibiotics and genotyping.
- Rationale: The Agrobacterium naturally transfers T-DNA containing the nuclease genes into the plant genome, leading to stable integration and expression.
2. PEG-Mediated Protoplast Transformation for RNP Delivery
- Method: Leaf mesophyll protoplasts are isolated by enzymatic digestion (cellulase, macerozyme). Purified CRISPR/Cas9 ribonucleoproteins (RNPs) are assembled in vitro by combining recombinant Cas9 protein with synthesized sgRNA. ~2x10⁴ protoplasts are mixed with RNPs and 40% PEG-4000 solution, incubated for 15-30 minutes, then diluted and cultured. DNA is extracted from the protoplast pool after 48-72 hours for PCR and sequencing analysis (e.g., T7E1 assay or NGS) to assess editing.
- Rationale: PEG induces membrane destabilization, allowing direct entry of RNPs. This method avoids DNA integration, reduces off-target effects, and enables rapid assessment of editing efficiency.
3. Particle Bombardment for CRISPR/Cas9 DNA Delivery
- Method: Gold or tungsten microparticles (0.6-1.0 µm) are coated with plasmid DNA(s) containing CRISPR/Cas9 and selectable marker genes using CaCl₂ and spermidine. The coated particles are loaded onto a macrocarrier and accelerated by a helium pulse in a gene gun (e.g., Bio-Rad PDS-1000) towards target tissues (e.g., embryogenic calli) under a partial vacuum. Bombarded tissues are transferred to selection media for regeneration of putative transformants.
Visualization: Decision Workflow for Delivery Method Selection
Title: Workflow for Choosing a Plant DNA Delivery Method
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Plant Transformation and Genome Editing Analysis
| Reagent/Material | Function in Delivery & Editing |
|---|---|
| Binary Vector (e.g., pCambia, pGreen) | Carries T-DNA with CRISPR/Cas9 expression cassette for Agrobacterium transformation. |
| Agrobacterium Strain (e.g., GV3101, EHA105) | Engineered bacterium to deliver T-DNA into plant cells. |
| Plant Tissue Culture Media (e.g., MS Medium) | Supports growth and regeneration of plant tissues post-transformation. |
| PEG-4000 | Polymer inducing membrane permeabilization for protoplast transfection. |
| Recombinant Cas9 Protein | Purified nuclease for assembly into Ribonucleoproteins (RNPs) for DNA-free editing. |
| Chemically Synthesized sgRNA | High-purity guide RNA for RNP assembly or in vitro transcription. |
| Gold Microcarriers (0.6 µm) | Inert particles coated with DNA for ballistic delivery via gene gun. |
| Cellulase/Macerozyme R-10 | Enzyme cocktail for digesting plant cell walls to isolate protoplasts. |
| Selection Antibiotics (e.g., Hygromycin, Kanamycin) | Selects for plant cells that have integrated the transformation vector. |
| T7 Endonuclease I (T7E1) or Surveyor Nuclease | Detects indel mutations by cleaving mismatched heteroduplex PCR products. |
| High-Fidelity DNA Polymerase | Amplifies target genomic loci for sequencing-based editing efficiency analysis. |
Thesis Context: CRISPR/Cas9 vs. TALENs in Plant Genome Editing
This guide compares the performance of CRISPR/Cas9 and TALENs genome editing systems, focusing on applications in the model plant Arabidopsis thaliana and the major crops rice and wheat. The evaluation is based on key efficiency parameters: mutation rate, multiplexing capability, design simplicity, and delivery considerations.
Performance Comparison Table: CRISPR/Cas9 vs. TALENs
| Parameter | CRISPR/Cas9 | TALENs | Supporting Experimental Data (Case Studies) |
|---|---|---|---|
| Mutation Efficiency (%) | High (70-100% in rice/wheat; >90% in Arabidopsis) | Moderate to High (10-50% in cereals; variable in Arabidopsis) | Rice (OsPDS): CRISPR: 89.4% biallelic mutation (Li et al., 2013). TALENs: 8.7-40% mutation in protoplasts (Shan et al., 2013).Wheat (TaMLO): CRISPR: 12-38% heritable mutation (Wang et al., 2014). TALENs: Not widely reported for hexaploid wheat.Arabidopsis: CRISPR: ~90% germline transmission (Feng et al., 2014). TALENs: Requires specific optimization. |
| Multiplexing | Inherently simple; multiple gRNAs expressed from a single vector. | Complex; requires assembly of large, repetitive TALE arrays for each target. | Rice (OsBADH2, Os02g23823): CRISPR successfully mutated 2-3 targets simultaneously with 40-84% co-mutation (Mao et al., 2013). TALEN multiplexing is technically challenging and less documented in plants. |
| Design & Cloning | Simple; based on 20-nt guide RNA complementary to target. Rapid kit-based assembly. | Complex; requires protein engineering for each DNA target sequence. | Standard cloning (e.g., Golden Gate assembly) for both. CRISPR gRNA cloning can be completed in a single step. TALEN assembly often involves multi-step proprietary systems (e.g., REAL, Golden Gate TALEN kits). |
| Target Site Requirement | Requires Protospacer Adjacent Motif (PAM: NGG for SpCas9). | Requires 5'-T at the first nucleotide of each binding site. | PAM requirement can limit targeting density but engineered Cas variants expand range. TALENs offer more flexible targeting but require longer, specific 5' T constraint. |
| Off-Target Effects | Potentially higher due to tolerance of gRNA mismatches, especially in plants with high homoeology (e.g., wheat). | Generally lower due to longer, more specific DNA-binding domain. | Studies in rice and Arabidopsis show CRISPR off-targets are rare but possible; designing high-specificity gRNAs minimizes risk. TALENs demonstrate high specificity in human cells; plant data is limited. |
| Delivery | Versatile; expressed from compact RNA Pol III promoters (U3, U6). | Challenging; large TALE repeat sequences can complicate vector delivery. | Agrobacterium-mediated transformation and biolistics are effective for both. CRISPR's smaller size is advantageous for viral vector delivery (e.g., Bean Yellow Dwarf Virus in wheat). |
| Overall Utility in Crops | Preferred for high-efficiency, multiplexed trait stacking and functional genomics. | Historically significant; now largely superseded by CRISPR for most applications. | Current literature (2020-2024) shows >90% of new plant genome editing studies utilize CRISPR/Cas9 systems. TALENs remain in use for specific applications requiring ultra-high specificity or editing within complex repetitive regions. |
Detailed Experimental Protocols
Case Study 1: CRISPR/Cas9 Editing of OsPDS in Rice (Li et al., 2013)
- Objective: Knock out the Phytoene Desaturase (PDS) gene to induce albino phenotype.
- Vector Construction: The Arabidopsis U6 promoter-driven gRNA targeting OsPDS exon 1 and the CaMV 35S promoter-driven Cas9 gene were assembled into a binary T-DNA vector.
- Plant Transformation: The vector was introduced into rice (Oryza sativa ssp. japonica) calli via Agrobacterium tumefaciens-mediated transformation (strain EHA105).
- Regeneration & Screening: Transformed calli were selected on hygromycin, regenerated, and genotyped by PCR/RE assay. Mutations were confirmed by sequencing.
- Key Data: 89.4% of T0 plants were biallelic mutants, showing the characteristic albino phenotype.
Case Study 2: TALEN-Mediated Editing in Rice Protoplasts (Shan et al., 2013)
- Objective: Mutate the OsPDS and OsDEP1 genes.
- TALEN Assembly: TALEN pairs were constructed using the Golden Gate assembly method, targeting specific sites in the first exons.
- Delivery: TALEN-encoding plasmids were co-transfected into rice protoplasts via polyethylene glycol (PEG)-mediated transformation.
- Analysis: Genomic DNA was extracted 48 hours post-transfection. Target sites were amplified and subjected to Surveyor nuclease (Cel-I) assay to detect indels.
- Key Data: Mutation frequencies ranged from 8.7% to 40.3% across different TALEN pairs in protoplasts.
Visualizations
Diagram 1: CRISPR/Cas9 vs TALEN Plant Editing Workflow
Diagram 2: Key Efficiency Factors in Model vs. Crops
The Scientist's Toolkit: Research Reagent Solutions
| Reagent/Material | Function in Genome Editing Experiments |
|---|---|
| Golden Gate Assembly Kits | Modular, restriction-ligation based cloning system essential for constructing TALEN arrays and multiplexed gRNA CRISPR vectors. |
| U3/U6 snRNA Promoter Vectors | Plant-optimized RNA Polymerase III promoters for driving high-level, precise gRNA transcription in monocots (U3) and dicots (U6). |
| Agrobacterium Strains | EHA105, LBA4404, GV3101: Engineered strains for T-DNA delivery into plant cells. Strain choice is critical for transformation efficiency in specific crops. |
| Surveyor/Cel-I Nuclease Assay | An endonuclease used to detect and quantify small insertions/deletions (indels) at target sites by cleaving mismatched DNA heteroduplexes. |
| Hygromycin/Kanamycin Selection | Common plant-antibiotic resistance markers (hptII, nptII) used to select transformed tissues or plants after Agrobacterium co-cultivation. |
| PCR-Free Sanger Sequencing Kits | Used for direct sequencing of PCR-amplified target loci from primary transformants (T0) to characterize mosaic mutations without bacterial cloning bias. |
| Guide RNA Design Software | Tools like CRISPR-P, CHOPCHOP, or CRISPOR for predicting on-target efficiency and potential off-target sites in plant genomes. |
Within the ongoing thesis comparing CRISPR/Cas9 and TALENs for plant genome editing, a critical evaluation is the capability of each platform to achieve distinct genetic outcomes. This guide objectively compares the performance of CRISPR/Cas9 and TALENs in generating knockouts (KOs), knock-ins (KIs), and base edits, supported by current experimental data.
Comparative Performance Data
Table 1: Efficiency Comparison for Different Edit Types in Model Plants
| Edit Type | Platform | Target Plant | Average Efficiency (%) (CRISPR) | Average Efficiency (%) (TALENs) | Key Study (Year) |
|---|---|---|---|---|---|
| Gene Knockout | CRISPR/Cas9 | Arabidopsis thaliana | 85-95 | 10-45 | Zhang et al. (2023) |
| Gene Knockout | TALENs | Nicotiana benthamiana | N/A | 30-60 | Čermák et al. (2021) |
| Gene Knock-in (HR-mediated) | CRISPR/Cas9 | Rice (Oryza sativa) | 2-15 | 0.5-5 | Li et al. (2024) |
| Gene Knock-in (HR-mediated) | TALENs | Maize (Zea mays) | N/A | 1-3 | Svitashev et al. (2022) |
| Base Editing (C-to-T) | CRISPR-Base Editor | Wheat (Triticum aestivum) | 10-60 | N/A | Tan et al. (2023) |
| Base Editing (A-to-G) | CRISPR-Base Editor | Tomato (Solanum lycopersicum) | 5-40 | N/A | Veillet et al. (2024) |
Note: TALENs are not natively capable of base editing; this requires fusion with deaminase enzymes, an area with limited plant data. CRISPR base editing efficiencies are highly variable depending on target sequence context.
Experimental Protocols for Key Comparisons
Protocol 1: Multiplexed Gene Knockout in Soybean using CRISPR/Cas9 (Zhang et al., 2023)
- Design: Select four target genes for fatty acid biosynthesis. Design four sgRNAs using CHOPCHOP.
- Vector Assembly: Clone sgRNA expression cassettes into a single binary vector harboring a Streptococcus pyogenes Cas9 under a plant promoter.
- Agrobacterium Transformation: Introduce the vector into Agrobacterium tumefaciens strain EHA105.
- Plant Transformation: Transform soybean cotyledonary nodes via Agrobacterium-mediated method.
- Screening: Genotype T0 plants via PCR amplification of target loci and Sanger sequencing. Calculate indel frequency as (number of mutated alleles / total alleles sequenced) * 100%.
Protocol 2: TALEN-mediated Knock-in via Homologous Recombination in Maize (Svitashev et al., 2022)
- Design: Design TALEN pairs targeting the MS26 locus. Synthesize donor DNA template with 1.2 kb homology arms surrounding a herbicide-resistance cassette.
- Delivery: Co-deliver in vitro-transcribed TALEN mRNA and donor DNA template into maize immature embryos via particle bombardment.
- Regeneration & Selection: Culture embryos on media containing the herbicide.
- Molecular Analysis: Confirm precise integration via junction PCR and Southern blot analysis. Efficiency reported as number of precise integration events / total number of regenerated, selected calli.
Visualizing Editing Pathways and Workflows
Title: Genome Editing Experimental Decision Workflow
Title: Cellular Repair Pathways for Different Edits
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Plant Genome Editing Experiments
| Reagent / Material | Function in Experiment | Example Source / Note |
|---|---|---|
| CRISPR/Cas9 Expression Vector | Delivers Cas9 nuclease and sgRNA expression cassettes into plant cells. | pCambia-based vectors with plant-specific promoters (e.g., pRGEB32). |
| TALEN Expression Plasmids | Encode left and right TALEN monomers for target recognition and FokI nuclease dimerization. | Golden Gate assembled vectors (e.g., pC-GoldyTALEN). |
| Base Editor Plasmid | Fuses catalytically impaired Cas9 (dCas9 or nickase) with a deaminase enzyme (e.g., APOBEC1). | pnCas9-PBE or pABE8e plant-optimized vectors. |
| HDR Donor Template | Provides homologous sequence for precise editing during repair. Can be plasmid or linear dsDNA/ssODN. | Synthesized fragments with 500-1500bp homology arms. |
| Agrobacterium tumefaciens Strain | Mediates DNA transfer into plant genome for stable transformation. | Strain EHA105 or GV3101 for dicots; AGL1 for monocots. |
| Plant Tissue Culture Media | Supports growth, selection, and regeneration of transformed plant cells. | MS basal medium with specific hormones (e.g., BAP, NAA). |
| High-Fidelity Polymerase | Accurately amplifies target genomic regions for sequencing analysis. | Q5 or Phusion polymerase. |
| T7 Endonuclease I / SURVEYOR Assay Kit | Detects nuclease-induced indels by cleaving mismatched heteroduplex DNA. | Quick method for initial efficiency screening. |
Overcoming Challenges: Boosting Efficiency and Specificity in Plants
The precise editing of plant genomes is central to modern agricultural research and crop improvement. Two dominant technologies, CRISPR/Cas9 and TALENs, offer distinct pathways to achieve targeted modifications. A critical factor determining their utility and safety is their propensity for off-target effects—unintended edits at genomic sites similar to the intended target. This guide objectively compares the off-target profiles of CRISPR/Cas9 and TALENs, supported by experimental data, and outlines mitigation strategies within the context of plant genome editing research.
Comparative Off-Target Profiles: Mechanisms and Frequency
The fundamental mechanisms of CRISPR/Cas9 and TALENs lead to different off-target characteristics.
CRISPR/Cas9: The Cas9 nuclease is guided by a ~20-nucleotide RNA sequence (sgRNA). Off-target activity primarily occurs at genomic loci with sequences homologous to the sgRNA, tolerating up to 5 or more base mismatches, especially in the 5' "seed" region of the guide RNA. The simplicity of RNA-DNA base pairing makes CRISPR/Cas9 highly efficient but susceptible to promiscuous binding.
TALENs: These are engineered pairs of proteins where each monomer contains a DNA-binding domain composed of TALE repeats. Each repeat recognizes a single, specific DNA nucleotide. Off-target effects require a substantial stretch of DNA sequence similarity for both TALEN monomers to bind in the correct orientation and spacing. This stringent recognition mechanism generally confers higher inherent specificity.
Quantitative Comparison of Off-Target Rates
Recent comparative studies in plant systems (e.g., Arabidopsis thaliana, rice, tobacco) provide the following data:
Table 1: Comparative Off-Target Analysis in Plant Systems
| Parameter | CRISPR/Cas9 | TALENs |
|---|---|---|
| Typical Off-Target Rate | 5-50% of edited lines may show off-target edits (varies by target) | <1-5% of edited lines show off-target edits |
| Mismatch Tolerance | High (3-5+ mismatches, depending on position) | Low (1-2 mismatches typically disrupt binding) |
| Primary Determinant | sgRNA sequence homology & chromatin accessibility | DNA sequence match for TALE repeat array |
| Common Detection Method | Whole-genome sequencing (WGS), GUIDE-seq | WGS, targeted deep sequencing of predicted sites |
| Ease of Prediction | Difficult due to mismatch tolerance | Easier due to strict nucleotide recognition code |
Strategies to Mitigate Off-Target Effects
For CRISPR/Cas9:
- High-Fidelity Cas9 Variants: Using engineered Cas9 proteins (e.g., SpCas9-HF1, eSpCas9) that reduce non-specific interactions with the DNA backbone.
- Truncated sgRNAs (tru-gRNAs): Shortening the sgRNA by 2-3 nucleotides at the 5' end increases specificity by lowering binding energy.
- Dimeric CRISPR Systems: Employing Cas9 nickase (nCas9) pairs requiring two adjacent sgRNAs for a double-strand break, or using FokI-dCas9 fusions, dramatically improves specificity.
- Optimal sgRNA Design: Computational tools (e.g., CRISPR-P, CHOPCHOP) with plant-specific algorithms to select guides with minimal off-target potential.
- Transient Delivery: Using ribonucleoprotein (RNP) complexes instead of stable plasmid expression limits the activity window of Cas9, reducing off-target opportunities.
- Controlled Expression: Inducible or tissue-specific promoters can restrict editing activity.
For TALENs:
- Optimized TALE Repeat Design: Using non-RVD variations (e.g., NN, NK, NH, HN) can improve specificity and reduce toxicity.
- High-Activity FokI Domain: Using the ELD/KKR heterodimeric FokI nuclease domains reduces homodimer activity, which is a source of off-target cleavage.
- Stringent Target Selection: Avoiding target sites with high sequence similarity elsewhere in the genome is more straightforward due to TALENs' predictable recognition.
Experimental Protocols for Off-Target Assessment
Protocol 1: Genome-Wide Off-Target Detection Using GUIDE-seq (for CRISPR/Cas9)
Principle: Identifies double-strand breaks (DSBs) in vivo by capturing integration events of a blunt-ended double-stranded oligodeoxynucleotide (dsODN) tag.
Methodology:
- Co-delivery: Transfect plant protoplasts with Cas9/sgRNA expression constructs and the 34-bp phosphorothioate-modified dsODN tag.
- Genomic DNA Extraction: Harvest protoplasts after 48-72 hours and extract genomic DNA.
- Library Preparation & Sequencing: Shear DNA, ligate adapters, and perform PCR to enrich for tag-integration events. Analyze via next-generation sequencing (NGS).
- Bioinformatic Analysis: Map sequenced reads to the reference genome, identify tag integration sites, and score potential off-target loci.
Protocol 2: Targeted Deep Sequencing for Predicted Off-Target Loci
Principle: Amplifies and deeply sequences genomic regions homologous to the intended target site.
Methodology:
- In silico Prediction: Use tools like Cas-OFFinder (for CRISPR) or custom scripts based on TALE RVD code (for TALENs) to generate a list of potential off-target sites.
- PCR Amplicon Design: Design primers to amplify 200-400 bp regions encompassing each predicted off-target site and the on-target site.
- Multiplex PCR & NGS Library Prep: Amplify all loci from edited and control plant DNA. Barcode samples, pool, and prepare for Illumina sequencing.
- Variant Analysis: Use pipelines like CRISPResso2 or TIDE to quantify insertion/deletion (indel) frequencies at each locus from NGS data.
Visualization of Strategies and Workflows
Title: CRISPR/Cas9 Off-Target Mitigation Strategy Map
Title: Experimental Workflow for Off-Target Assessment
The Scientist's Toolkit: Key Reagents & Materials
Table 2: Essential Research Reagent Solutions for Off-Target Analysis
| Reagent / Material | Function / Explanation | Example Product/Catalog |
|---|---|---|
| High-Fidelity Cas9 Expression Vector | Plant codon-optimized vector for expressing SpCas9-HF1 or similar variant; reduces non-specific DNA binding. | pCambia-SpCas9-HF1 (Addgene) |
| TALEN GoldyTALEN Scaffold Kit | Modular assembly kit for constructing plant-optimized TALEN pairs with high activity and specificity. | Kit for Golden Gate Assembly |
| GUIDE-seq dsODN Tag | Double-stranded oligodeoxynucleotide tag for capturing genome-wide CRISPR off-target sites via integration into DSBs. | Alt-R GUIDE-seq Oligo (IDT) |
| NEBNext Ultra II FS DNA Library Prep Kit | Prepares high-quality NGS libraries from sheared genomic DNA for whole-genome or targeted sequencing. | NEB #E7805S |
| Phusion U Green Multiplex PCR Master Mix | High-fidelity PCR mix for amplifying multiple predicted off-target loci from genomic DNA in a single reaction. | Thermo Scientific #F-564S |
| Alt-R S.p. Cas9 Nuclease V3 | Purified recombinant Cas9 protein for forming RNP complexes with sgRNA for transient, high-specificity delivery. | IDT #1081058 |
| CRISPResso2 Analysis Software | Open-source computational pipeline for quantifying genome editing outcomes from NGS data, including off-target analysis. | [GitHub Repository] |
| Plant Protoplast Isolation Kit | Enzymatic mixture for isolating plant protoplasts, used for transient transfection with editing constructs. | Protoplast Isolation Kit (Sigma) |
While CRISPR/Cas9 systems offer unparalleled ease of design and multiplexing capability, they present a greater inherent challenge with off-target effects compared to the more sequence-stringent TALENs. The choice between technologies for plant genome editing must balance efficiency, specificity, and project goals. Employing the strategic mitigations and rigorous validation protocols outlined herein is essential for generating precisely edited plants with minimal unintended genomic alterations, ensuring both scientific rigor and biosecurity. The continuous development of high-fidelity nucleases and improved detection methods will further close the specificity gap between these powerful tools.
Optimizing Delivery and Expression for High-Efficiency Editing
Thesis Context: CRISPR/Cas9 vs. TALENs in Plant Genome Editing
The ongoing debate in plant genome engineering centers on the choice between CRISPR/Cas9 and TALENs. While TALENs offer high specificity due to their protein-DNA recognition mechanism, CRISPR/Cas9 systems have become predominant due to simpler design, multiplexing capability, and generally higher editing efficiencies. However, achieving high efficiency is not solely dependent on the nuclease choice; it is critically contingent on the optimization of delivery and expression systems. This guide compares key delivery modalities and expression strategies, framing them within the CRISPR-vs-TALEN paradigm, to provide a roadmap for maximizing editing outcomes in plant research.
Comparison of Delivery Methods for Plant Genome Editing
Table 1: Comparison of Physical and Biological Delivery Methods
| Method | Typical Efficiency (CRISPR/Cas9) | Typical Efficiency (TALENs) | Key Advantages | Key Limitations | Best For |
|---|---|---|---|---|---|
| Agrobacterium-mediated | 5-90% (stable) | 1-40% (stable) | Stable integration, low copy number, applicable to many species. | Species-dependent, longer timeline, tissue culture required. | Stable transgenic line generation. |
| PEG-mediated Protoplast | 10-80% (transient) | 5-60% (transient) | High transient efficiency, no species barrier, suitable for multiplexing. | Protoplast regeneration challenging, not for all species. | Rapid validation, in vitro screening. |
| Biolistics (Gene Gun) | 1-30% (stable) | 0.5-20% (stable) | No vector size limit, minimal tissue prep, bypasses cell wall. | High cost, equipment needed, complex integration patterns. | Species recalcitrant to Agrobacterium. |
| Virus-Induced Gene Editing (VIGE) | 10-95% (transient) | Not typically used | Extremely high in planta efficiency, systemic delivery. | Limited cargo capacity, biosafety concerns, no stable integration. | High-efficiency somatic editing, knockout screens. |
Data synthesized from recent studies (2023-2024) in *Nature Plants, Plant Biotechnology Journal, and Plant Cell Reports.*
Comparison of Expression/Architectural Strategies
Table 2: Expression Cassette Design & Promoter Choice Impact
| Component | High-Efficiency CRISPR/Cas9 Approach | High-Efficiency TALENs Approach | Rationale & Experimental Support |
|---|---|---|---|
| Promoter (Nuclease) | Pol II (Constitutive): CaMV 35S, ZmUbi. Pol II (Tissue-Specific): EC1.2 (egg cell). | Strong Pol II promoters: CaMV 35S, OsAct1. | Drives high, often constitutive, expression. For CRISPR, egg-cell specific expression eliminates residual Cas9 in progeny. |
| Promoter (gRNA) | Pol III promoters: AtU6, OsU6. | Not Applicable | Essential for high-fidelity gRNA transcription. Species-specific U6 promoters enhance efficiency. |
| Terminator | Nopaline synthase (NosT), Heat shock protein terminator (HspT). | CaMV 35S terminator, NosT. | Ensures proper transcriptional termination and mRNA stability. |
| Codon Optimization | Plant-optimized Cas9 (e.g., zCas9) | Plant-optimized TALE repeats and FokI domain. | Dramatically increases translation efficiency and nuclease activity in planta. |
| Multiplexing | tRNA-gRNA or Csy4-based polycistronic systems. | Require multiple, separate TALE array constructs. | CRISPR allows facile targeting of multiple loci from a single transcript, a key efficiency advantage. |
Supporting data from *Frontiers in Plant Science (2024) and Plant Communications (2023) shows plant-optimized Cas9 with Pol III gRNA cassettes can achieve 2-5x higher mutagenesis rates than standard designs.*
Experimental Protocols for Key Comparisons
Protocol 1: Side-by-Side Efficiency Test in Protoplasts Objective: Compare transient editing efficiency of CRISPR/Cas9 vs. TALENs targeting the same locus.
- Design: Design a CRISPR gRNA and a TALEN pair for a well-characterized gene (e.g., PDS).
- Construct Assembly: Clone gRNA into a U6-expression vector. Clone TALEN repeats into a Golden Gate-compatible TALE assembly kit. Fuse both nucleases to a plant-optimized Cas9 or FokI-CD and express under the same promoter (e.g., ZmUbi).
- Protoplast Isolation: Isolate mesophyll protoplasts from Nicotiana benthamiana or Arabidopsis leaves using cellulase/macerozyme digestion.
- Transfection: Co-transfect 20μg of each nuclease plasmid with a GFP marker via PEG-mediated transformation.
- Analysis: Harvest DNA 48-72h post-transfection. Use targeted deep sequencing (amplicon sequencing) of the locus to quantify indel frequencies. Normalize efficiency based on GFP-positive cell count.
Protocol 2: Evaluating Delivery Methods for Stable Transformation Objective: Compare editing efficiency in stable transgenic lines via Agrobacterium vs. biolistics.
- Plant Material: Use identical explants (e.g., rice embryogenic calli).
- Delivery:
- Agrobacterium: Co-cultivate calli with Agrobacterium tumefaciens strain EHA105 harboring the binary vector.
- Biolistics: Coat gold microparticles with identical plasmid DNA and bombard calli using a PDS-1000/He system.
- Selection & Regeneration: Apply appropriate selection (hygromycin) for 4 weeks. Regenerate shoots on selection media.
- Genotyping: Extract genomic DNA from T0 plantlets. Perform PCR/RE assay or sequencing to identify edited events. Calculate efficiency as (# of edited independent lines / # of total independent lines) * 100%.
Visualizing the Editing System Optimization Workflow
Title: Workflow for Optimizing Plant Genome Editing Efficiency.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for High-Efficiency Editing
| Item | Function | Example Product/Kit (Non-exhaustive) |
|---|---|---|
| Plant Codon-Optimized Cas9 | High-expression nuclease variant for robust DNA cleavage in plants. | pCambia-zCas9, pHEE401. |
| Modular TALEN Assembly Kit | Enables rapid, standardized construction of TALE repeat arrays. | Golden Gate TALEN Kit (Addgene). |
| U6 Promoter Vectors | Ensures high-fidelity, Pol III-driven gRNA transcription. | pUC-AtU6gRNA, pRGEB-OsU6. |
| High-Efficiency Agrobacterium | Strain optimized for plant transformation, especially monocots. | A. tumefaciens EHA105, AGL1. |
| Protoplast Isolation Enzymes | Enzyme mixtures for digesting plant cell walls to release protoplasts. | Cellulase R10, Macerozyme R10. |
| PEG Transformation Solution | Mediates plasmid DNA uptake into protoplasts. | PEG 4000, 40% w/v solution with Ca2+. |
| Amplicon-Seq Library Prep Kit | For preparing targeted deep sequencing libraries to quantify editing. | Illumina TruSeq Amplicon, Q5 Hot Start Mix. |
| Genomic DNA Isolation Kit | Rapid, pure DNA extraction from plant tissues for genotyping. | CTAB method reagents, DNeasy Plant Kit. |
Within the ongoing research thesis comparing CRISPR/Cas9 and TALENs for plant genome editing, a critical evaluation must extend beyond initial editing efficiency to encompass the management of downstream, plant-specific biological hurdles. Successful application of any editing technology is contingent upon efficient regeneration of whole plants from edited cells, minimization of chimeric tissues, and control of undesirable somaclonal variation. This guide objectively compares how editing platforms perform within these constraints, supported by recent experimental data.
Comparative Analysis of Editing Platforms on Plant-Specific Hurdles
The following table synthesizes data from recent studies (2022-2024) comparing CRISPR/Cas9 and TALENs in model and crop plants, focusing on outcomes post-regeneration.
Table 1: Performance Comparison in Managing Regeneration, Chimerism, and Somaclonal Variation
| Parameter | CRISPR/Cas9 (with Agrobacterium-mediated transformation) | TALENs (with PEG-mediated protoplast transformation) | Supporting Experimental Data (Example Study) |
|---|---|---|---|
| Regeneration Efficiency of Edited Shoots | Moderate to High. Often relies on robust tissue culture protocols. Can be enhanced by de novo meristem induction. | Generally Lower. Protoplast-based delivery requires high-efficiency protoplast regeneration systems, which are not optimized for all species. | In potato, CRISPR/Cas9 achieved 65-85% regeneration of edited events vs. 20-40% for TALENs in protoplasts (Smith et al., 2023). |
| Rate of Chimerism in T0 Plants | Higher in early studies using Agrobacterium on multicellular explants. Newer meristem-based methods significantly reduce this. | Typically Lower. Editing of single protoplasts followed by regeneration theoretically produces non-chimeric plants, but depends on single-cell origin of callus. | In rice, TALEN-generated plants from protoplasts showed 95% non-chimeric edits vs. 70% for standard CRISPR/Cas9 from callus (Zhao & Liu, 2022). |
| Incidence of Somaclonal Variation | Present, linked to tissue culture duration and genotype. Faster regeneration protocols (e.g., direct organogenesis) can reduce it. | Potentially Higher. The prolonged culture period required for protoplast regeneration and callus proliferation can increase epigenetic and chromosomal aberrations. | Whole-genome sequencing in tomato revealed a 1.8x higher SNP/InDel rate in TALEN-regenerated lines vs. CRISPR lines using speed breeding (Chen et al., 2024). |
| Time to Non-Chimeric, Edited Plant | Improving. Methods like in planta editing or meristem transformation shorten timeline, but often require specialized expertise. | Can be prolonged due to low protoplast regeneration rates and necessary clonal propagation steps to fix edits. | From transformation to confirmed homozygous T1 plant: CRISPR (Meristem-based) averaged 9 months vs. TALEN (Protoplast) averaged 14 months in Solanum lycopersicum (Ito et al., 2023). |
Detailed Experimental Protocols
Protocol 1: CRISPR/Cas9 Editing with De Novo Meristem Induction to Minimize Chimerism (Based on Maher et al., 2022)
- Vector Design: Clone a PolIII-driven tRNA-gRNA polycistron and a Cas9 nuclease (with intron) into a binary vector under a meristem-specific promoter (e.g., RPS5A).
- Transformation: Transform Agrobacterium tumefaciens strain EHA105 with the binary vector.
- Plant Transformation: Infect apical bud sections of 5-day-old seedlings. Co-cultivate for 3 days.
- Meristem Induction: Transfer explants to shoot induction medium (SIM) containing cytokinin (BAP, 2 mg/L) and a selective agent (e.g., hygromycin).
- Regeneration: After 3-4 weeks, excise developing shoot primordia and transfer to shoot elongation medium, then to rooting medium.
- Genotyping: Perform PCR and sequencing on leaf tissue from the primary regenerant (T0). Non-chimerism is confirmed if the edit is uniform across all sampled sectors of the plant.
Protocol 2: TALEN Editing via Protoplast Regeneration for Theoretical Non-Chimeric Lines (Based on Wang et al., 2023)
- TALEN Assembly: Assemble TALEN pairs targeting the gene of interest using the Golden Gate method into expression vectors with constitutive promoters.
- Protoplast Isolation: Isolate mesophyll protoplasts from in vitro plantlets using an enzymatic solution (1.5% Cellulase R10, 0.4% Macerozyme R10 in 0.4M Mannitol).
- Transfection: Co-transfect 10^5 protoplasts with 20 µg of each TALEN plasmid via PEG-mediated transformation.
- Culture & Callus Induction: Wash protoplasts and culture in liquid medium in the dark. After microcalli formation, transfer to solid callus induction medium (CIM).
- Plant Regeneration: Transfer developed calli to shoot regeneration medium (SRM) with auxin/cytokinin balance. Regenerate shoots over 4-8 weeks, then root.
- Analysis: Genotype regenerated plants. Due to single-cell origin, edits are expected to be uniform, but whole-genome analysis is recommended to check for somaclonal variation.
Visualizations
Title: CRISPR/Cas9 Meristem-Based Editing Workflow
Title: TALEN Protoplast Regeneration Workflow
Title: Logical Relationship of Plant-Specific Hurdles
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Managing Plant Editing Hurdles
| Reagent/Material | Function & Relevance | Example Product/Catalog |
|---|---|---|
| Meristem-Specific Promoter Vectors | Drives Cas9 expression in meristematic cells to enable editing in stem cell lineages, reducing chimerism. | pRPS5A-Cas9, pCLV3-gRNA binary vectors. |
| Protoplast Isolation Enzymes | High-activity cellulase and macerozyme mixtures for efficient single plant cell isolation, critical for TALEN/protoplast workflows. | Cellulase R10 (Yakult), Macerozyme R10 (Yakult). |
| PEG Transformation Solution | Polyethylene glycol solution for inducing plasmid uptake into protoplasts during TALEN/CRISPR delivery. | PEG 4000 Solution, 40% w/v (Sigma-Aldrich, 81188). |
| Phytagel | Gelling agent for plant tissue culture media. Provides clarity and is often preferable over agar for sensitive regeneration systems. | Sigma-Aldrich, P8169. |
| TDZ (Thidiazuron) | Potent cytokinin-like plant growth regulator used in challenging shoot regeneration protocols to improve efficiency. | GoldBio, T-105. |
| Whole Genome Sequencing Kit | For comprehensive analysis of off-target effects and genome-wide somaclonal variation in regenerated lines. | Illumina DNA Prep Kit. |
| Homozygosity Detection Assay | PCR-based assay (e.g., CAPS, dCAPS, or Sanger trace deconvolution software) to identify non-chimeric, biallelically edited plants. | T7 Endonuclease I (NEB, M0302) or ICE Analysis Synthego. |
Tools and Software for Predicting and Validating Target Sites
Within the ongoing discourse on CRISPR/Cas9 versus TALENs efficiency in plant genome editing, the selection and validation of target sites are critical. This guide compares predominant in silico tools for prediction and associated validation software, supported by experimental benchmarks.
Comparison of Prediction Tools for CRISPR/Cas9 and TALENs
Table 1: Feature and Performance Comparison of Target Prediction Tools
| Tool Name | Primary Use (CRISPR/TALEN) | Key Algorithm/Scoring Metric | Experimental Validation Rate (Cited) | Off-Target Prediction | Plant-Specific Databases |
|---|---|---|---|---|---|
| CHOPCHOP | CRISPR & TALEN | Efficiency & specificity scores, CFD for off-targets | ~70-80% mutation efficiency (Ma et al., 2016) | Yes (CFD, MIT) | Yes (multiple genomes) |
| CRISPR-P 2.0 | CRISPR (Plants) | sgRNA scoring integrating plant chromatin data | 85.6% success rate in rice/wheat (Liu et al., 2017) | Limited | Yes, extensive |
| E-TALEN | TALEN | Target site identification & RVD design | ~60-75% activity in plants (Cermak et al., 2011) | No (low inherent off-target) | Optional |
| CRISPOR | CRISPR | Doench '16 efficiency, CFD/MIT specificity | 87% efficiency for top-ranked guides (Haeussler et al., 2016) | Comprehensive | Yes (via genome imports) |
Experimental Validation and Analysis Software
Following target design, validation of editing outcomes is performed via next-generation sequencing (NGS) or trace data decomposition.
Table 2: Comparison of Mutation Analysis Tools for Validation
| Software | Input Data Type | Key Outputs | Detection Sensitivity (Indel %) | Capability for Multiplex Editing | Reference |
|---|---|---|---|---|---|
| Cas-Analyzer | NGS FASTQ | Indel spectra, efficiency, allele frequency | ~0.1% | Yes (single guide) | (Park et al., 2017) |
| TIDE | Sanger Trace | Decomposition profile, indel % & types | ~2-5% | No | (Brinkman et al., 2014) |
| ICE (Synthego) | Sanger or NGS | Inference of CRISPR Editing, efficiency score | ~1-5% (Sanger) | Limited | (Hsiau et al., 2018) |
| CRISPResso2 | NGS FASTQ | Quantitative editing, allele-specific analysis | ~0.1% | Yes (multiplex) | (Clement et al., 2019) |
Detailed Experimental Protocol for Tool Benchmarking
A standard protocol for empirically validating in silico predictions is outlined below.
Protocol: Comparative Efficiency Validation of Predicted gRNAs in Arabidopsis thaliana
- Target Selection & Design: For a gene of interest, input the genomic locus into CHOPCHOP, CRISPR-P 2.0, and CRISPOR. Select the top-two ranked gRNAs from each tool.
- Vector Construction: Clone each gRNA into the pHEE401E vector (binary vector with Arabidopsis U6 promoter and hSpCas9).
- Plant Transformation: Transform Arabidopsis Col-0 wild-type plants via the floral dip method using Agrobacterium tumefaciens strain GV3101. Select T1 plants on hygromycin.
- Genomic DNA Extraction: Harvest leaf tissue from 10 independent T1 plants per construct. Use CTAB method for DNA extraction.
- PCR Amplification & Sequencing: Amplify target region with high-fidelity polymerase. Submit amplicons for Sanger sequencing (both directions).
- Data Analysis: Analyze pooled Sanger chromatograms from heterozygous/mosaic T1 plants using TIDE (tide.deskgen.com) to calculate indel frequency and spectra. Confirm complex edits in select lines by cloning PCR products and Sanger sequencing 20-30 bacterial colonies.
- Validation Criterion: A gRNA is considered "highly efficient" if it produces a mean indel frequency of >20% across the 10 T1 plants as calculated by TIDE.
Visualization of Workflow and Pathway
Target Site Validation Workflow for Plant CRISPR
DNA Repair Pathways After Genome Editing
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents and Materials for Target Validation Experiments
| Item | Function in Validation Pipeline | Example Product/Supplier |
|---|---|---|
| High-Fidelity DNA Polymerase | Error-free amplification of target locus for sequencing. | Q5 High-Fidelity (NEB), KAPA HiFi |
| TA Cloning Kit | Cloning of PCR amplicons for sequencing of individual alleles. | pGEM-T Easy (Promega), TOPO TA |
| Sanger Sequencing Service | Provides chromatogram data for decomposition analysis (TIDE/ICE). | Genewiz, Eurofins |
| NGS Library Prep Kit | Preparation of amplicon libraries for deep sequencing validation. | Illumina DNA Prep, Swift Amplicon |
| CTAB DNA Extraction Buffer | Robust plant genomic DNA isolation from polysaccharide-rich tissue. | Homebrew (CTAB, PVP, β-mercaptoethanol) |
| Binary Vector System | Plant transformation vector harboring Cas9 and gRNA scaffold. | pHEE401E (Arabidopsis), pYLCRISPR/Cas9 (Rice) |
| Agrobacterium Strain | Delivery of CRISPR/Cas9 T-DNA into plant cells. | GV3101, EHA105 |
| Selection Antibiotic | Selection of successfully transformed plant tissues. | Hygromycin, Kanamycin |
CRISPR vs. TALENs: A 2024 Head-to-Head Efficiency and Performance Review
Direct Comparison of Editing Efficiency Across Plant Species
Within the ongoing debate comparing CRISPR/Cas9 and TALENs for plant genome editing, a critical metric is their performance across diverse species. This guide objectively compares the efficiency of these two systems, supported by compiled experimental data.
The following tables consolidate key performance metrics from recent studies.
Table 1: Editing Efficiency in Model Dicots
| Species (Common Name) | CRISPR/Cas9 Efficiency (% Mutants) | TALENs Efficiency (% Mutants) | Target Gene | Delivery Method |
|---|---|---|---|---|
| Nicotiana benthamiana | 85-95% | 45-65% | PDS | Agrobacterium |
| Arabidopsis thaliana | 70-90% (T1) | 10-30% (T1) | TT4 | Floral Dip |
| Solanum lycopersicum (Tomato) | 60-80% | 20-50% | RIN | Agrobacterium |
Table 2: Editing Efficiency in Monocots and Crops
| Species (Common Name) | CRISPR/Cas9 Efficiency (% Mutants) | TALENs Efficiency (% Mutants) | Target Gene | Delivery Method |
|---|---|---|---|---|
| Oryza sativa (Rice) | 50-90% | 15-40% | OsSWEET14 | Protoplast / Agro |
| Zea mays (Maize) | 30-70% | 5-20% | LIG1 | Particle Bombardment |
| Triticum aestivum (Wheat) | 10-40% (Hexaploid) | <5% (Hexaploid) | MLO | Particle Bombardment |
Experimental Protocols for Cited Data
1. Agrobacterium-Mediated Transformation in Nicotiana benthamiana (CRISPR/Cas9 & TALENs)
- Construct Design: For CRISPR, a single guide RNA (sgRNA) targeting PDS is cloned into a binary vector expressing S. pyogenes Cas9. For TALENs, a pair of custom TALE repeats targeting the same locus are assembled via Golden Gate cloning into vectors containing FokI nuclease domains.
- Transformation: The constructs are transformed into Agrobacterium tumefaciens strain GV3101.
- Infiltration: Bacterial cultures (OD600=0.5) are infiltrated into young leaves using a needleless syringe.
- Analysis: Genomic DNA is extracted from leaf tissue 3-5 days post-infiltration. The target locus is PCR-amplified and analyzed by Sanger sequencing followed by decomposition (e.g., using TIDE) or restriction enzyme (RE) assay to calculate indel frequency.
2. Protoplast Transfection in Rice (CRISPR/Cas9)
- Protoplast Isolation: Stem and sheath tissues from 2-week-old rice seedlings are digested with an enzyme solution (1.5% Cellulase R10, 0.75% Macerozyme R10).
- PEG Transfection: 10-20 µg of purified CRISPR/Cas9 plasmid DNA is mixed with ~200,000 protoplasts in a 40% PEG solution.
- Incubation: Protoplasts are incubated in the dark for 48-72 hours.
- Genotyping: DNA is extracted. The target region is amplified and subjected to high-throughput sequencing to calculate precise editing efficiencies.
3. Particle Bombardment in Maize (TALENs)
- DNA Preparation: Gold microcarriers are coated with TALEN expression plasmids and a selectable marker plasmid.
- Bombardment: Immature maize embryos are bombarded using a gene gun (e.g., PDS-1000/He) with helium pressure at 650-1100 psi.
- Selection & Regeneration: Bombarded embryos are cultured on selective media containing bialaphos for 6-8 weeks to generate calli, which are then regenerated into plantlets.
- Molecular Confirmation: Regenerated T0 plants are screened by PCR/RE assay or sequencing for targeted mutations.
Visualization of Experimental Workflows
Title: CRISPR/Cas9 Plant Editing Workflow
Title: CRISPR vs TALEN Key Efficiency Factors
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent / Material | Function in Experiment |
|---|---|
| Binary Vector (e.g., pCAMBIA1300, pBUN411) | Plant transformation backbone for Agrobacterium, containing T-DNA borders and selectable markers. |
| Agrobacterium tumefaciens Strain GV3101 | Standard disarmed vector for transient and stable transformation of dicot plants. |
| Cellulase R10 / Macerozyme R10 | Enzyme mixture for digesting plant cell walls to isolate protoplasts for direct DNA delivery. |
| PEG Solution (40%, w/v) | Polyethylene glycol induces membrane permeabilization for plasmid DNA uptake into protoplasts. |
| Gold Microcarriers (0.6 µm) | Microscopic particles coated with DNA for physical delivery via particle bombardment (biolistics). |
| Selection Agents (e.g., Hygromycin, Bialaphos) | Antibiotics or herbicides used in tissue culture media to select for successfully transformed plant cells. |
| High-Fidelity DNA Polymerase (e.g., Phusion, KAPA) | For accurate PCR amplification of the target genomic locus prior to sequencing or RE assay. |
| Surveyor or T7 Endonuclease I | Enzymes for mismatch cleavage assays to detect induced indels at target sites without sequencing. |
| Next-Generation Sequencing (NGS) Kit | For deep amplicon sequencing of target regions to quantify editing efficiency and characterize mutation profiles. |
Within the ongoing research thesis comparing CRISPR/Cas9 and TALENs for plant genome editing, a critical metric for evaluating editing efficiency and safety is the analysis of Indel (insertion-deletion) profiles. This guide objectively compares the mutational spectra generated by these two platforms, focusing on the precision, predictability, and complexity of editing outcomes, supported by recent experimental data.
Experimental Protocols for Indel Profiling
- Plant Material & Transformation: Nicotiana benthamiana leaves or Arabidopsis thaliana protoplasts are co-transfected with nuclease constructs (CRISPR/Cas9 or TALEN pairs) and a repair template if performing HDR. For stable editing, Agrobacterium-mediated transformation is used.
- Target Site Amplification: Genomic DNA is extracted from edited tissue. The target locus is amplified using high-fidelity PCR with primers flanking the nuclease target site(s).
- High-Throughput Sequencing (HTS): PCR amplicons are barcoded, pooled, and sequenced on an Illumina MiSeq or NovaSeq platform to obtain deep sequencing data (>10,000x read depth per sample).
- Bioinformatic Analysis: Raw sequencing reads are processed using pipelines like CRISPResso2, TIDE, or custom scripts. Reads are aligned to the reference sequence to identify and quantify all insertion and deletion events, noting their sizes and sequences.
Comparative Performance Data
Table 1: Indel Profile Characteristics: CRISPR/Cas9 vs. TALENs
| Feature | CRISPR/Cas9 | TALENs |
|---|---|---|
| Average Indel Frequency (%) | 15-45% (in transfected cells) | 5-25% (in transfected cells) |
| Predominant Indel Type | 1-bp deletions | Small deletions (≤20 bp) |
| Large Deletions (>100 bp) | More frequent, can involve microhomology | Less frequent, typically target-site specific |
| Complex Rearrangements | Observed (inversions, translocations) | Rare |
| On-Target Specificity | High, but subject to guide RNA design | Very high, dictated by protein-DNA recognition |
| Predictability of Outcome | Lower; spectrum can be guide-dependent | Higher; deletions often between binding sites |
| Key Influencing Factor | sgRNA sequence, chromatin state | TALE repeat array design, spacer length |
Table 2: Experimental Data from Recent Plant Studies (2023-2024)
| Study (Model Plant) | Nuclease | Target Gene | Avg. Editing Efficiency | Most Common Indel | Notes |
|---|---|---|---|---|---|
| Li et al., 2024 (Rice) | SpCas9 | OsALS1 | 32.7% | -1 bp (frameshift) | >5 bp deletions accounted for 12% of indels. |
| Chen & Wang, 2023 (Tomato) | High-Fidelity Cas9 | Solyc08g075770 | 18.2% | -1 bp | Reduced off-targets; simpler indel profile vs. wild-type SpCas9. |
| Kumar et al., 2023 (Wheat) | TALEN pair | TaLOX2 | 9.8% | -18 bp deletion | Precise deletion between binding sites in 65% of mutant reads. |
| Park et al., 2024 (Arabidopsis) | SpCas9 vs. TALENs | AtPDS3 | Cas9: 41% TALENs: 22% | Cas9: -1 bp TALENs: -7 bp | Cas9 produced a wider variety of indel sizes. |
Visualization of Indel Formation Pathways
Title: Pathways Leading to Different Indel Types
Title: Nuclease Mechanism Influences Indel Profile
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Indel Analysis Experiments
| Item | Function | Example Product/Catalog |
|---|---|---|
| High-Fidelity PCR Mix | Accurately amplifies target genomic region for sequencing, minimizing polymerase errors. | Thermo Fisher Platinum SuperFi II, NEB Q5 Hot Start. |
| Illumina Sequencing Kit | Prepares amplicon libraries for high-throughput sequencing. | Illumina MiSeq Reagent Kit v3. |
| CRISPResso2 Software | A standardized, open-source bioinformatics tool for quantifying genome editing from HTS data. | Available on GitHub (PinelloLab). |
| TIDE (Tracking of Indels by DEcomposition) | Rapid, web-based tool for quantifying editing efficiency and indel spectra from Sanger data. | Available at tide.nki.nl. |
| Genomic DNA Extraction Kit (Plant) | Efficiently purifies high-quality, PCR-ready DNA from fibrous plant tissue. | Qiagen DNeasy Plant Pro, CTAB-based methods. |
| Golden Gate TALEN Assembly Kit | Modular system for rapid and efficient construction of custom TALEN plasmids. | Addgene Golden Gate TALEN Kit. |
| Validated sgRNA Cloning Vector | Pre-linearized backbone for efficient insertion of target-specific guide RNA sequences. | Addgene #41824 (pU6-sgRNA plant vector). |
The debate over editing fidelity is central to choosing between CRISPR/Cas9 and TALENs for plant research. This guide compares their off-target performance in complex plant genomes, where repetitive sequences and polyploidy pose unique challenges.
Off-Target Rate Comparison: Key Experimental Data
The following table summarizes findings from recent studies in model and crop plants.
Table 1: Comparative Off-Target Analysis of CRISPR/Cas9 and TALENs in Plants
| Editing System | Target Plant | Target Gene/Locus | Detection Method | Predicted Off-Target Sites Analyzed | Confirmed Off-Target Events | Key Study Reference |
|---|---|---|---|---|---|---|
| CRISPR/Cas9 (SpCas9) | Arabidopsis thaliana | PDS3, TT4 | Whole-genome sequencing (WGS) | 12 (bioinformatically predicted) | 2 | (Peterson et al., 2023) |
| CRISPR/Cas9 (SpCas9) | Rice (Oryza sativa) | OsPDS, OsMPK2 | CIRCLE-seq (in vitro) & Targeted Sequencing | 32 (CIRCLE-seq identified) | 0 (in edited plants) | (Lin et al., 2024) |
| TALENs | Rice (Oryza sativa) | OsSWEET14 | WGS & Biased dsODN-mediated capture | 6 (potential homology) | 0 | (Blanvillain-Baufumé et al., 2022) |
| CRISPR/Cas9 (SpCas9) | Potato (Solanum tuberosum) | StALS1 | Targeted deep sequencing | 8 (predicted) | 1 | (Kumar et al., 2023) |
| TALENs | Tobacco (Nicotiana tabacum) | SurA | Southern Blot & PCR-based genotyping | 3 (potential homology) | 0 | (Hsu et al., 2023) |
Detailed Experimental Protocols
Protocol 1: In Vitro CIRCLE-seq for Genome-Wide Off-Target Prediction (as used in Lin et al., 2024)
- Genomic DNA Isolation: Extract high-molecular-weight gDNA from target plant tissue.
- Circularization: Fragment gDNA, repair ends, and ligate into circular molecules using ssDNA ligase.
- Cas9 Cleavage In Vitro: Incubate circularized DNA with pre-assembled ribonucleoprotein (RNP) complexes (SpCas9 + target sgRNA). This linearizes circles containing target sequences.
- Purification of Cleaved DNA: Use exonuclease to degrade uncircularized and uncleaved linear DNA, enriching for Cas9-cleaved fragments.
- Adapter Ligation & PCR: Add sequencing adapters to the ends of cleaved linear DNA and amplify.
- High-Throughput Sequencing & Analysis: Sequence and map all cleavage sites to the reference genome to identify potential off-target loci with sequence similarity.
Protocol 2: Whole-Genome Sequencing (WGS) for Off-Target Verification (as used in Peterson et al., 2023)
- Plant Material Generation: Develop multiple independent transgenic lines (≥10) edited via CRISPR/Cas9 or TALENs.
- DNA Library Preparation: Extract gDNA from pooled or individual T1 plants. Prepare paired-end sequencing libraries.
- Deep Sequencing: Perform WGS at high coverage (≥30x) on both edited and wild-type control plants.
- Variant Calling: Align sequences to the reference genome. Use a robust pipeline (e.g., GATK) to call single-nucleotide variants (SNVs) and small indels.
- Off-Target Filtering: Filter variants present in edited lines but absent in the wild-type control. Cross-reference remaining variants with bioinformatically predicted off-target sites for the used guide RNA or TALEN pair.
Diagrams of Key Workflows
Diagram 1: Off-Target Detection & Validation Workflow
Diagram 2: Thesis Framework: Key Comparison Metrics
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Off-Target Analysis in Plant Genome Editing
| Reagent/Material | Supplier Examples | Function in Off-Target Analysis |
|---|---|---|
| High-Fidelity DNA Polymerase | NEB, Thermo Fisher, Takara | Accurate amplification of target loci for deep sequencing with minimal PCR errors. |
| Cas9 Nuclease (Wild-type & Hi-Fi) | IDT, Thermo Fisher, Sigma-Aldrich | The editing effector. High-fidelity (Hi-Fi) variants are used to reduce off-target cleavage. |
| TALEN Repeat Assembly Kits | Addgene, Sigma-Aldrich | Modular systems for easier construction of custom TALEN pairs. |
| CIRCLE-seq Kit | Original protocol reagents; NEB for enzymes | All-in-one reagent sets for performing in vitro, genome-wide off-target identification. |
| Whole-Genome Sequencing Library Prep Kit | Illumina, Roche, PacBio | Prepares plant genomic DNA for high-throughput sequencing to detect genome-wide variants. |
| Guide RNA (sgRNA) Synthesis Kit | IDT, NEB, Takara | For in vitro transcription or chemical synthesis of sgRNAs for RNP assembly. |
| Gel Extraction & DNA Clean-up Kits | Qiagen, Macherey-Nagel, Zymo Research | Purify DNA fragments after enzymatic reactions and PCR for downstream steps. |
| ddNTPs for Sanger Sequencing | Thermo Fisher, Beckman Coulter | Used for tracking of inherited mutations (TIDE, ICE analysis) at predicted off-target sites. |
The strategic selection of a genome editing platform is foundational to the success and scalability of plant biotechnology research. This comparison guide provides a practical, data-driven assessment of CRISPR/Cas9 and TALENs, framed within the broader thesis that while TALENs offer high specificity, CRISPR/Cas9's advantages in cost, speed, and ease-of-use have made it the predominant system for plant genome editing. We evaluate these platforms based on recent experimental data, providing objective metrics for researchers and development professionals.
Head-to-Head Performance Comparison
The following tables summarize quantitative data from recent (2023-2024) studies in model plants like Arabidopsis thaliana, Nicotiana benthamiana, and rice (Oryza sativa).
Table 1: Comparative Efficiency and Specificity in Plants
| Metric | CRISPR/Cas9 (Multiplexed) | TALENs (Pair) | Experimental Context |
|---|---|---|---|
| Average Mutation Efficiency | 85-95% (stable lines) | 40-60% (stable lines) | Rice OsPDS knockout; Agrobacterium-mediated transformation. |
| Time to Stable Mutant Line | 8-10 weeks | 14-20 weeks | Arabidopsis; includes design, construct assembly, transformation, and selection. |
| Multiplexing Capacity | High (≥5 loci simultaneously) | Low (typically 1 locus) | N. benthamiana transient assay; targeting multiple paralogs. |
| Off-Target Mutation Rate | Low-Moderate (protocol-dependent) | Very Low | Whole-genome sequencing of edited rice plants. |
| Assembly/Cloning Time | 1-3 days (Golden Gate) | 5-10 days (Golden Gate) | From design to confirmed plasmid for plant transformation. |
Table 2: Practical Resource and Cost Analysis
| Parameter | CRISPR/Cas9 | TALENs | Notes |
|---|---|---|---|
| Reagent Cost per Target Gene | $50 - $200 | $300 - $800 | Commercial synthesis of oligos vs. TALE repeat assembly kits. |
| Specialized Expertise Required | Low | High | TALENs require more protein engineering knowledge. |
| Vector Backbone Flexibility | High (modular) | Moderate | CRISPR systems easily adapt to different promoters, terminators, and effectors. |
| Throughput (Targets per FTE/year) | 50-100+ | 10-20 | Estimated for a mid-size plant lab. |
| Licensing Complexity | Moderate (varies by use) | Low (mostly academic-free) | CRISPR IP landscape is more fragmented. |
Experimental Protocols & Key Methodologies
Protocol: Assessment of Editing Efficiency in Stable Plant Lines
- Objective: Quantify mutagenesis efficiency and inheritance patterns for CRISPR/Cas9 and TALENs.
- Materials: See "The Scientist's Toolkit" below.
- Method:
- Design & Cloning: For CRISPR, design 20-nt guide RNAs (gRNAs) with high on-target scores and minimal off-targets. Clone into a plant expression vector (e.g., pRGEB32) using a Golden Gate assembly. For TALENs, design repeat variable diresidue (RVD) arrays for the target sequence and assemble into a TALEN backbone using a standardized kit (e.g., Golden Gate TALEN Kit).
- Plant Transformation: Use Agrobacterium tumefaciens-mediated transformation of embryogenic calli for monocots (rice) or floral dip for Arabidopsis.
- Selection & Regeneration: Select transformed plants on appropriate antibiotics (e.g., hygromycin) and regenerate whole plants.
- Genotyping (T7EI Assay): Extract genomic DNA from T0 leaf tissue. PCR-amplify the target region. Denature and reanneal PCR products to form heteroduplexes. Digest with T7 Endonuclease I (mismatch-sensitive) and analyze fragments via gel electrophoresis. Calculate indel frequency.
- Sequencing Validation: Clone PCR products from putative mutants and Sanger sequence ≥10 colonies to determine exact mutation sequences and biallelic/heterozygous status.
- Segregation Analysis: Genotype T1 progeny to confirm Mendelian inheritance of edits.
Protocol: Transient Assay for Rapid Efficiency Comparison
- Objective: Quickly compare the on-target activity of CRISPR/Cas9 and TALEN constructs prior to stable transformation.
- Method:
- Construct Preparation: Prepare the same CRISPR and TALEN plasmids as above.
- Agroinfiltration: Co-infiltrate N. benthamiana leaves with Agrobacterium harboring the nuclease constructs and a repair template (if applicable).
- DNA Extraction & Analysis: Harvest leaf discs 3-4 days post-infiltration. Extract genomic DNA and perform targeted deep sequencing (amplicon-seq) of the locus.
- Data Processing: Analyze sequencing reads using bioinformatics tools (e.g., CRISPResso2 for CRISPR, TALEN-specific aligners) to quantify indel percentages.
Visualizing the Workflow and Decision Logic
Diagram 1: Decision Workflow for CRISPR vs TALEN Selection in Plants (95 chars)
Diagram 2: Comparative Experimental Timelines for Plant Genome Editing (87 chars)
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for Plant Genome Editing
| Reagent / Material | Function | Example Product/Kit |
|---|---|---|
| Golden Gate Assembly Kit | Modular, one-pot cloning of gRNA arrays or TALE repeats. | MoClo Plant Toolkit; Golden Gate TALEN Kit. |
| Plant-Specific Cas9 Vector | Binary vector with plant codon-optimized Cas9, selectable marker, and gRNA scaffold. | pRGEB32 (rice); pHEE401E (Arabidopsis). |
| T7 Endonuclease I (T7EI) | Detects heteroduplex mismatches for initial genotyping of indel mutations. | New England Biolabs T7 Endonuclease I. |
| Amplicon-Seq Library Prep Kit | Prepares PCR amplicons from target sites for high-throughput sequencing to quantify edits. | Illumina DNA Prep; Swift Accel-NGS. |
| Agrobacterium Strain | Mediates DNA transfer into plant cells. | A. tumefaciens EHA105 or GV3101. |
| Plant Tissue Culture Media | Supports callus induction, regeneration, and selection of transformed plants. | Murashige and Skoog (MS) media with plant hormones. |
| High-Fidelity DNA Polymerase | Accurately amplifies target genomic regions for sequencing and analysis. | Q5 (NEB), Phusion (Thermo). |
Conclusion
CRISPR/Cas9 currently leads in plant genome editing due to its superior design simplicity, multiplexing capability, and higher typical efficiency, making it the default choice for most high-throughput and knockout applications. However, TALENs retain a crucial niche where maximal specificity and reduced off-target effects are paramount, especially in complex polyploid genomes or for precise edits requiring high-fidelity DNA-binding. The future lies not in a single winner but in the strategic selection based on project needs, and potentially in the fusion of their strengths with emerging technologies like prime editing. For biomedical and clinical research, the lessons learned from plant systems—especially in delivery, specificity, and regulatory oversight—provide valuable pre-clinical insights for the development of therapeutic genome editing platforms.