Precision Plant Breeding: Comparing Base Editing and Traditional CRISPR-Cas9 for Crop Improvement
This article provides a comparative analysis of base editing and traditional CRISPR-Cas9 systems for crop genome editing, tailored for researchers, scientists, and drug development professionals.
Precision Plant Breeding: Comparing Base Editing and Traditional CRISPR-Cas9 for Crop Improvement
Abstract
This article provides a comparative analysis of base editing and traditional CRISPR-Cas9 systems for crop genome editing, tailored for researchers, scientists, and drug development professionals. It explores the foundational principles, mechanism of action, and key differences between these technologies. The content delves into specific methodologies, applications in enhancing yield and stress tolerance, and protocols for targeted crop gene modification. It addresses critical challenges such as off-target effects, efficiency optimization, and delivery barriers. Finally, the article offers a rigorous comparative framework covering precision, scope, regulatory considerations, and validation metrics to inform technology selection and strategic research planning in agricultural biotechnology.
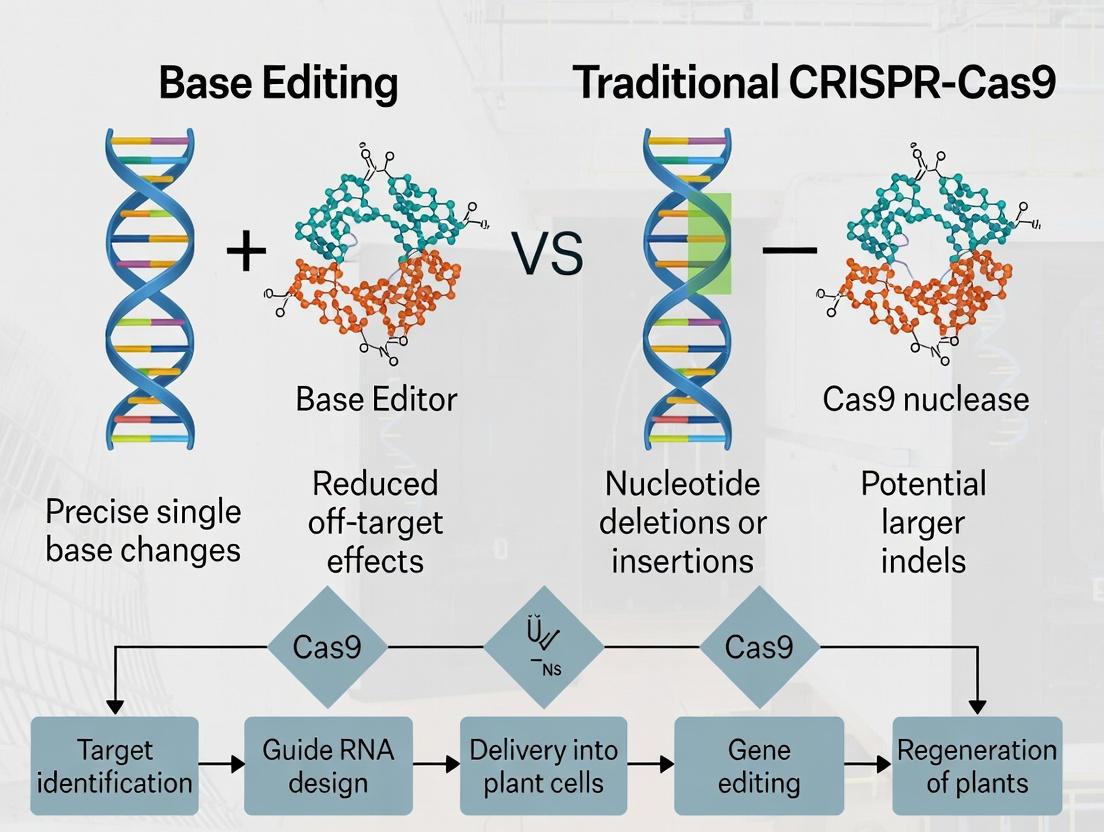
Understanding the Core Mechanisms: How Base Editors and CRISPR-Cas9 Differ at the Molecular Level
Within the context of crop research, the choice between traditional CRISPR-Cas9 nucleases and modern base editors represents a pivotal strategic decision. This whitepaper provides an in-depth technical comparison of these molecular tool architectures, focusing on their mechanisms, applications, and experimental implementation for precise genetic modification in plants.
Core Mechanisms and Architectures
CRISPR-Cas9 Nuclease System
The canonical Streptococcus pyogenes CRISPR-Cas9 system functions as a programmable DNA endonuclease. The guide RNA (gRNA) directs the Cas9 protein to a specific genomic locus via Watson-Crick base pairing, where Cas9 induces a double-strand break (DSB). In crops, repair of this break through error-prone non-homologous end joining (NHEJ) leads to gene knockouts, while homology-directed repair (HDR) can facilitate precise gene insertion or correction, though with low efficiency in plants.
Base Editor Architectures
Base editors are fusion proteins that combine a catalytically impaired Cas9 (nickase or dead) with a nucleobase deaminase enzyme. They enable direct, irreversible chemical conversion of one base pair to another without requiring a DSB or donor DNA template. Two primary classes are used in crops:
- Cytosine Base Editors (CBEs): Convert C•G to T•A.
- Adenine Base Editors (ABEs): Convert A•T to G•C. Recent advances include Dual Base Editors and Glycosylase Base Editors (GBEs) that expand editable sequences and enable C-to-G transversions.
Comparative Quantitative Analysis
Table 1: Core Performance Metrics in Crop Systems
| Parameter | CRISPR-Cas9 Nuclease | Cytosine Base Editor (CBE) | Adenine Base Editor (ABE) |
|---|---|---|---|
| Primary Edit | Indels (NHEJ), Precise edits (HDR) | C•G to T•A | A•T to G•C |
| Typical Efficiency in Plants | 0.5-20% (HDR: <1-5%) | 1-50% (avg. 10-30%) | 1-40% (avg. 5-20%) |
| DSB Formation | Yes | No | No |
| Off-target (DNA) Frequency | Moderate to High | Generally Lower | Generally Lower |
| Product Purity (%) | Low for HDR, Mixed for NHEJ | High (>99% desired base change in many studies) | High (>99% desired base change) |
| Key Limitation | Low HDR efficiency, PDSB toxicity | Restricted to C within activity window (protospacer positions 4-8, ~5nt window) | Restricted to A within activity window (protospacer positions 4-8, ~5nt window) |
| Common Applications in Crops | Gene knockouts, Large deletions, Gene insertion (complex) | Point mutation correction, Creating herbicide resistance alleles, Inducing premature stop codons | Point mutation correction, Creating gain-of-function alleles |
Table 2: Analysis of Edit Outcomes in a Model Crop Study (Rice OsALS Gene)
| Editing Tool | Target Base | Edit Efficiency (%) | Desired Product (Herbicide Res.) Purity (%) | Unwanted Indel Frequency (%) |
|---|---|---|---|---|
| Cas9-HDR | C to T (P171) | 2.1 | 31 | 65 |
| CBE (rAPOBEC1-nCas9) | C to T (P171) | 41.7 | 98.2 | 1.1 |
| ABE (TadA-nCas9) | A to G (W548) | 22.5 | 99.5 | 0.3 |
Detailed Experimental Protocols
Protocol 1: Agrobacterium-Mediated Transformation for CRISPR-Cas9 Editing in Rice
Objective: Generate stable knockout mutant lines.
- Vector Construction: Clone gene-specific gRNA(s) into a binary vector (e.g., pRGEB32) expressing Cas9 and the gRNA under Ubi and U6 promoters, respectively.
- Plant Material: Use embryogenic calli derived from mature seeds of Oryza sativa spp. japonica.
- Agrobacterium Transformation: Transform Agrobacterium tumefaciens strain EHA105 with the binary vector via electroporation.
- Co-cultivation: Infect rice calli with Agrobacterium suspension (OD600=0.8-1.0) for 15-30 minutes, then co-cultivate on solid N6 medium for 3 days at 25°C.
- Selection & Regeneration: Transfer calli to selection medium containing hygromycin (50 mg/L) and cefotaxime (250 mg/L) for 3-4 weeks. Regenerate shoots on regeneration medium with the same antibiotics.
- Genotyping: Extract genomic DNA from T0 plantlets. Perform PCR on target locus and sequence amplicons (Sanger or NGS) to detect indels.
Protocol 2: Protoplast Transfection for Rapid Validation of Base Editors in Wheat
Objective: Quantify base editing efficiency and profile prior to stable transformation.
- Plasmid Preparation: Purify high-quality plasmid DNA encoding the base editor (e.g., BE3 for CBE) and the specific gRNA.
- Protoplast Isolation: Isolate mesophyll protoplasts from 10-day-old wheat seedling leaves using enzymatic digestion (1.5% Cellulase R10, 0.75% Macerozyme R10 in 0.6M mannitol) for 6 hours in the dark.
- PEG-Mediated Transfection: Mix 10 µg of total plasmid DNA (molar ratio 1:1 of editor:gRNA) with 200 µL of protoplast suspension (10^5 cells). Add an equal volume of 40% PEG4000 solution, incubate for 15 minutes.
- Culture & Harvest: Dilute with W5 solution, culture in the dark at 25°C for 48-72 hours.
- DNA Extraction & Analysis: Harvest protoplasts, extract genomic DNA. Amplify target locus by PCR and submit for high-throughput amplicon sequencing (Illumina MiSeq). Analyze sequencing data with tools like CRISPResso2 or BE-Analyzer to calculate editing efficiency and byproduct spectra.
Key Signaling and Workflow Diagrams
Title: CRISPR-Cas9 Gene Editing Workflow in Crops
Title: Base Editor Mechanism Without Double-Strand Breaks
Title: Decision Logic for Selecting CRISPR Tools in Crop Research
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for CRISPR/Cas and Base Editing in Plants
| Reagent / Material | Function in Experiments | Key Consideration for Crops |
|---|---|---|
| Binary Vectors (e.g., pRGEB, pYLCRISPR) | Agrobacterium T-DNA vectors for stable plant transformation. Contain plant promoters (Ubi, U6), Cas9/BE, and gRNA scaffold. | Must be compatible with the specific crop (monocot/dicot) and transformation method. |
| Cas9 & Base Editor Plasmids | Source of nuclease or editor coding sequence. Common: SpCas9, BE4max, ABE8e. | Codon optimization for the host plant species is critical for high expression. |
| gRNA Cloning Kits | Streamline the insertion of target-specific 20nt sequences into expression vectors. | Golden Gate or BsaI-based modular systems allow multiplexing of several gRNAs. |
| Agrobacterium Strains (EHA105, GV3101) | Mediate DNA delivery into plant genomes via T-DNA transfer. | Strain choice affects transformation efficiency in different species (e.g., EHA105 for rice). |
| Protoplast Isolation Enzymes (Cellulase, Macerozyme) | Digest plant cell walls to release intact protoplasts for transient transfection assays. | Enzyme concentration and incubation time must be optimized for each plant tissue. |
| PEG Transformation Solution (PEG4000) | Facilitates plasmid uptake into protoplasts during transient transfection for rapid testing. | High-purity PEG is essential for high transfection efficiency and low cytotoxicity. |
| Next-Generation Sequencing (NGS) Kits | For deep sequencing of amplicons to quantify editing efficiency, purity, and off-targets. | Amplicon-EZ or similar services are standard for analyzing editing outcomes from bulk tissue. |
| HDR Donor Templates | Single-stranded oligodeoxynucleotides (ssODNs) or double-stranded DNA for precise edits with Cas9-HDR. | For plants, long ssODNs or plasmid-based donors with long homologies are often used, though efficiency remains low. |
Within the context of modern crops research, the debate between traditional CRISPR-Cas9 and base editing hinges on the fundamental mechanism of DNA alteration. This whitepaper details the double-strand break (DSB) paradigm, the canonical pathway through which traditional CRISPR-Cas9 induces genetic changes. For researchers engineering crop traits—such as herbicide tolerance, disease resistance, or improved yield—understanding the precision, efficiency, and potential unintended consequences of DSB repair is paramount when selecting a genome-editing tool.
The Core Mechanism: DSB Induction and Repair Pathways
The CRISPR-Cas9 system from Streptococcus pyogenes induces a targeted DSB, a severe form of DNA damage. The cellular response to this break dictates the genetic outcome. Two primary, competing endogenous repair pathways resolve DSBs: Non-Homologous End Joining (NHEJ) and Homology-Directed Repair (HDR).
Non-Homologous End Joining (NHEJ)
NHEJ is an error-prone, dominant pathway in most plant cells, particularly in non-dividing cells. It directly ligates the broken DNA ends, often resulting in small insertions or deletions (indels) at the junction. This is the primary mechanism for generating gene knockouts in crops.
Key Steps:
- Recognition: The Ku70/Ku80 heterodimer binds the DSB ends.
- End Processing: Nucleases (e.g., Artemis) may trim the DNA ends, creating microhomologies.
- Ligation: DNA Ligase IV/XRCC4 complex seals the break.
Homology-Directed Repair (HDR)
HDR is a precise, template-dependent pathway active primarily in the S/G2 phases of the cell cycle. It requires a donor DNA template with homology arms flanking the target site to introduce specific edits (e.g., point mutations, gene insertions).
Key Steps:
- Resection: The MRN complex (Mre11-Rad50-Nbs1) resects 5' ends to generate 3' single-stranded DNA overhangs.
- Strand Invasion: The recombinase RAD51 facilitates invasion of the homologous donor template by the single-stranded DNA.
- Synthesis & Resolution: DNA polymerase extends the invading strand using the donor sequence, and the intermediate is resolved, incorporating the desired edit.
Visualizing the DSB Repair Pathway Decision Logic
Diagram Title: DSB Repair Pathway Decision Logic
Key Experimental Protocol: Analyzing CRISPR-Cas9 Editing Outcomes in Crop Protoplasts
This protocol is used to assess the efficiency and spectrum of edits (indels) generated via the NHEJ pathway.
Materials:
- Target crop protoplasts
- Plasmid DNA encoding Cas9 and sgRNA or pre-formed Cas9-gRNA RNP complexes
- PEG transformation solution
- Nuclei lysis buffer
- PCR primers flanking the target genomic locus
- High-fidelity DNA polymerase
- Gel electrophoresis and purification kit
- T7 Endonuclease I (T7EI) or sequencing analysis software.
Procedure:
- Design & Delivery: Design sgRNA targeting the gene of interest. Introduce CRISPR-Cas9 components into isolated protoplasts via PEG-mediated transfection.
- Incubation: Incubate protoplasts for 48-72 hours to allow for editing and repair.
- Genomic DNA Extraction: Harvest protoplasts and extract genomic DNA.
- PCR Amplification: Amplify the target region from treated and control samples.
- Heteroduplex Formation: Denature and re-anneal PCR products to form heteroduplexes between wild-type and edited strands.
- Mismatch Detection:
- T7EI Assay: Digest heteroduplexed DNA with T7EI, which cleaves mismatched sites. Analyze fragments by gel electrophoresis. Indel frequency is estimated from band intensities.
- Sequencing Analysis: Clone PCR products or perform deep amplicon sequencing. Align sequences to the reference to identify exact indel sequences and their frequencies.
- Data Analysis: Quantify editing efficiency (% of indels) and characterize the spectrum of mutations (distribution of insertion vs. deletion sizes).
Quantitative Data: DSB Repair Outcomes in Model Crops
Table 1: Typical NHEJ-Mediated Editing Efficiencies and Indel Spectra in Crops Using Traditional CRISPR-Cas9
| Crop Species | Target Tissue | Delivery Method | Average Indel Efficiency (%) | Predominant Indel Type (Size Range) | HDR Efficiency (with donor) (%) | Reference Key Findings |
|---|---|---|---|---|---|---|
| Oryza sativa (Rice) | Protoplasts | RNP Transfection | 15-40% | 1-bp deletions (-1) | 0.5-2% | NHEJ highly active; HDR very inefficient in the absence of cell cycle synchronization. |
| Zea mays (Maize) | Immature Embryos | Agrobacterium | 10-30% | Small deletions (-1 to -10 bp) | <1% | Complex, multi-bp deletions common. HDR requires specialized donor design. |
| Nicotiana benthamiana | Leaf Disks | Agrobacterium (Transient) | 20-60% | Mixture of insertions & deletions | ~5% (transient) | High transient expression enables measurable HDR in somatic cells. |
| Triticum aestivum (Wheat) | Callus Cells | Biolistics | 5-25% | Large deletions (>20 bp) | Extremely low | Polyploid genome complexity can lead to large chromosomal deletions at DSB sites. |
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for DSB-Paradigm CRISPR-Cas9 Experiments in Crops
| Reagent / Material | Function & Role in DSB Paradigm | Example Supplier / Note |
|---|---|---|
| High-Efficiency Cas9 Expression Vector | Drives constitutive or tissue-specific expression of the Cas9 nuclease to create the DSB. | pCambia-Cas9, pUC-Cas9 vectors optimized for monocots/dicots. |
| sgRNA Cloning Kit | Streamlines the cloning of target-specific guide RNA sequences into expression vectors. | Commercial Golden Gate or BsaI-based assembly kits. |
| Cas9-gRNA Ribonucleoprotein (RNP) Complex | Pre-formed, purified Cas9 protein + sgRNA. Enables rapid, transient DSB induction without DNA integration, reducing off-target potential. | Synthesized in-lab or purchased from protein production services. |
| T7 Endonuclease I (T7EI) | Detection reagent for identifying mismatches in heteroduplex DNA, used to quantify NHEJ-derived indel efficiency. | Available from multiple molecular biology suppliers (NEB, Thermo Fisher). |
| Surveyor Nuclease (Cel I) | Alternative to T7EI for mismatch detection, often used for validation. | |
| Homology-Directed Repair (HDR) Donor Template | Single-stranded oligodeoxynucleotide (ssODN) or double-stranded DNA (dsDNA) with homologous arms and desired edit. Essential for precise editing via HDR. | Ultramer oligonucleotides or cloned plasmid donors. |
| Next-Generation Sequencing (NGS) Library Prep Kit for Amplicons | Enables deep sequencing of target loci to comprehensively characterize the full spectrum of NHEJ/HDR outcomes at nucleotide resolution. | Illumina TruSeq, Nextera Flex kits. |
| Protoplast Isolation & Transfection Kit | System for delivering CRISPR components into plant cells without the use of Agrobacterium, useful for rapid efficacy testing. | Cellulase & pectinase enzyme mixes, PEG-based transfection solutions. |
This whitepaper details the chemical conversion approach of base editing, a precision genome editing technology that enables direct, irreversible conversion of one target DNA base pair to another without generating double-strand breaks (DSBs). Framed within the broader thesis comparing base editing to traditional CRISPR-Cas9 in crop research, this method offers a paradigm shift by minimizing unintended mutagenic outcomes and enabling precise single-nucleotide polymorphisms (SNPs) critical for crop trait enhancement. For researchers and drug development professionals, this guide provides an in-depth technical examination of the core machinery, quantitative performance, and detailed protocols.
Core Mechanism & Architecture
Base editors (BEs) are fusion proteins comprising a catalytically impaired CRISPR-Cas protein (nCas9 or dCas9) tethered to a nucleobase deaminase enzyme. The system is guided by a single guide RNA (sgRNA) to a target genomic locus. Upon binding, the deaminase domain acts on a single-stranded DNA bubble within the R-loop, chemically converting a specific base (e.g., cytidine to uridine, or adenosine to inosine) within a narrow editing window. This edit is then processed by cellular DNA repair mechanisms or replication to permanently install the desired point mutation.
Diagram 1: Cytidine Base Editor Mechanism
Quantitative Performance Data
Base editing outcomes are quantified by editing efficiency (percentage of target alleles modified), product purity (percentage of desired product among edited alleles), and indel frequency. The following tables summarize key performance metrics from recent studies in plant and mammalian systems.
Table 1: Comparison of Major Base Editor Systems
| Base Editor System | Core Deaminase | Target Conversion | Theoretical Editing Window | Typical Efficiency in Plants (Range) | Indel Rate (%) |
|---|---|---|---|---|---|
| BE1/BE2 | rAPOBEC1 | C•G to T•A | ~5 nucleotides (positions 4-8) | 1-10% | <0.1 |
| BE3/BE4 | rAPOBEC1 | C•G to T•A | ~5 nucleotides (positions 4-8) | 10-50% | 0.1-1.0 |
| BE4max | rAPOBEC1 | C•G to T•A | ~5 nucleotides (positions 4-8) | Up to 70% | ~0.1 |
| AID-A variants | PmCDA1, AID | C•G to T•A | ~5 nucleotides | 5-40% | <1.0 |
| ABE7.10 | TadA*7.10 | A•T to G•C | ~5 nucleotides (positions 4-8) | 10-50% | <0.1 |
| ABE8e | TadA*8e | A•T to G•C | ~5 nucleotides (positions 4-8) | Up to 80% | <0.1 |
| CGBE | rAPOBEC1 + UGI | C•G to G•C | ~5 nucleotides | 5-30% | 1-5 |
Table 2: Base Editing vs. Traditional CRISPR-Cas9 HDR in Crops
| Parameter | Base Editing | Traditional CRISPR-Cas9 + HDR |
|---|---|---|
| Primary Mechanism | Chemical deamination & repair | DSB, HDR with donor template |
| Efficiency for Point Mutations | High (often >30%) | Very Low (<5% in plants) |
| Product Purity | High (often >90% of edits are desired) | Low (frequent indels from NHEJ) |
| DSB Formation | No | Yes (required) |
| Donor DNA Template Required | No | Yes |
| Applicability in Non-dividing Cells | Yes | Limited |
| Major Byproducts | Unintended base edits within window | Indels (from NHEJ) |
| Ideal Use Case | Installing known SNPs (e.g., herbicide resistance), correcting point mutations | Large insertions, deletions, or transgene integration |
Detailed Experimental Protocol: Plant Protoplast Base Editing
This protocol outlines the steps for assessing base editor activity in plant protoplasts, a common preliminary assay.
Materials:
- Plant expression vectors for base editor (BE) and sgRNA.
- Target plant seedlings (e.g., Arabidopsis, rice, tobacco).
- Enzymes for cell wall digestion (Cellulase R-10, Macerozyme R-10).
- Mannitol solution (0.6 M).
- PEG solution (40% PEG 4000, 0.2 M mannitol, 0.1 M CaCl2).
- MMg solution (0.4 M mannitol, 15 mM MgCl2, 5 mM MES, pH 5.7).
- WI solution (0.5 M mannitol, 20 mM KCl, 4 mM MES, pH 5.7).
- DNA extraction kit.
- PCR reagents and primers flanking target site.
- Sequencing platform (Sanger or NGS).
Procedure:
Vector Construction:
- Clone the sequence encoding the desired BE (e.g., BE3, ABE) into a plant expression vector with a strong constitutive promoter (e.g., CaMV 35S, ZmUbi).
- Clone the target-specific sgRNA sequence into a compatible expression vector under a Pol III promoter (e.g., AtU6).
Protoplast Isolation (for leaf tissue):
- Slice 1-2g of young leaf tissue into 0.5-1mm strips.
- Incubate strips in 20ml of enzyme solution (1.5% Cellulase R-10, 0.4% Macerozyme R-10, 0.4M mannitol, 20mM KCl, 20mM MES pH 5.7, 10mM CaCl2, 0.1% BSA) for 4-6 hours in the dark with gentle shaking.
- Filter the digest through a 70μm nylon mesh into a 50ml tube.
- Centrifuge at 100xg for 3 minutes to pellet protoplasts. Gently resuspend in 10ml of W5 solution (154mM NaCl, 125mM CaCl2, 5mM KCl, 5mM glucose, 2mM MES pH 5.7). Centrifuge again.
- Resuspend the final pellet in MMg solution. Count protoplast density (aim for ~2x10^5/ml).
PEG-Mediated Transfection:
- Aliquot 100μl of protoplast suspension (2x10^4 cells) into a 2ml tube.
- Add 10μg of total plasmid DNA (typically a 1:1 molar ratio of BE and sgRNA vectors).
- Add an equal volume (110μl) of PEG solution. Mix gently by inversion.
- Incubate at room temperature for 15-20 minutes.
- Gradually dilute with 1ml of WI solution, mixing gently.
- Centrifuge at 100xg for 3 minutes. Remove supernatant.
- Resuspend in 1ml of WI solution and transfer to a multi-well plate. Incubate in the dark at 22-25°C for 48-72 hours.
Genomic DNA Extraction & Analysis:
- Harvest protoplasts by centrifugation. Extract genomic DNA using a standard kit.
- Perform PCR amplification of the target genomic region.
- Quantification: Use Sanger sequencing followed by decomposition trace analysis (e.g., using EditR or BEAT) or high-throughput sequencing (amplicon-seq) to calculate editing efficiency and identify byproducts.
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Material | Provider Examples | Function in Base Editing Experiments |
|---|---|---|
| Base Editor Plasmids | Addgene (BE3, BE4, ABE7.10, ABE8e), commercial agro kits | Source of the base editor fusion protein gene for expression in target cells. |
| sgRNA Cloning Kits | ToolGen, Synthego, IDT | Streamlined systems for cloning custom sgRNA sequences into expression vectors. |
| PCR Cloning Kits (Golden Gate, Gibson) | NEB, Takara Bio | For assembling multi-component genetic constructs, such as fusing deaminase variants to Cas proteins. |
| Plant Protoplast Isolation Kits | Sigma-Aldrich, homemade preparations | Provide purified plant cells for rapid transient transfection and editing efficiency testing. |
| PEG Transfection Reagents | Sigma-Aldrich (PEG 4000) | Facilitates plasmid DNA uptake into protoplasts. |
| Next-Generation Sequencing Kits | Illumina (MiSeq), PacBio | For deep sequencing of target amplicons to precisely quantify editing efficiency, identify byproducts, and assess off-target effects. |
| EditR Software / BE-Analyzer | Open source, lab-developed | Computational tools for analyzing Sanger sequencing chromatograms to calculate base editing efficiency. |
| Cell-Penetrating Peptide (CPP) Conjugates | Commercial synthesizers | For delivering base editor ribonucleoprotein (RNP) complexes into cells, reducing off-target DNA exposure. |
Pathway & Workflow Visualization
Diagram 2: Crop Base Editing Workflow
Diagram 3: DNA Repair Pathway Comparison
Deaminase-mediated base editing represents a precise, efficient, and DSB-free alternative to traditional CRISPR-Cas9 for installing point mutations in crops. Its superior efficiency and product purity for SNP introduction accelerate functional genomics and the development of crops with improved yield, resilience, and nutritional quality. While challenges such as off-target editing and sequence context limitations persist, ongoing engineering of deaminases, Cas variants, and delivery methods continues to expand the capabilities of this transformative technology within the plant biotech toolkit.
Within the accelerating field of crop genome engineering, the debate between base editing and traditional CRISPR-Cas9 hinges on the precise function and configuration of core molecular components. Traditional CRISPR-Cas9 relies on creating double-strand breaks (DSBs), which are repaired by error-prone non-homologous end joining (NHEJ), often leading to unpredictable indels. Base editing, in contrast, directly converts one target DNA base pair to another without a DSB, enabling precise point mutations critical for crop improvement. This technical guide provides an in-depth comparison of the three core components—guide RNA, Cas protein variants, and enzyme fusions—that define the efficacy, specificity, and applicability of these systems in plant research.
Guide RNA: Specificity and Scaffolding
The guide RNA (gRNA) is the targeting module, consisting of a CRISPR RNA (crRNA) sequence that base-pairs with the target DNA and a scaffold that binds the Cas protein. In base editors, the gRNA must position the enzyme fusion complex with sub-nanometer precision adjacent to the target base (within a defined "activity window").
Key Design Parameters:
- Spacer Sequence (crRNA): 20-nt length is standard; requires a protospacer adjacent motif (PAM) immediately downstream for Cas9 binding.
- Scaffold: Often engineered for enhanced stability in planta.
- Activity Window Positioning: The target base must reside within a narrow window (typically positions 4-8 for cytosine base editors, CBE; positions 4-10 for adenine base editors, ABE) counting from the PAM-distal end.
Table 1: gRNA Design Considerations for Crop Systems
| Parameter | Traditional CRISPR-Cas9 | Base Editing (CBE/ABE) | Rationale for Difference |
|---|---|---|---|
| Target Site | Any base within spacer | Bases within a specific "activity window" | Deaminase enzyme must have steric access to the target base. |
| PAM Requirement | Strict (e.g., NGG for SpCas9) | Strict, but more restrictive | PAM defines the location of the activity window relative to the target base. |
| Off-target Concern | High; tolerates mismatches, especially distal to PAM | Generally lower for DNA deamination, but RNA off-targets possible | Cas9 variants with higher fidelity are often employed. |
| Delivery in Crops | Often as a Pol III-transcribed unit from a U6/U3 promoter. Codon-optimization essential. | Same as traditional, but expression levels critical for balancing efficiency vs. off-target effects. | Plant codon optimization and promoter choice affect complex formation. |
Cas Protein Variants: PAM Expansion and Fidelity
The Cas protein confers DNA binding and cleavage. Variants are engineered to alter PAM specificity, reduce off-target effects, or eliminate nuclease activity for base editing.
Table 2: Key Cas Protein Variants in Crop Research
| Variant | Parent | Key Feature | Primary Application | Example PAM |
|---|---|---|---|---|
| SpCas9 | S. pyogenes Cas9 | Standard nuclease | Traditional KO, DSB-mediated HDR | NGG |
| SpCas9-HF1 | SpCas9 | High-fidelity; reduced non-specific DNA contacts | Traditional KO with lower off-targets | NGG |
| SpCas9-NG | SpCas9 | Relaxed PAM (NG) | Expands target range for KO and base editing | NG |
| xCas9(3.7) | SpCas9 | Broad PAM recognition (NG, GAA, GAT) | Expands target range | NG, GAA, GAT |
| SpCas9n (D10A) | SpCas9 | Nickase; creates single-strand break | Base editor fusion (reduces indels) | NGG |
| nSpCas9 | SpCas9 | Fully nuclease-dead (dCas9) | Transcriptional regulation, prime editor fusion | NGG |
| SaCas9 | S. aureus Cas9 | Smaller size; different PAM | Useful for AAV delivery in animals, tested in plants | NNGRRT |
Enzyme Fusions: The Catalytic Core of Base Editors
Base editors are fusion proteins that link a catalytically impaired Cas protein (nCas9 or dCas9) to a nucleobase deaminase enzyme. This is the definitive component differentiating base editing from traditional CRISPR.
Table 3: Core Base Editor Architectures
| Editor Type | Cas Component | Fused Enzyme | Chemical Conversion | Theoretical Outcome |
|---|---|---|---|---|
| Cytosine Base Editor (CBE) | nCas9 (D10A) | Cytidine Deaminase (e.g., rAPOBEC1, PmCDA1) | C•G → U•G → T•A | C-to-T (or G-to-A) |
| Adenine Base Editor (ABE) | nCas9 (D10A) | Engineered tRNA Adenosine Deaminase (TadA*) | A•T → I•T → G•C | A-to-G (or T-to-C) |
| Dual Base Editor | nCas9 | Fusion of deaminases (e.g., CBE+ABE) | C-to-T & A-to-G simultaneously | Combined transition mutations |
Experimental Protocol: Assessing Base Editing Efficiency in Protoplasts
A standard transient assay to quantify editing components' performance.
1. Materials: Plant expression vectors for base editor and gRNA, plant protoplasts (e.g., from rice or Arabidopsis), PEG transformation solution, DNA extraction kit, PCR reagents, sequencing primers.
2. Methodology:
- Construct Assembly: Clone your target gRNA into a plant expression vector (e.g., driven by AtU6 promoter). Use a vector expressing the base editor (BE) under a strong constitutive promoter (e.g., 35S or ZmUbi).
- Protoplast Isolation & Transfection: Isolate protoplasts from etiolated seedlings using cellulase and macerozyme digestion. Co-transfect 10-20 µg of BE plasmid and 5-10 µg of gRNA plasmid into ~10⁶ protoplasts using PEG-mediated transformation.
- Incubation: Incubate transfected protoplasts in the dark at room temperature for 48-72 hours.
- Genomic DNA Extraction: Harvest protoplasts and extract genomic DNA.
- Target Amplification: PCR-amplify the target genomic region from transfected and control samples.
- Editing Analysis: Submit PCR products for Sanger sequencing. Analyze chromatograms for base conversion signatures using tools like BEAT or EditR. For high-resolution data, perform next-generation amplicon sequencing.
3. Key Metrics: Calculate editing efficiency as (% of sequencing reads showing the desired base conversion) at the target position within the activity window.
Title: Base Editing Workflow in Plant Protoplasts
Title: Base Editor Component Assembly & Action
The Scientist's Toolkit: Key Research Reagent Solutions
| Reagent/Material | Supplier Examples | Primary Function in Base Editing Research |
|---|---|---|
| High-Fidelity DNA Polymerase | NEB (Q5), Thermo Fisher | Error-free amplification of gRNA and target sequences for cloning and analysis. |
| T7 Endonuclease I / Surveyor Nuclease | NEB, IDT | Detects indel mutations from traditional CRISPR-Cas9 (not for base editing analysis). |
| Plant Genomic DNA Extraction Kit | Qiagen, MP Biomedicals | Clean gDNA isolation from transfected tissues for PCR and sequencing. |
| Next-Generation Sequencing Kit (Amplicon) | Illumina, Swift Biosciences | Quantifies base editing efficiency and byproducts at high depth and accuracy. |
| PEG Transformation Solution | Sigma-Aldrich, homemade (40% PEG) | Mediates plasmid DNA delivery into plant protoplasts. |
| Cellulase & Macerozyme R-10 | Yakult Pharmaceutical | Enzymatic digestion of plant cell walls for protoplast isolation. |
| Modular Cloning System (e.g., Golden Gate) | Addgene (MoClo), kits available | Enables rapid, standardized assembly of multiple gRNA and effector constructs. |
| Chemically Competent E. coli | NEB Stable, Thermo Fisher | Transformation of base editor plasmids, which can be large and unstable. |
| Sanger Sequencing Service | Genewiz, Eurofins | Initial, cost-effective screening for editing events at target loci. |
| BE-Analyzer Web Tool | Publicly available (benchling.com) | Designs optimal gRNAs by predicting location within base editor activity window. |
The strategic selection and optimization of guide RNA, Cas protein variants, and enzyme fusions directly dictate the success of base editing versus traditional CRISPR-Cas9 in crops. Base editors, by obviating the need for DSBs and HDR, offer a more predictable and efficient route for installing agronomically valuable point mutations. However, their application is constrained by PAM requirements, a narrow activity window, and potential for off-target edits. Ongoing research focuses on developing next-generation editors with expanded targeting scope, minimized off-target effects, and tailored delivery systems for stable crop transformation, solidifying their central role in the future of precision crop breeding.
The application of genome editing in crop improvement hinges on the ability to generate predictable and precise genetic modifications. Traditional CRISPR-Cas9 systems, employing a Cas9 nuclease and a guide RNA (gRNA), create double-strand breaks (DSBs) that are repaired primarily by error-prone non-homologous end joining (NHEJ) or homology-directed repair (HDR). This results in two primary classes of molecular outputs: insertion-deletion mutations (Indels) and, less frequently, precise HDR-mediated edits. In contrast, base editing technologies, which fuse a catalytically impaired Cas protein (nCas9 or dCas9) to a deaminase enzyme, facilitate direct, irreversible chemical conversion of one base pair to another without creating a DSB, enabling efficient generation of precise point mutations. This whitepaper provides an in-depth technical comparison of these key molecular outputs, framed within the evolving paradigm of crop genome engineering.
Core Mechanisms and Molecular Outputs
Traditional CRISPR-Cas9 Pathway Outputs
Indels via NHEJ: The predominant repair pathway for Cas9-induced DSBs in plants is NHEJ. This often results in small insertions or deletions at the break site, leading to frameshifts and gene knockouts.
Precise Edits via HDR: When a donor DNA template with homology arms is co-delivered, the DSB can be repaired via HDR. This allows for precise nucleotide changes, gene insertions, or gene replacements. However, HDR efficiency in plants is typically very low (<10%) and is often confounded by predominant NHEJ activity.
Base Editing Outputs
Precise Point Mutations: Base editors (BEs) achieve targeted base conversions without DSBs. Cytosine Base Editors (CBEs) facilitate C•G to T•A conversions, while Adenine Base Editors (ABEs) facilitate A•T to G•C conversions. The output is a precise, predictable point mutation within a defined editing window (typically 4-5 nucleotides wide) with minimal indel byproducts.
Quantitative Data Comparison
Table 1: Comparative Output Metrics of CRISPR-Cas9 Nuclease vs. Base Editors in Model Crops
| Parameter | Traditional CRISPR-Cas9 (NHEJ) | Traditional CRISPR-Cas9 (HDR) | Cytosine Base Editor (CBE) | Adenine Base Editor (ABE) |
|---|---|---|---|---|
| Primary Output | Indels (Knockout) | Precise sequence change/insertion | C•G to T•A point mutation | A•T to G•C point mutation |
| Typical Efficiency in Crops | High (10-80%) | Very Low (0.1-10%) | Moderate to High (1-60%)* | Moderate to High (1-40%)* |
| DSB Formation | Yes | Yes | No (Uses nCas9) | No (Uses nCas9) |
| Donor Template Required | No | Yes (dsODN or plasmid) | No | No |
| Byproduct Formation (Indels) | N/A | Common (from NHEJ) | Low (<1-5%) | Very Low (<1%) |
| Multiplexing Capability | High | Low (due to donor complexity) | High | High |
| Example Crop Application | Knockout of susceptibility genes | Herbicide resistance allele swap | Creating stop codons (C to T) | Creating favorable amino acid changes (A to G) |
*Efficiency is highly dependent on promoter, construct design, and delivery method. Data compiled from recent studies in rice, wheat, and tomato (2023-2024).
Experimental Protocols
Protocol for Analyzing Indels and HDR in CRISPR-Cas9 Experiments
Title: Amplicon Sequencing for Mutation Characterization in Regenerated Plantlets
Materials: Leaf tissue from T0 plant, DNA extraction kit, high-fidelity PCR polymerase, gene-specific primers flanking target site, gel purification kit, NGS library prep kit, bioinformatics pipeline (e.g., CRISPResso2).
Method:
- Genomic DNA Extraction: Isolate high-quality gDNA from ~100mg of leaf tissue.
- PCR Amplification: Design primers ~150-200bp upstream/downstream of target. Perform PCR with high-fidelity polymerase to generate amplicon for sequencing.
- Amplicon Purification: Gel-purify the PCR product to ensure specificity.
- Next-Generation Sequencing (NGS) Library Prep: Use a dual-indexing strategy to barcode samples. Pool equimolar amounts of each amplicon library.
- Sequencing: Run on an Illumina MiSeq or NovaSeq platform (2x250bp or 2x150bp).
- Bioinformatic Analysis: Process reads through CRISPResso2 to quantify percentages of: a) wild-type sequence, b) indels (NHEJ), c) precise HDR events (alignment to donor template). Filter for reads with ≥10x coverage.
Protocol for Quantifying Base Editing Efficiency and Purity
Title: Targeted Sequencing Analysis for Base Editor Performance
Materials: As in Protocol 4.1.
Method:
- Follow steps 1-5 from Protocol 4.1.
- Bioinformatic Analysis (Specialized): Process reads through base editor-specific tools (e.g., BEAT or CRISPResso2 in base editor mode).
- Quantify the percentage of reads with intended C-to-T or A-to-G conversion within the editing window.
- Precisely quantify the percentage of reads with any indels (byproduct).
- Calculate the "product purity" ratio: (Reads with intended base edit) / (Reads with intended base edit + Reads with indels).
Diagrams
Title: CRISPR-Cas9 vs Base Editing Mechanism Comparison
Title: Mutation Analysis Workflow for Regenerated Plants
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Crop Genome Editing & Analysis
| Reagent / Material | Function in Experiment | Key Consideration for Crops |
|---|---|---|
| High-Efficiency Cas9 Vector (e.g., pRGEB32, pYLCRISPR/Cas9) | Drives expression of Cas9 nuclease. | Use plant-codon optimized Cas9 with strong, constitutive promoters (e.g., ZmUbi, OsActin). |
| Base Editor Vector (e.g., pnCas9-PBE, pABE8e) | Drives expression of nCas9-deaminase fusion. | Select editor with appropriate editing window and PAM compatibility (SpCas9-NG, xCas9). |
| gRNA Cloning Kit / Golden Gate Assembly System | For modular assembly of single or multiplexed gRNA expression cassettes. | Ensures precise, high-throughput construct assembly. Pol III promoters (OsU3, OsU6) are standard. |
| HDR Donor Template (ssODN or dsDNA with homology arms) | Provides repair template for precise edits via HDR. | For plants, dsDNA with 500-1000bp homology arms is typical. Chemical modification can enhance stability. |
| Plant Transformation Reagents (Agrobacterium strains, Biolistic gun, PEG for protoplasts) | Delivery of editing machinery into plant cells. | Choice depends on crop species (Agrobacterium for dicots, biolistics for monocots like wheat). |
| High-Fidelity PCR Polymerase (e.g., Phusion, KAPA HiFi) | Amplification of target locus for sequencing analysis without introducing errors. | Critical for generating clean amplicons for NGS to avoid false-positive mutation calls. |
| NGS Mutation Detection Service/Pipeline (e.g., Amplicon-EZ, CRISPResso2 cloud) | Accurate quantification of editing outcomes (indels, base edits, HDR). | Outsourcing can be efficient; in-house pipelines require bioinformatics expertise. |
| Selectable Marker / Reporter System (e.g., GFP, hygromycin resistance) | Enriches for transformed cells/plants in tissue culture. | Screenable markers (GFP) allow early selection; antibiotics require optimized crop-specific protocols. |
Protocols and Pipelines: Implementing Base Editing and CRISPR-Cas9 in Crop Systems
Vector Design and Construct Assembly for Plant Transformation
Within the accelerating field of crop genome engineering, the debate between Base Editing and traditional CRISPR-Cas9 hinges on the precision, efficiency, and outcome of the edit. Traditional CRISPR-Cas9 induces double-strand breaks (DSBs), relying on error-prone repair pathways like Non-Homologous End Joining (NHEJ) or Homology-Directed Repair (HDR). Base editing, in contrast, directly converts one base pair to another without a DSB, offering a more predictable and cleaner edit but within a narrower window. The success of both modalities is fundamentally dependent on the initial step: the design and assembly of the transformation vector. This guide provides a technical deep-dive into the modern principles of plant vector construction, optimized for contemporary editing tools.
Core Vector Components and Design Considerations
A plant transformation vector is a recombinant DNA construct that delivers the gene-editing machinery into the plant cell and ensures its stable integration or transient expression. Key components must be carefully selected and assembled.
Essential Genetic Modules
- Promoter(s): Drives expression of the editing machinery. Constitutive promoters like CaMV 35S or Ubiquitin are common, but tissue-specific or inducible promoters can reduce off-target effects and cytotoxicity.
- Gene-Editing Cassette: Encodes the core effector protein.
- For Traditional CRISPR-Cas9: A gene for a nuclease-active Cas9 (e.g., SpCas9).
- For Base Editing: A gene for a nickase Cas9 (nCas9) or deaminase-fused dead Cas9 (dCas9) coupled with a cytidine (CBE) or adenine (ABE) deaminase.
- Guide RNA (gRNA) Expression Unit: Typically uses a RNA Pol III promoter (e.g., AtU6) to drive expression of a single-guide RNA (sgRNA) targeting the genomic locus of interest.
- Selectable Marker Gene: Essential for identifying transformed tissue (e.g., hptII for hygromycin resistance, bar for glufosinate resistance).
- Plant Regulatory Sequences: Including a terminator/polyadenylation signal (e.g., NOS terminator) for each expressed gene.
- Bacterial Backbone: Contains an origin of replication (ori) and bacterial selection marker (e.g., spectinomycin resistance) for propagation in E. coli and Agrobacterium.
Backbone Choice: Binary Vectors forAgrobacterium-Mediated Transformation
Agrobacterium tumefaciens-mediated transformation (the most common method for dicots and many monocots) requires a T-DNA binary vector system. The T-DNA region, flanked by left and right borders (LB, RB), is transferred into the plant genome.
Quantitative Comparison: Vector Design for Cas9 vs. Base Editing
The choice of editor imposes specific design constraints, as summarized in the table below.
Table 1: Vector Design Parameters for Traditional CRISPR-Cas9 vs. Base Editing
| Parameter | Traditional CRISPR-Cas9 Vector | Base Editing Vector | Rationale |
|---|---|---|---|
| Core Effector | Nuclease-active Cas9 (e.g., SpCas9) | Nickase Cas9 (nCas9) or dead Cas9 (dCas9) fused to deaminase (e.g., rAPOBEC1 for CBE, TadA for ABE) | Base editors require catalytically impaired Cas9 to avoid DSBs while maintaining targeting. |
| gRNA Design | Target sequence + NGG (SpCas9 PAM) | Target sequence + PAM, with edit window (typically positions 4-8 for CBE, 4-7 for ABE) within spacer. | The editable "window" is restricted by deaminase activity range relative to PAM. |
| Repair Template | Optional, co-delivered for HDR-mediated precise edits. | Not required. The deamination chemistry directly converts C to T (or G to A) or A to G (or T to C). | Base editing is a standalone chemical conversion without relying on endogenous repair templates. |
| Multiplexing Strategy | Multiple sgRNAs expressed from individual Pol III promoters or as a tRNA-gRNA array. | Similar multiplexing possible, but deaminase activity window must be considered for each target. | Multiplexed base editing can introduce multiple point mutations simultaneously. |
| Primary Outcome | Indels (NHEJ) or precise sequence insertion (HDR). | Point mutation (transition: C->T, A->G) without indels. | Reduces genomic scarring and unpredictable mutations common with NHEJ. |
| Typical Edit Efficiency (in plants) | Highly variable: 0-60% for NHEJ, <5% for HDR. | Often higher and more consistent for point mutations: 10-50% (without selection). | Avoidance of DSB and complex repair pathways leads to more predictable and efficient outcomes. |
Construct Assembly: Modern Methodologies
Golden Gate and Gibson Assembly are the dominant methods for modular, scarless vector construction.
Protocol: Modular Assembly via Golden Gate Cloning
This method uses Type IIS restriction enzymes (e.g., BsaI, BpiI) which cut outside their recognition site, allowing for the assembly of multiple fragments in a defined order.
Detailed Protocol:
- Design & Amplify Modules: Design all vector components (promoters, effectors, gRNAs, terminators) as separate modules flanked by appropriate Type IIS sites (e.g., BsaI sites with 4bp overhangs). Generate modules via PCR or gene synthesis.
- Prepare Reaction Mix:
- Equimolar mix of each DNA module (e.g., 50 fmol each).
- 1x T4 DNA Ligase Buffer.
- 10 U BsaI-HFv2 (or similar Type IIS enzyme).
- 400 U T4 DNA Ligase.
- Nuclease-free water to 20 µL.
- Run Thermo-Cycling Digestion/Ligation: Program a thermocycler: 37°C for 5 min (digestion) -> 16°C for 5 min (ligation). Repeat for 30-50 cycles. Final step: 50°C for 5 min (to inactivate BsaI), 80°C for 5 min (to inactivate ligase).
- Transformation: Transform 2-5 µL of the reaction into competent E. coli. Select on appropriate antibiotic.
- Validation: Screen colonies by colony PCR and/or restriction digest. Validate final construct by Sanger sequencing.
Experimental Workflow: From Design to Transformed Plant
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Vector Construction and Plant Editing
| Item | Function/Description | Example Product/Supplier |
|---|---|---|
| Type IIS Restriction Enzymes | Core enzyme for Golden Gate assembly. Creates unique, non-palindromic overhangs. | BsaI-HFv2, BpiI (NEB); Esp3I (Thermo). |
| High-Fidelity DNA Polymerase | For error-free amplification of vector modules and verification PCRs. | Q5 (NEB), Phusion (Thermo), KAPA HiFi (Roche). |
| T4 DNA Ligase | Joins DNA fragments with compatible overhangs during assembly. | T4 DNA Ligase (NEB, Thermo). |
| Commercial DNA Assembly Mix | Pre-mixed enzymes for simplified Gibson or Golden Gate assembly. | NEBuilder HiFi DNA Assembly Mix (NEB), Gibson Assembly Master Mix (NEB). |
| Competent E. coli | For plasmid propagation and cloning. High-efficiency strains are crucial for complex assemblies. | NEB Stable, NEB 5-alpha (NEB); DH5α (Thermo). |
| Competent Agrobacterium | For plant transformation. Electrocompetent strains are standard. | A. tumefaciens GV3101, EHA105 (various suppliers). |
| Plant Selection Antibiotic | Selects for transformed plant tissue. Choice depends on the marker gene in the vector. | Hygromycin B, Glufosinate-ammonium (GoldBio, Sigma). |
| Modular Cloning Toolkit | Pre-made collections of standardized DNA parts (promoters, terminators, markers) for plant vector assembly. | MoClo Plant Toolkit (Addgene); GoldenBraid system. |
| gRNA Design & Off-Target Prediction Software | In silico tools for selecting specific sgRNAs and assessing potential off-target sites. | CRISPR-P, CHOPCHOP, Cas-OFFinder. |
Within the accelerating field of plant genome engineering, the selection of an optimal delivery method is as critical as the choice of editing machinery. The paradigm shift from traditional CRISPR-Cas9, which creates double-strand breaks (DSBs) and relies on error-prone repair, to more precise base editing (BE) and prime editing (PE) systems, imposes new technical constraints on delivery. Base editors, comprising a catalytically impaired Cas protein fused to a deaminase enzyme, facilitate direct, irreversible conversion of one base pair to another without inducing DSBs. This whitepaper provides an in-depth technical analysis of three cornerstone delivery methodologies—Agrobacterium-mediated transformation, biolistics, and protoplast transfection—framed within the context of deploying base editing versus traditional CRISPR-Cas9 systems in crop research.
Agrobacterium tumefaciens-Mediated Transformation
Agrobacterium utilizes a natural bacterial vector system to transfer DNA (T-DNA) into the plant genome. For genome editing, the gene(s) of interest are cloned between the T-DNA borders.
Technical Protocol for CRISPR/BE Delivery viaAgrobacterium
- Vector Construction: Clone the expression cassette(s) for the base editor (e.g., BE4max) or SpCas9, along with the guide RNA (sgRNA), into a binary vector (e.g., pCAMBIA1300) between the left and right T-DNA borders. For base editing, ensure the promoter (e.g., pUBI, p35S) is compatible with the deaminase component.
- Strain Preparation: Electroporate the recombinant binary vector into a disarmed A. tumefaciens strain (e.g., EHA105, GV3101).
- Plant Material Preparation: Surface-sterilize explants (e.g., cotyledons, immature embryos) of the target crop species.
- Co-cultivation: Immerse explants in the Agrobacterium suspension (OD~600~ = 0.5-1.0) for 10-30 minutes, then co-cultivate on solid medium for 2-3 days.
- Wash & Selection: Wash explants with sterile water containing a bactericide (e.g., cefotaxime) and transfer to selection medium containing an appropriate antibiotic or herbicide to select for transformed tissue.
- Regeneration & Screening: Regenerate whole plants from putative transgenic calli. Screen regenerants via PCR and sequencing of the target locus to identify edits.
Advantages & Limitations in Base Editing Context
- Advantages: Stable integration is ideal for creating heritable edits and transgenic lines. Low copy number integration reduces complexity. Well-established for many dicots.
- Limitations: The T-DNA transfer process can be slow, potentially allowing for prolonged expression of the editor, which may increase off-target effects—a critical consideration for both Cas9 and BE. Host range limitations exist, especially for monocots. The regeneration process is genotype-dependent and can be lengthy.
Biolistics (Particle Bombardment)
Biolistics involves the acceleration of microscopic, DNA-coated gold or tungsten particles into plant cells using a gene gun, enabling direct physical DNA delivery.
Technical Protocol for CRISPR/BE Delivery via Biolistics
- Microcarrier Preparation: Suspend 1.0 µm gold particles in 100% ethanol, vortex, and pellet. Wash sequentially in sterile water and 50% glycerol. Resuspend in 50% glycerol.
- DNA Coating: For 50 µg of gold, add 10 µg of purified plasmid DNA (encoding BE or Cas9+sgRNA), 50 µl of 2.5M CaCl~2~, and 20 µl of 0.1M spermidine. Vortex for 10 minutes. Pellet, wash with 100% ethanol, and resuspend in 100% ethanol.
- Macrocarrier Loading: Pipette the DNA-gold suspension onto the center of a macrocarrier membrane and allow to dry.
- Target Tissue Preparation: Place embryogenic calli or immature embryos on osmotic treatment medium (e.g., with high sucrose or mannitol) 4-24 hours pre-bombardment.
- Bombardment: Perform bombardment under a partial vacuum (e.g., 28 in Hg) using appropriate pressure (e.g., 1100 psi) and distance (e.g., 9 cm) settings (PDS-1000/He system).
- Post-Bombardment Recovery: Keep tissue on osmotic medium for 12-24 hours, then transfer to standard regeneration/selection medium.
Advantages & Limitations in Base Editing Context
- Advantages: Species- and genotype-independent. Can deliver multiple plasmids simultaneously (e.g., BE + multiple sgRNAs). No vector size constraints. Suitable for organelles.
- Limitations: High cost of equipment. Can cause significant cell damage. Often results in complex, multi-copy integration patterns, complicating molecular analysis. Transient expression can be less efficient than stable integration for achieving edits.
Protoplast Transfection
This method involves the isolation of plant cells whose cell walls have been enzymatically removed, followed by direct introduction of DNA or ribonucleoprotein (RNP) complexes via chemical (PEG) or electrical (electroporation) means.
Technical Protocol for BE RNP Delivery via Protoplast Transfection
- Protoplast Isolation: Slice young leaves into thin strips. Digest in an enzyme solution (e.g., 1.5% cellulase, 0.4% macerozyme, 0.4M mannitol, pH 5.7) for 4-16 hours with gentle shaking.
- Protoplast Purification: Filter the digest through a 70 µm mesh. Pellet protoplasts by centrifugation (100 x g, 5 min). Wash twice with W5 solution (154mM NaCl, 125mM CaCl~2~, 5mM KCl, 2mM MES, pH 5.7). Resuspend in MMg solution (0.4M mannitol, 15mM MgCl~2~, 4mM MES, pH 5.7). Count and adjust density to 1-2 x 10^6^/mL.
- RNP Complex Assembly: For base editing RNPs, pre-complex purified Cas9 nickase (nCas9) protein with chemically synthesized sgRNA (and possibly purified deaminase if not a single protein) at a molar ratio of 1:2-1:5 in nuclease-free buffer. Incubate 10-15 min at room temperature.
- PEG-Mediated Transfection: Mix 100 µL of protoplast suspension with 10-20 µL of RNP complex (or plasmid DNA). Add an equal volume of 40% PEG-4000 solution (in 0.2M mannitol, 0.1M CaCl~2~). Mix gently and incubate for 15-30 min.
- Dilution & Culture: Dilute the transfection mix stepwise with W5 solution. Pellet protoplasts and resuspend in culture medium. Incubate in the dark for 48-72 hours for editing to occur.
- DNA Extraction & Analysis: Harvest protoplasts, extract genomic DNA, and analyze the target locus via PCR/restriction enzyme (RE) assay or next-generation sequencing (NGS).
Advantages & Limitations in Base Editing Context
- Advantages: Enables high-efficiency, transgene-free editing when using RNP delivery—ideal for regulatory and breeding applications. Rapid assessment of editing efficiency (days vs. months). Minimizes off-targets due to short RNP activity.
- Limitations: Regeneration of fertile plants from protoplasts is extremely difficult for many major crops. Technically challenging, requiring optimized isolation protocols. Edits are only in somatic cells until a regeneration system is established.
Comparative Data Analysis
Table 1: Quantitative Comparison of Delivery Methods for Genome Editing
| Parameter | Agrobacterium | Biolistics | Protoplast Transfection (RNP) |
|---|---|---|---|
| Typical Editing Efficiency* | 1-50% (transgenic lines) | 0.1-10% (transient/stable) | 40-80% (transient, somatic cells) |
| Time to Regenerated Plant | 3-12 months | 6-12 months | Often not applicable (lack of regeneration) |
| Transgene Integration Frequency | Low copy (1-3) | High, complex arrays | Zero (with RNP delivery) |
| Ideal for Stable/Heritable Lines | Excellent | Good | Poor |
| Ideal for Transient Assays | Poor | Fair | Excellent |
| Species Versatility | Broad, but limited in monocots | Universal | Broad, but regeneration limited |
| Throughput Potential | Medium | Low | High (for screening) |
| Relative Cost | Low | High (equipment) | Medium |
*Efficiency varies dramatically by species, tissue, and target locus. Base editor efficiency is generally lower than traditional Cas9 knockout efficiency for the same delivery method.
Table 2: Suitability for Traditional CRISPR-Cas9 vs. Base Editing Systems
| Delivery Method | Traditional CRISPR-Cas9 (DSB) | Base Editing (No DSB) | Prime Editing (No DSB) |
|---|---|---|---|
| Agrobacterium | Excellent for knockouts. Prolonged expression can increase off-targets. | Suitable for heritable base changes. Prolonged deaminase activity may increase unintended bystander edits. | Suitable, but large plasmid size (~9kb) can be challenging for T-DNA transfer. |
| Biolistics | Good, but repair of multiple DSBs can cause chromosomal rearrangements. | Suitable, especially for organelles. Particle damage can reduce viable edited cells. | Suitable, but large plasmid size may reduce coating/co-delivery efficiency. |
| Protoplast RNP | Excellent for knockouts. Short-lived activity reduces off-targets. | Optimal for control. Precise RNP dosing minimizes bystander edits. Enables rapid BE optimization. | Challenging due to the large, multi-component PE protein and pegRNA requirements. |
The Scientist's Toolkit: Essential Reagents & Materials
Research Reagent Solutions for Delivery & Analysis
| Item | Function & Relevance |
|---|---|
| Binary Vectors (e.g., pCAMBIA, pGreen) | Backbone for Agrobacterium T-DNA construction. Contain plant selection markers (e.g., hptII, bar). |
| Disarmed A. tumefaciens Strains | Engineered for plant transformation without causing disease. Common: GV3101 (for Arabidopsis), EHA105/AGL1 (for monocots). |
| Gold Microcarriers (1.0 µm) | Inert, high-density particles for coating DNA in biolistics. Superior to tungsten for reproducibility. |
| PEG-4000 (40% solution) | Induces membrane fusion and pore formation for chemical transfection of protoplasts. |
| Cellulase/Macerozyme R-10 | Enzyme cocktails for digesting plant cell walls to isolate viable protoplasts. |
| PureBase Editor Protein (nCas9-Deaminase) | Recombinant, nuclease-free base editor protein for assembly into RNPs for protoplast transfection. |
| Synthetic sgRNA (chemically modified) | High-purity, stabilized guide RNA for RNP assembly; reduces degradation in protoplasts. |
| Deep Sequencing Kit (Amplicon) | For NGS-based quantification of editing efficiency, bystander edits, and off-target effects at predicted sites. |
| T7 Endonuclease I (T7E1) / Surveyor Assay | Mismatch-specific nucleases for quick, gel-based detection of indels from traditional CRISPR, less effective for base edits. |
| High-Sucrose Osmoticum Media | Used pre-/post-bombardment to protect cells from osmotic shock and improve viability. |
Experimental Workflow Visualizations
The trajectory of crop genome engineering toward precision base and prime editing necessitates a re-evaluation of delivery methodologies. Agrobacterium remains the workhorse for generating stable, heritable edits but requires careful monitoring of editor expression duration to minimize unwanted bystander mutations. Biolistics offers unparalleled versatility, especially for difficult-to-transform crops and organellar editing, albeit with trade-offs in integration complexity. Protoplast transfection, particularly with RNP complexes, emerges as the superior method for the rapid, transgene-free optimization and functional validation of base editors, circumventing the pitfalls of prolonged editor activity and enabling high-throughput screening. The optimal choice is dictated by the specific crop, the desired outcome (transient assay vs. heritable line), and the inherent requirements of the precision editing tool itself. A synergistic approach, using protoplasts for rapid tool validation followed by Agrobacterium or biolistics for plant regeneration, is becoming a standard paradigm in advanced crop research laboratories.
Target Selection and gRNA Design Rules for Each Platform
The optimization of genome editing in crops requires a precise choice between base editing and traditional CRISPR-Cas9. This selection dictates the subsequent experimental parameters, most critically the strategy for target selection and guide RNA (gRNA) design. Traditional Cas9 induces double-strand breaks (DSBs), repaired by error-prone non-homologous end joining (NHEJ) or homology-directed repair (HDR). Base editors (BEs)—comprising cytosine base editors (CBEs) and adenine base editors (ABEs)—catalyze direct, irreversible conversion of C•G to T•A or A•T to G•C without a DSB, enabling precise single-nucleotide polymorphisms (SNPs). This guide details the distinct target selection and gRNA design rules for these platforms, a critical decision point within the broader crop engineering thesis where precision, efficiency, and off-target effects are paramount.
- Traditional CRISPR-Cas9 (SpCas9): Targets an NGG Protospacer Adjacent Motif (PAM). Requires gRNAs that place the cut site (typically between bases 17 and 18 upstream of the PAM) at the intended disruption locus. Ideal for gene knock-outs, large deletions, and, with a donor template, knock-ins.
- Cytosine Base Editors (e.g., BE4, hA3A-BE3): Typically use a Cas9 nickase (nCas9) fused to a cytidine deaminase. The editable window is a narrow zone (typically positions 4-8, or 4-10, counting the PAM as 21-23) within the single-stranded DNA bubble created by the Cas9-gRNA complex. The target C must be within this window and preferably have an optimal sequence context.
- Adenine Base Editors (e.g., ABE8e): Use nCas9 fused to an engineered tRNA adenosine deaminase. They operate within a similar but not identical single-stranded window (typically positions 4-9). The target A must be within this window.
Quantitative Comparison: Platform-Specific Design Rules
Table 1: Core Design Parameters for CRISPR Platforms in Crops
| Parameter | Traditional Cas9 (SpCas9) | Cytosine Base Editor (CBE) | Adenine Base Editor (ABE) |
|---|---|---|---|
| Primary PAM | NGG (canonical) | NGG (derived from SpCas9) | NGG (derived from SpCas9) |
| Editing Outcome | DSB → Indels (NHEJ) or HDR | C•G to T•A conversion | A•T to G•C conversion |
| Critical Target Window | Cut site (~-3 to -4 from PAM) | R-loop "Activity Window" (e.g., positions 4-10, 5-7 optimal) | R-loop "Activity Window" (e.g., positions 4-9, A5-A7 optimal) |
| Key Sequence Context | Avoid homopolymers near cut site | Avoid >1 consecutive C in window; optimal 5' TC or AC context for rAPOBEC1-based editors | No strict context, but efficiency varies |
| Off-Target Concern | DSB at genomic sites with similar sequence | Bystander edits (Cs in window), DNA/RNA off-target deamination | Bystander edits (As in window), generally lower RNA off-target |
| Typical Efficiency in Crops | 1-30% (HDR <1%) | 10-50% (product purity varies) | 10-40% (high product purity) |
| Optimal Use Case | Gene knock-out, large deletions, fragment insertion | Correcting C•G to T•A SNPs, introducing stop codons (CAA->TAA, CAG->TAG) | Correcting A•T to G•C SNPs, introducing start codons, reverting T•A to C•G |
Table 2: Design Rule Checklist for Crop gRNA Design
| Rule | Traditional Cas9 | Base Editor (CBE/ABE) | Rationale |
|---|---|---|---|
| 1. PAM Presence | Must have NGG at target locus. | Must have NGG at target locus, positioned so the target base(s) fall in the activity window. | Cas9 binding prerequisite. |
| 2. On-Target Efficiency | Predict using tools (e.g., CRISPR-P, CHOPCHOP). GC content 40-60%. | Predict using BE-specific tools (e.g., BE-HIVE, BE-DESIGN). | Impacts editing rates. BEs have different sequence preferences. |
| 3. Specificity (Off-Target) | Minimize hits with ≤3 mismatches in seed region (PAM-proximal 12 nt). | Critical: Also minimize bystander C/A bases within the activity window. | Reduces genomic off-target DSBs. Reduces unintended on-target base conversions. |
| 4. Genomic Context | Avoid repetitive regions, centromeres. Check for SNP in gRNA in varietal lines. | Essential: Check for SNP in the activity window in varietal lines. | Ensures unique targeting. Prevents gRNA mis-binding or editing the wrong allele. |
| 5. Biological Outcome | For KO: target early constitutive exon, ensure frameshift. For HDR: cut close to edit. | Ensure base change creates desired amino acid change or regulatory element. Verify via codon usage table. | Maximizes functional impact. Ensures intended biological result. |
Experimental Protocols
Protocol 1: In Silico gRNA Design and Selection Workflow
- Sequence Acquisition: Retrieve the genomic sequence of the target gene from a crop-specific database (e.g., MaizeGDB, Rice Genome Annotation Project).
- PAM Identification:
- For Cas9: Scan for all 5'-NGG-3' sites in the sense and antisense strands of the target exon/region.
- For BE: Identify NGG sites where the target SNP or nucleotide is positioned within the editor's activity window (positions ~4-10 upstream of the PAM).
- gRNA Candidate Generation: Extract the 20-nt protospacer sequence immediately 5' to each PAM.
- Specificity Analysis: Input each 20-nt spacer into a specificity tool (e.g., Cas-OFFinder) against the relevant crop genome. Filter out gRNAs with potential off-target sites with ≤3 mismatches, especially in the seed region.
- Efficiency Prediction: Score remaining gRNAs using a validated algorithm (e.g., DeepSpCas9 for Cas9; BE-DESIGN for BEs). Select top 3-5 candidates.
- Final Validation: Manually inspect candidates for genomic uniqueness, absence of homopolymers, and optimal GC content. For BEs, analyze bystander positions.
Protocol 2: Validation of gRNA Efficacy (Agrobacterium-mediated Transformation in Rice)
- Vector Construction: Clone selected gRNA sequences into a plant-appropriate expression vector (e.g., pRGEB32 for Cas9, pnCas9-PBE or pnCas9-ABE for BEs) using Golden Gate or restriction-ligation.
- Plant Material: Use embryonic calli from a model rice variety like Nipponbare.
- Transformation: Introduce the vector into Agrobacterium tumefaciens strain EHA105. Co-cultivate rice calli with Agrobacterium for 3 days.
- Selection & Regeneration: Transfer calli to selection media containing hygromycin for 2-3 weeks. Regenerate shoots and then roots on appropriate media.
- Genotyping (T0 Plants): a. Extract genomic DNA from leaf tissue. b. PCR-amplify the target region (primers ~150bp flanking the edit site). c. For Cas9: Analyze by Sanger sequencing followed by decomposition tools (TIDE, ICE) or next-generation sequencing (NGS) to quantify indel frequencies. d. For BE: Perform Sanger sequencing. The presence of overlapping peaks at the target base(s) indicates editing. Quantify precise base conversion efficiency and bystander edits via NGS amplicon sequencing.
Mandatory Visualization
Title: CRISPR Platform Selection Logic for Crop Editing
Title: Universal gRNA Design & Selection Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Target Selection & gRNA Validation in Crops
| Reagent / Material | Function & Importance in Workflow |
|---|---|
| Crop-Specific Genomic Database (e.g., Phytozome, Gramene) | Source of accurate reference sequences and gene annotations for protospacer and PAM identification. |
| gRNA Design Software (e.g., CRISPR-P 2.0, CHOPCHOP, BE-DESIGN) | Algorithms to predict on-target efficiency and off-target sites, tailored for plant genomes and base editors. |
| Specificity Checker (e.g., Cas-OFFinder) | Identifies potential genomic off-target sites by allowing mismatches, critical for minimizing unintended edits. |
| Plant CRISPR Vector (e.g., pRGEB32, pYLCRISPR/Cas9Pubi-B) | Binary vectors containing Pol III promoters for gRNA(s) and plant-specific promoters for Cas9/BE expression. |
| Agrobacterium tumefaciens Strain (e.g., EHA105, GV3101) | Standard vector for stable transformation of many crop species, particularly monocots and dicots. |
| High-Fidelity DNA Polymerase (e.g., Q5, KAPA HiFi) | For error-free amplification of target loci from plant genomic DNA for cloning and genotyping. |
| NGS Amplicon Sequencing Service | Provides quantitative, base-resolution data on editing efficiency, product purity, and bystander edits. |
| Sanger Sequencing Deconvolution Tool (e.g., ICE, TIDE, BE-Analyzer) | Enables rapid, cost-effective quantification of editing outcomes from Sanger chromatograms. |
Screening and Selection Strategies for Edited Plant Lines
Within the broader thesis contrasting base editing with traditional CRISPR-Cas9 for crop improvement, the development of robust screening and selection strategies is paramount. While traditional CRISPR-Cas9 relies on generating double-strand breaks (DSBs) and subsequent repair via error-prone non-homologous end joining (NHEJ) or homology-directed repair (HDR), base editors enable precise, predictable nucleotide conversions without DSBs. This fundamental difference necessitates distinct and often more refined downstream screening approaches to identify desired edits efficiently and distinguish them from unintended off-target effects.
Core Screening Methodologies: A Comparative Framework
Primary Screening for Targeted Edits
The initial step involves detecting the presence of edits at the target locus.
For Traditional CRISPR-Cas9 Lines:
- Method: PCR amplification of the target region followed by restriction fragment length polymorphism (PCR-RFLP) if the edit disrupts a natural site, or by mismatch cleavage assays.
- Key Assay: T7 Endonuclease I (T7EI) or Surveyor Nuclease assay. These enzymes cleave heteroduplex DNA formed by annealing wild-type and mutated strands, indicating indel presence.
- Protocol (T7EI Assay):
- Isolate genomic DNA from putative edited and control plants.
- PCR-amplify the target locus (200-500 bp amplicon).
- Denature and re-anneal PCR products to form heteroduplexes.
- Digest with T7EI enzyme (NEB) at 37°C for 15-60 min.
- Analyze fragments on a 2-3% agarose gel. Cleaved products confirm mutation.
For Base-Edited Lines:
- Method: High-resolution melting (HRM) analysis or Sanger sequencing followed by decomposition tools.
- Key Assay: HRM can detect single-nucleotide changes. Sanger sequencing traces are analyzed using tools like BEAT or EditR to quantify base conversion efficiency.
- Protocol (HRM Screening):
- Perform PCR on genomic DNA using a saturating DNA dye (e.g., EvaGreen).
- Run on a real-time PCR instrument with HRM capability.
- After amplification, incrementally increase temperature from 65°C to 95°C while monitoring fluorescence.
- Analyze melting curve shapes. Altered curves indicate a sequence variant (base edit).
Advanced Genotyping and Sequence Verification
Definitive confirmation requires sequencing.
- Sanger Sequencing & Deconvolution: Standard for initial validation. For base editing, trace files must be analyzed with decomposition software to detect sub-stoichiometric editing.
- Next-Generation Sequencing (NGS): Essential for comprehensive analysis.
- Amplicon-Seq: Deep sequencing of PCR-amplified target loci. Reveals editing efficiency, precise edit identity, and potential small indels.
- Whole-Genome Sequencing (WGS): Gold standard for identifying off-target effects genome-wide, crucial for both CRISPR-Cas9 and base editor lines.
Selection Strategies: Enriching for Edited Cells/Plants
- Visual Markers (e.g., GFP, YFP): Co-expressed with editors to transiently identify transformed tissue.
- Antibiotic/Herbicide Resistance: Selectable marker genes (e.g., hptII, bar) integrated with the editor enable survival of transformed tissue on selective media.
- PCR-based Selection: Using allele-specific primers that amplify only the edited sequence.
- Protoplast Enrichment: Fluorescence-activated cell sorting (FACS) of protoplasts expressing fluorescent markers linked to the editing machinery.
Quantitative Comparison of Key Outcomes
Table 1: Typical Efficiency and Outcome Profiles of Editing Systems in Plants
| Parameter | Traditional CRISPR-Cas9 (NHEJ) | Base Editing (Cytosine or Adenine) |
|---|---|---|
| Primary Edit Type | Small insertions/deletions (indels) | Precise C•G to T•A or A•T to G•C conversions |
| Typical On-Target Efficiency | 1-30% (varies by species, target) | 0.1-50% (highly variable, often lower than Cas9) |
| Product Purity | Mixed population of indels at locus | Primarily targeted point mutation; can have bystander edits |
| DSB Formation | Yes (inherent) | No (theoretical) |
| Common Off-Target Effects | Indels at off-target sites with high homology | Point mutations at off-target sites; can be RNA-dependent |
| Ideal Screening Method | T7EI/Surveyor + Amplicon-Seq | HRM + Amplicon-Seq + Trace Decomposition |
Table 2: Analysis Methods for Edited Plant Lines
| Method | Detects | Throughput | Cost | Best For |
|---|---|---|---|---|
| T7EI / Surveyor | Indel presence (binary) | Medium | Low | Primary screen for CRISPR-Cas9 NHEJ lines |
| HRM Analysis | Sequence variation (SNPs/indels) | High | Low | Primary screen for base-edited or HDR lines |
| Sanger Sequencing | Exact sequence at target | Low | Medium | Confirmation, low-plex validation |
| Amplicon-Seq | Exact sequence, efficiency, heterogeneity | High | Medium-High | Definitive on-target analysis, efficiency quantitation |
| Whole-Genome Seq | All mutations genome-wide | Low | High | Comprehensive off-target assessment, event characterization |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Screening and Selection
| Item | Function & Application | Example Vendor(s) |
|---|---|---|
| T7 Endonuclease I | Mismatch-specific nuclease for detecting indels via cleavage assay. | New England Biolabs (NEB), Integrated DNA Technologies (IDT) |
| Surveyor Nuclease | Alternative mismatch-specific nuclease for indel detection. | IDT |
| EvaGreen Dye | Saturating DNA dye for HRM analysis, enables high-resolution melt curves. | Biotium |
| Phusion U Green Mix | High-fidelity PCR master mix optimized for HRM. | Thermo Fisher Scientific |
| ZymoBIOMICS DNA Kit | Reliable genomic DNA extraction from plant tissue for PCR. | Zymo Research |
| NEBNext Ultra II FS | Library preparation kit for amplicon or whole-genome sequencing. | NEB |
| Guide-it Genotype Confirmation Kit | Streamlined kit for PCR and T7EI analysis of CRISPR edits. | Takara Bio |
| BEAT (Base Editing Analysis Toolkit) | Web-based tool for deconvoluting Sanger traces from base-edited samples. | Public Software |
| EditR Software | Tool for quantifying base editing efficiency from Sanger sequencing data. | Public Software |
Experimental Protocols in Detail
Protocol 1: Comprehensive Amplicon-Seq for On-Target Analysis
- Genomic DNA Extraction: Use a CTAB-based method or commercial kit for high-quality gDNA.
- Primary PCR: Amplify target locus(s) with gene-specific primers containing overhangs for subsequent indexing. Use high-fidelity polymerase (e.g., Q5). Cycle: 98°C 30s; 35 cycles of (98°C 10s, 65°C 20s, 72°C 20s); 72°C 2 min.
- Clean-up: Purify PCR products with magnetic beads (e.g., SPRIselect).
- Indexing PCR: Add dual indices and Illumina sequencing adapters via a second, limited-cycle PCR.
- Clean-up & Pool: Purify indexed libraries, quantify, and pool equimolarly.
- Sequencing: Run on an Illumina MiSeq or iSeq with paired-end 2x150 or 2x250 cycles.
- Bioinformatics: Process with CRISPResso2, BEAT-AMPLICON, or a custom pipeline (align to reference, call variants, quantify editing efficiency).
Protocol 2: Phenotypic Selection for Herbicide Resistance (e.g.,bargene)
- Transformation: Use Agrobacterium or biolistics to deliver editing construct with linked bar gene (confers resistance to glufosinate/Basta).
- Primary Selection: Culture explants/tissue on medium containing glufosinate ammonium (e.g., 3-5 mg/L). Subculture surviving tissue every 2 weeks.
- Regeneration: Transfer resistant calli to regeneration medium with the same selective agent.
- Plantlet Screening: Transfer regenerated shoots to rooting medium with selection.
- Molecular Confirmation: Perform PCR for bar gene presence and genotype the target locus to confirm linkage between the edit and the selectable marker.
Workflow Visualizations
Screening & Selection Workflow for Edited Plants
Genotyping Strategy for Base-Edited Lines
This whitepaper presents an in-depth technical analysis of trait improvement in crops, framed within a critical evaluation of base editing versus traditional CRISPR-Cas9 systems. The precision of base editing—allowing for targeted point mutations without double-strand DNA breaks (DSBs)—offers distinct advantages for specific applications, which are explored through the following case studies.
Case Study: Herbicide Resistance via Acetolactate Synthase (ALS) Gene Modification
Scientific Context & Objective
The ALS enzyme is a target for sulfonylurea and imidazolinone herbicides. A single nucleotide change (C to T) at a specific position in the ALS gene can confer resistance by altering the herbicide-binding site. Traditional CRISPR-Cas9-mediated homology-directed repair (HDR) for this conversion is inefficient in plants. Base editing, specifically employing a cytosine base editor (CBE), provides a precise, DSB-free alternative.
Experimental Protocol
- Plant Material: Oryza sativa (Rice) protoplasts and calli.
- Vector Design: A CBE (rAPOBEC1-nCas9-UGI) was fused to a plant codon-optimized nCas9 (D10A). The sgRNA was designed to target the specific cytosine within the ALS gene (e.g., position 572 in OsALS).
- Delivery: Agrobacterium tumefaciens-mediated transformation of rice callus.
- Selection & Screening: Calli were selected on herbicide-containing medium. Surviving lines were genotyped via Sanger sequencing followed by deconvolution (ICE analysis or amplicon deep sequencing) to quantify editing efficiency.
- Phenotyping: T1 plants were sprayed with field-relevant doses of the target herbicide, and injury scores were recorded over 21 days.
Table 1: Efficiency of ALS Herbicide-Resistance Introduction in Rice
| Editing System | nCas9 Variant | Base Editor Type | Target Change (C->T) | Editing Efficiency (% in T0) | Homozygous Mutants in T1 (%) | Herbicide Survival Rate (T1) |
|---|---|---|---|---|---|---|
| Traditional CRISPR-Cas9 | SpCas9 (WT) | HDR-dependent | C->T | < 2.0% | ~0% | < 5% |
| Base Editing | nCas9 (D10A) | CBE (AID-UGI) | C->T | ~45.2% | ~18.7% | ~92.5% |
| Base Editing | nCas9 (D10A) | CBE (rAPOBEC1-UGI) | C->T | ~53.8% | ~22.1% | ~95.0% |
Pathway & Workflow Diagram
Title: Workflow for Base Editing Herbicide Resistance in Rice
Case Study: Nutritional Quality Enhancement via Pro-Vitamin A (β-Carotene) Biosynthesis
Scientific Context & Objective
In rice endosperm, β-carotene biosynthesis is limited. The enzyme β-carotene hydroxylase (BCH1) converts β-carotene into less nutritionally potent xanthophylls. Introducing a loss-of-function mutation (bch1) via a precise G:C to A:T transition in a key exon can reduce this conversion, increasing β-carotene accumulation. An Adenine Base Editor (ABE) is ideally suited for this gain-of-nutrient trait.
Experimental Protocol
- Plant Material: Zea mays (Maize) immature embryos.
- Vector Design: An ABE (TadA-8e-nCas9-NLS) was used. The sgRNA targeted the adenine within the ZmBCH1 gene's protospacer.
- Delivery: Particle bombardment (biolistics) of maize embryos.
- Screening: Primary screening via fluorescence marker (RFP) linked to the editor. T0 plants were genotyped using high-throughput amplicon sequencing.
- Quantification: HPLC analysis of carotenoid profiles in T1 and T2 seeds from homozygous edited lines versus wild-type.
Table 2: Enhancement of β-Carotene in Maize via BCH1 Knockout
| Editing System | Target Change | Editing Efficiency (% in T0) | BCH1 Protein Reduction (%) | β-Carotene Increase (μg/g DW) | Lutein/Zeaxanthin Change |
|---|---|---|---|---|---|
| Traditional CRISPR-Cas9 | Indel (Frameshift) | ~70% | ~99% | +5.2 ± 1.1 | -40% |
| Base Editing (ABE) | A:T -> G:C (Stop Codon) | ~28% | ~95% | +4.8 ± 0.9 | -38% |
| Base Editing (ABE) | A:T -> G:C (Missense) | ~35% | ~80% | +3.5 ± 0.7 | -25% |
Carotenoid Pathway & Editing Strategy Diagram
Title: ABE Knockout of BCH1 to Increase β-Carotene in Maize
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Trait Improvement via Base Editing
| Reagent / Material | Function in Experiment | Example Vendor/Product (Illustrative) |
|---|---|---|
| Cytosine Base Editor (CBE) Plasmid | Encodes the fusion protein (e.g., rAPOBEC1-nCas9-UGI) for C->T (or G->A) conversion. | Addgene (#124163; BE4max). |
| Adenine Base Editor (ABE) Plasmid | Encodes the fusion protein (e.g., TadA-8e-nCas9) for A->G (or T->C) conversion. | Addgene (#138489; ABE8e). |
| Plant Codon-Optimized nCas9 (D10A) | The nickase scaffold for base editor fusion; critical for plant expression. | Integrated into above plasmids. |
| sgRNA Cloning Vector | Backbone for assembling and expressing the target-specific sgRNA. | pYPQ141 (U6 promoter vector). |
| Agrobacterium Strain | For stable transformation in dicots/monocots (e.g., rice). | EHA105 or LBA4404. |
| Biolistics/Gene Gun System | For delivery into recalcitrant species (e.g., maize, wheat). | Bio-Rad PDS-1000/He. |
| Herbicide Selection Agent | For in vitro selection of edited cells (e.g., Bispyribac-sodium for ALS). | Sigma-Aldrich (CHEBI). |
| HPLC System with Diode Array | For quantitative analysis of nutritional compounds (carotenoids, vitamins). | Agilent 1260 Infinity II. |
| Amplicon Deep Sequencing Kit | For high-throughput, quantitative assessment of editing efficiency and purity. | Illumina MiSeq Nano Kit. |
| Deconvolution Software (ICE/Synthego) | To analyze Sanger sequencing traces from heterogeneous edited populations. | ICE v2.0 (Synthego). |
Comparative Analysis: Base Editing vs. Traditional CRISPR-Cas9
Table 4: Core Technical Comparison for Trait Improvement Applications
| Parameter | Traditional CRISPR-Cas9 (NHEJ/HDR) | Base Editing (CBE/ABE) | Implication for Trait Development |
|---|---|---|---|
| DNA Lesion | Creates a DSB. | Creates a single-strand nick or no break (PE). | Base editing reduces genomic stress and off-target indels. |
| Primary Outcome | Indels (NHEJ) or precise template-dependent repair (HDR). | Directed point mutation within a ~5-nt window without a donor. | Base editing is superior for point mutations conferring herbicide resistance or altering enzyme function. |
| Efficiency for Point Mutations | Very low for HDR (<5% in plants). | Consistently high (10-50% in plants). | Faster generation of non-transgenic, point-mutated lines. |
| Product Purity | Mixed outcomes (indels + HDR). | Clean, predictable point changes with minimal indels. | Simpler segregation and regulatory characterization. |
| Limitations | PAM dependency, donor template required for HDR. | Restricted to specific base changes (C->T, A->G, etc.) within editing window. Potential for bystander edits. | Base editing is not suitable for gene knock-ins or deletions. Window and bystander effects require careful design and screening. |
Decision Workflow Diagram
Title: Decision Workflow for Choosing Base Editing vs. CRISPR-Cas9
Base editing represents a paradigm shift for specific trait improvement applications in crops, particularly where herbicide resistance or nutritional quality hinges on precise point mutations. The case studies demonstrate its superior efficiency and product purity compared to traditional CRISPR-Cas9 HDR for these ends. However, the choice of editing platform must be driven by the specific genetic change required. The integration of base editing into the plant biotechnology toolkit enables a more precise, efficient, and predictable path to developing improved crop varieties.
Overcoming Technical Hurdles: Maximizing Efficiency and Specificity in Crop Editing
Within the ongoing thesis comparing base editing and traditional CRISPR-Cas9 for crop genome engineering, the paramount challenge remains the accurate identification and mitigation of off-target effects. While base editors offer reduced indel formation compared to Cas9 nuclease, they are not immune to off-target DNA or RNA editing. This guide provides a technical framework for the computational prediction and subsequent empirical validation of off-target sites, a critical pathway for developing precise, safe, and commercially viable edited crops.
Computational Prediction of Off-Target Sites
Computational tools predict potential off-target loci by scanning the genome for sequences homologous to the single guide RNA (sgRNA). Predictions are based on sequence similarity, often allowing for mismatches, bulges, and GC content variations.
Key Algorithms and Tools
- Cas-OFFinder: Allows searches for off-targets with user-defined numbers of mismatches and DNA/RNA bulge lengths across multiple genomes.
- CRISPOR: Integrates off-target prediction (using MIT and cutting frequency determination [CFD] scores) with on-target efficiency scoring.
- BE-Designer & BE-Offinder (from BEEP): Specialized tools for base editor off-target prediction, considering the protospacer adjacent motif (PAM) requirements of nickase Cas9 (e.g., NG for SpCas9) and the wider editing window.
Data Presentation: Comparison of Off-Target Prediction Tools
The following table summarizes the core features and outputs of leading computational tools.
Table 1: Feature Comparison of Primary Off-Target Prediction Tools
| Tool Name | Primary Use Case | Allowed Variations | Key Output Metrics | Reference Genome Support |
|---|---|---|---|---|
| Cas-OFFinder | CRISPR-Cas9 & Base Editor prediction | Mismatches (up to 7), DNA/RNA bulges | List of genomic coordinates & sequence alignment | Customizable (local installation) |
| CRISPOR | CRISPR-Cas9 design & validation | Mismatches (weighted by CFD score) | MIT specificity score, CFD off-target score, efficiency predictions | Pre-loaded major crop genomes |
| BE-Offinder | Base Editor specific prediction | Mismatches within extended sgRNA region | List of potential off-target sites with PAM compatibility | Compatible with BLAST-indexed genomes |
Experimental Protocol: In Silico Off-Target Screening Workflow
- Input: Define the 20-nt sgRNA spacer sequence and the required PAM (e.g., NGG for SpCas9, NG for SpCas9-nickase).
- Tool Selection: For base editing studies, initiate analysis with BE-Offinder. For traditional CRISPR-Cas9, use CRISPOR or Cas-OFFinder.
- Parameter Setting: Set mismatch tolerance (typically 3-5) and bulge allowance (0-1). For crops, specify the correct genome assembly (e.g., Maize B73 RefGen_v4, Rice IRGSP-1.0).
- Execution: Run the genome-wide scan.
- Output Analysis: Rank predicted off-target sites by similarity score (e.g., CFD score). Prioritize sites within gene coding regions, especially those with functional annotations. Export a list of top candidate loci (e.g., top 10-20) for empirical validation.
Empirical Validation of Predicted Off-Targets
Computational predictions require rigorous experimental confirmation. The current gold standard is sequencing-based detection.
Key Validation Methods
- Targeted Deep Sequencing (Amplicon-Seq): High-depth sequencing of PCR amplicons spanning predicted off-target loci.
- Whole-Genome Sequencing (WGS): Unbiased detection of off-target effects genome-wide, though cost and data complexity are higher.
- Circularization for In Vitro Reporting of Cleavage Effects (CIRCLE-seq): An in vitro, cell-free, high-sensitivity method to identify nuclease-dependent off-target sites.
Experimental Protocol: Targeted Deep Sequencing for Off-Target Validation
This protocol validates off-targets in edited plant calli or regenerated tissues.
Materials:
- Genomic DNA from edited and wild-type control plants.
- Predesigned PCR primers flanking each predicted off-target locus (~200-300 bp amplicon).
- High-fidelity PCR master mix.
- Library preparation kit for NGS (e.g., Illumina).
- Next-generation sequencer.
Procedure:
- DNA Extraction: Isolate high-quality gDNA from edited and control samples.
- PCR Amplification: For each off-target locus, perform PCR using locus-specific primers with overhang adapters for NGS.
- Library Preparation: Index PCR products, purify, and pool equimolar amounts to create a sequencing library.
- Sequencing: Run the pooled library on an Illumina MiSeq or HiSeq platform to achieve high coverage (>10,000x per amplicon).
- Data Analysis: Align reads to the reference genome. Use variant calling tools (e.g., GATK) to identify single nucleotide variants (SNVs) or indels at each locus. Compare variant frequency in edited samples versus wild-type controls to confirm off-target editing.
Data Presentation: Sample Off-Target Validation Results
The following table illustrates hypothetical data from an amplicon-seq experiment comparing SpCas9 and an ABE base editor targeting the same OsALS gene in rice.
Table 2: Empirical Off-Target Editing Frequencies for OsALS Targeting in Rice
| Predicted Off-Target Locus (Chr.Position) | Mismatch Count | Gene Context | SpCas9 Editing Frequency (%) | ABE (Adenine Base Editor) Editing Frequency (%) | Validated? |
|---|---|---|---|---|---|
| On-Target: Chr02:1324578 | 0 | OsALS Exon | 92.5 | 68.2 (A•T to G•C) | Yes |
| OT-1: Chr06:4532109 | 3 | Intergenic | 1.7 | 0.05 | Yes |
| OT-2: Chr09:8712345 | 4 | Non-coding RNA | 0.8 | 0.02 | Yes |
| OT-3: Chr11:3298765 | 2 | OsPDS Intron | 15.3 | 0.31 | Yes |
| OT-4: Chr05:1120987 | 5 | Intergenic | 0.0 | 0.0 | No |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Off-Target Analysis
| Item | Function | Example Product/Catalog |
|---|---|---|
| High-Fidelity DNA Polymerase | Accurate amplification of off-target loci for sequencing. | NEB Q5 Hot Start, Takara PrimeSTAR GXL. |
| NGS Library Prep Kit | Preparing barcoded amplicon libraries for Illumina sequencing. | Illumina DNA Prep, Nextera XT. |
| Genomic DNA Extraction Kit (Plant) | High-yield, PCR-grade gDNA from tough plant tissues. | DNeasy Plant Pro, CTAB-based methods. |
| BE or Cas9 Expression Plasmid | Delivery of editing machinery into plant cells. | pRGEB32 (BE), pYAO:Cas9 (CRISPR). |
| Positive Control gDNA | gDNA from a sample with known off-target events for assay validation. | In-house generated or from collaborators. |
Visualizations
Diagram Title: Off-Target Analysis Workflow for Crop Editing
Diagram Title: Off-Target Mitigation in the Thesis Context
Strategies to Improve Editing Efficiency and On-Target Activity
The advent of CRISPR-Cas9 revolutionized genetic engineering, but its reliance on double-strand breaks (DSBs) introduces challenges, including error-prone repair and off-target effects. Base editing, a more recent technology, enables direct, irreversible conversion of one DNA base pair to another without inducing DSBs, offering a potentially cleaner and more predictable editing outcome. In crops research, precision is paramount to develop traits like herbicide resistance or drought tolerance without introducing genomic instability. This guide explores technical strategies to enhance the efficiency and specificity of both traditional CRISPR-Cas9 and base editing systems, framed within their comparative utility for crop genome engineering.
Core Mechanistic Comparison: DSBs vs. Chemical Conversion
Traditional CRISPR-Cas9 creates a blunt-ended DSB, repaired by non-homologous end joining (NHEJ) or homology-directed repair (HDR). This often leads to insertions/deletions (indels). Base editors fuse a catalytically impaired Cas protein (nickase or dead) to a nucleobase deaminase enzyme. Cytosine Base Editors (CBEs) convert C•G to T•A, while Adenine Base Editors (ABEs) convert A•T to G•C within a defined editing window.
Title: Workflow Comparison: CRISPR-Cas9 vs Base Editing
Quantitative Comparison of Editing Outcomes
Recent studies (2023-2024) in major crop models like rice, wheat, and tomato provide a clear performance snapshot.
Table 1: Comparative Performance Metrics in Crops (Representative Studies)
| Editing System | Avg. On-Target Efficiency* | Typical Indel Rate | Typical Off-Target Rate | Primary Outcome |
|---|---|---|---|---|
| CRISPR-Cas9 (NHEJ) | 10-60% (highly variable) | 30-90% of edits | 0.1-5.0% (sequence-dependent) | Insertions/Deletions |
| CRISPR-Cas9 (HDR) | 0.5-10% (often <2%) | N/A | 0.1-5.0% | Precise sequence replacement |
| Cytosine Base Editor (CBE) | 20-70% (can be >90%) | 0.1-10% (from nCas9 nick) | <0.1% (highly specific) | C•G to T•A conversion |
| Adenine Base Editor (ABE) | 30-80% (can be >90%) | 0.1-10% (from nCas9 nick) | <0.1% (highly specific) | A•T to G•C conversion |
| Dual Base Editor | 15-50% per base type | Similar to CBE/ABE | <0.1% | Combined C-to-T and A-to-G |
Efficiency measured as percentage of alleles successfully edited in transformed cells/plants. Data compiled from *Nature Plants, Plant Biotechnology Journal, Genome Biology (2023-2024).
Strategies to Improve Editing Efficiency
For CRISPR-Cas9 (Focus on HDR)
Protocol: Enhancing HDR in Crop Protoplasts via Synchronization & Inhibitors
- Material: Isolate protoplasts from target crop leaf tissue.
- Synchronization: Treat protoplasts with aphidicolin (12 µM for 24h) to arrest cells at S-phase, where HDR is more active.
- Ribonucleoprotein (RNP) Delivery: Electroporate with pre-assembled Cas9 protein + sgRNA RNP complexes (50-100 pmol each) and ~100 pmol dsDNA HDR donor template containing homologous arms (80-120 bp each).
- HDR Promotion: Post-electroporation, incubate protoplasts in medium containing an NHEJ inhibitor (e.g., SCR7, 1 µM) for 48-72h.
- Analysis: Harvest DNA and use droplet digital PCR (ddPCR) with dual-labeled probes to quantify precise HDR integration versus total allelic disruption.
For Base Editing
Protocol: Optimizing Base Editor Expression & Delivery in Plant Cells
- Construct Design: Use a plant-optimized, constitutive promoter (e.g., ZmUbi for monocots, AtUBQ10 for dicots) to express the base editor protein. Employ a Pol III promoter (U6, U3) for sgRNA expression.
- Codons & Localization: Ensure nuclear localization signals (NLS) are intact. Use plant-preferred codons for the deaminase domain.
- Delivery: For Agrobacterium-mediated transformation, use a T-DNA vector with the above constructs. For direct delivery, use RNP complexes of purified base editor protein + in vitro-transcribed sgRNA.
- sgRNA Tuning: Design sgRNAs where the target base(s) fall within positions 4-10 (most efficient window) of the protospacer relative to the PAM. Use online tools (BE-Hive, BE-Design) to predict efficiency.
- Variant Selection: Employ next-generation base editors (e.g., ABE8e for wider window/higher activity; engineered CBE variants with reduced off-target RNA editing).
Title: Base Editing Optimization Workflow for Crops
Strategies to Enhance On-Target Specificity
For CRISPR-Cas9
- High-Fidelity Cas9 Variants: Use SpCas9-HF1 or eSpCas9(1.1) to reduce non-specific DNA contacts.
- Truncated sgRNAs (tru-gRNAs): Use 17-18 nt guides instead of 20 nt, which can increase specificity by tolerating fewer mismatches.
- Dual Nickase Strategy (Paired nCas9): Use two offset sgRNAs with Cas9 nickase (D10A) to create staggered nicks, requiring two independent binding events for a DSB, dramatically improving specificity.
For Base Editing
- Narrow-Window Editors: Use engineered deaminases with narrower activity windows (e.g., YE1-CBE) to reduce bystander edits (conversion of non-target Cs within the window).
- Gam Protein Co-expression: Fuse or co-express the bacteriophage protein Gam, which binds DSB ends and inhibits NHEJ, reducing indel formation from the nCas9 nick.
- Temporal Control: Use chemically inducible or light-inducible systems to express base editors transiently, limiting off-target activity.
Table 2: Specificity-Enhancing Tools and Their Applications
| Tool/Strategy | Best Suited For | Key Mechanism | Impact on Efficiency |
|---|---|---|---|
| SpCas9-HF1 | CRISPR-Cas9 (NHEJ/HDR) | Reduced non-specific binding | Slight decrease |
| Paired Nickases | CRISPR-Cas9 (HDR focus) | Requires two binding events | Moderate decrease |
| YE1-CBE Variant | Base Editing (CBE) | Narrower deamination window | Moderate decrease |
| Gam Protein Fusion | Base Editing | Inhibits NHEJ at nick site | Increases product purity |
| Chemical Induction | Both systems | Limits editor lifetime | Controllable, often lower |
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Advanced Genome Editing in Crops
| Reagent / Material | Supplier Examples | Function in Experiment |
|---|---|---|
| High-Fidelity SpCas9 Expression Vector | Addgene (pKIR1.1-HF), Tsingke Bio | Provides the nuclease with reduced off-target binding for clean edits. |
| ABE8e & evoFERNY-CBE Plasmids | Addgene, Miao Lab Vectors | State-of-the-art base editors for higher efficiency and purity in plants. |
| NHEJ Inhibitor (SCR7, NU7026) | Sigma-Aldrich, Tocris | Shifts repair balance from NHEJ to HDR in CRISPR-Cas9 experiments. |
| Plant Protoplast Isolation Kit | CPEC, Sigma | Enables high-efficiency transient delivery of RNP complexes for rapid testing. |
| Droplet Digital PCR (ddPCR) Supermix | Bio-Rad | Allows absolute, sensitive quantification of editing efficiency and HDR events. |
| Next-Generation Sequencing Kit (Amplicon) | Illumina, PacBio | For deep sequencing of target loci to assess on-target edits and off-target effects. |
| Synchronous Plant Cell Culture Reagents | Phytotech Labs | Chemicals like aphidicolin for cell cycle synchronization to boost HDR. |
| Agrobacterium Strain LBA4404/pVS1 | Invitrogen, lab stocks | Robust strain for stable transformation of a wide range of crop species. |
In crops research, the choice between traditional CRISPR-Cas9 and base editing is dictated by the desired outcome. For gene knock-outs, high-efficiency NHEJ-based Cas9 remains optimal. For precise single-nucleotide changes (SNPs) responsible for agronomic traits, base editors offer superior efficiency and product purity without DSBs. The strategies outlined—from HDR enhancement protocols and editor variant selection to specificity-focused designs—provide a roadmap for researchers to maximize on-target activity. As the field evolves, integrating these approaches with novel delivery and regulation systems will be key to developing next-generation, precision-edited crops.
Addressing PAM Sequence Limitations and Target Site Availability
The comparative efficacy of Base Editing (BE) versus Traditional CRISPR-Cas9 in crop genome engineering hinges significantly on two interconnected constraints: Protospacer Adjacent Motif (PAM) sequence limitations and target site availability. Traditional Cas9 nucleases (e.g., SpCas9) require a canonical NGG PAM, which restricts the genomic loci that can be targeted. This limitation directly impacts the development of crops with precise, trait-specific edits. Base editors, which are fusion proteins of a Cas9 nickase and a deaminase, inherit the PAM requirement of their associated Cas protein, thereby facing the same fundamental targeting constraint. This technical guide explores current strategies to overcome these limitations, thereby expanding the editable genomic space for crop improvement. The ability to access a wider array of target sites is a critical determinant in choosing between BE for precise single-nucleotide polymorphisms (SNPs) and traditional Cas9 for knock-outs, especially for polygenic traits in complex crop genomes.
Quantitative Data on PAM Specificities & Targeting Scope
Recent data on engineered Cas variants and their implications for crop targeting are summarized below.
Table 1: PAM Specificities and Theoretical Targeting Scope in a Model Plant Genome
| Cas Protein / Variant | PAM Requirement | Source/Origin | Theoretical Targeting Density (sites per kb)* in Arabidopsis thaliana (Chromosome 1) | Primary Application Context |
|---|---|---|---|---|
| SpCas9 (Wild-type) | 5'-NGG-3' | S. pyogenes | ~1 site / 8 bp | Traditional KO, BE, PE |
| SpCas9-NG | 5'-NG-3' | Engineered from SpCas9 | ~1 site / 4 bp | BE for relaxed PAM |
| xCas9(3.7) | 5'-NG, GAA, GAT-3' | Engineered from SpCas9 | ~1 site / 3.2 bp | Broad PAM BE |
| SpRY | 5'-NRN > NYN-3' | Engineered from SpCas9 | ~1 site / 1.2 bp | Near-PAMless targeting |
| ScCas9 | 5'-NNG-3' | S. canis | ~1 site / 5 bp | BE with compact size |
| enAsCas12a | 5'-TTTV-3' | Engineered from Acidaminococcus sp. | ~1 site / 16 bp | Traditional KO, BE (AT-rich) |
| CjCas9 | 5'-NNNNRYAC-3' | Campylobacter jejuni | ~1 site / 32 bp | Viral vector delivery |
| Data compiled from recent literature (2023-2024). Targeting density is a theoretical calculation based on PAM occurrence in the specified sequence. |
Table 2: Comparison of Base Editor Systems Used in Crops (2020-2024)
| Base Editor System | Core Components | PAM Dependency | Edit Type | Reported Efficiency Range in Major Crops | Key Limitation |
|---|---|---|---|---|---|
| ABE8e | SpCas9-NG-nickase + TadA-8e | NG | A•T to G•C | Rice: 10-80% (depending on target) | Off-target RNA editing |
| APOBEC-nCas9-UGI | nSpCas9 + rAPOBEC1 | NGG | C•G to T•A | Wheat: 5-40% | Sequence context (5’TC motif) |
| SpRY-ABE8e | SpRY-nickase + TadA-8e | NRN/NYN | A•T to G•C | Tomato protoplasts: up to 45% | Increased DNA off-targets |
| enAsCas12a-ABE | enAsCas12a-nickase + TadA-8e | TTTV | A•T to G•C | Rice: 20-70% | Larger size, delivery challenge |
| Target-AID | nSpCas9 + PmCDA1 | NGG | C•G to T•A | Maize: 1-30% | High indel by-product |
Experimental Protocols for Evaluating New PAM Specificities in Crops
Protocol 3.1:In PlantaPAM Profiling for Novel Cas Variants
Objective: To empirically determine the functional PAM preferences of a newly engineered Cas nuclease or base editor in a crop system. Materials: Agrobacterium strain (e.g., LBA4404), plant expression vectors (pCambia series), target crop seedlings (e.g., rice Nipponbare), high-throughput sequencing platform. Method:
- Library Construction: Synthesize a randomized PAM library (e.g., NNNNNN) flanking a constant protospacer sequence and clone it into a plant transformation vector containing a GFP reporter gene, disrupting its coding sequence.
- Plant Transformation: Introduce the PAM library vector and a second vector expressing the novel Cas/BE protein into crop explants via Agrobacterium-mediated transformation.
- Selection & Sequencing: Harvest transformed tissue (e.g., calli), isolate genomic DNA, and amplify the integrated PAM library region via PCR. Perform deep sequencing (Illumina MiSeq) on both the input plasmid library and the genomic DNA output.
- Bioinformatic Analysis: Align sequences and calculate the enrichment/depletion of each PAM sequence in the output relative to the input. Generate a sequence logo (WebLogo) to visualize the validated functional PAM.
Protocol 3.2: High-Throughput Evaluation of Base Editor Targetability in a Crop Genome
Objective: To assess the functional availability of target sites for a given base editor across a genomic region of interest (e.g., a gene family). Materials: Prime editing or BE guide RNA library, pooled oligo synthesis, plant protoplasts or stable transformation system, NGS reagents. Method:
- gRNA Library Design: Design 100-500 gRNAs tiling across all genes in a family, considering both canonical and relaxed PAM variants. Include both sense and antisense strands.
- Multiplexed Delivery: Clone the pooled gRNA library into an appropriate expression system. Deliver the BE plasmid and the gRNA pool into crop protoplasts via PEG-mediated transfection or into callus via biolistics.
- DNA Extraction & Amplicon Sequencing: After 5-7 days, extract genomic DNA. Amplify target regions using indexed primers and perform deep sequencing.
- Analysis of Editing Efficiency: Align reads to the reference genome. Quantify base conversion percentages and indel frequencies at each target site. Correlate high-efficiency sites with sequence context features (e.g., local chromatin accessibility from public ATAC-seq data).
Visualization of Strategies and Workflows
Title: Strategies to Overcome PAM Limitations
Title: Decision Workflow for Target Site Selection
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Addressing PAM/Targeting Limitations
| Reagent / Material | Supplier Examples | Function in Research |
|---|---|---|
| SpCas9-NG & SpRY Expression Plasmids | Addgene (#137982, #139998), TaKaRa | Provide the core nuclease with relaxed PAM specificity for base editor fusion or traditional editing. |
| Modular Base Editor Cloning Kit (Golden Gate) | VectorBuilder, Twist Bioscience | Enables rapid assembly of custom BE using different deaminases and Cas variants. |
| Plant Codon-Optimized ABE8e & high-activity CBE | ABRC, Academia Sinica seed stocks | Ready-to-use plasmids for high-efficiency A-to-G or C-to-T editing in crops. |
| Arabidopsis U6-26 & Rice U3 snRNA Promoter Vectors | Addgene, personal labs | Vectors for expressing gRNA arrays in specific plant systems. |
| Plant in vivo PAM Screening Kit | Custom synthesis (IDT, GenScript) | Pre-made randomized PAM library constructs for empirical PAM determination in plants. |
| Chloroplast Transformation Kit (for bypassing nuclear delivery) | Bio-Rad, lab-specific protocols | Alternative delivery method to overcome size limitations of large Cas variants. |
| Nucleofection Reagent for Protoplasts | Lonza, Bio-Rad | Enables high-throughput transfection of BE/gRNA libraries for multiplex testing. |
| NGS-based Off-Target Analysis Kit (GUIDE-seq, CIRCLE-seq) | Integrated DNA Technologies | Critical for profiling the specificity of novel Cas/BE variants before plant regeneration. |
| Plant Hi-C or ATAC-seq Database Access | PlantDHS, NCBI SRA | In silico resource for predicting open chromatin regions more accessible to BE machinery. |
Optimizing Delivery and Regeneration for Stubborn Crop Species
This technical guide examines the pivotal challenge of transforming recalcitrant crop species, positioning novel base editing systems as a transformative solution when integrated with advanced delivery and regeneration protocols. Within the broader thesis contrasting base editing with traditional CRISPR-Cas9, we argue that the precision, reduced DNA damage, and simplified repair outcomes of base editors are particularly advantageous for species where cellular stress from double-strand breaks (DSBs) cripples regeneration. This document provides a contemporary, data-driven framework for researchers to overcome these barriers.
Traditional CRISPR-Cas9 relies on generating DSBs, repaired by error-prone non-homologous end joining (NHEJ) or homology-directed repair (HDR). In many agronomically vital perennial crops (e.g., cocoa, coffee, many fruit trees), DSB-induced stress exacerbates low transformation efficiency and somaclonal variation. Base editing—catalyzing direct, irreversible chemical conversion of one base pair to another (C•G to T•A or A•T to G•C) without a DSB—presents a paradigm shift. It minimizes genomic shock, making it ideally suited for stubborn species where the primary bottleneck is successful regeneration of edited, fertile plants.
Quantitative Landscape: CRISPR-Cas9 vs. Base Editing Efficiencies
Recent studies (2023-2024) highlight the differential performance of these systems in model and challenging crops.
Table 1: Comparative Editing & Regeneration Efficiencies in Selected Crops
| Crop Species (Stubborn Status) | Editing System | Target Gene | Max Editing Efficiency (% in T0) | Regeneration Rate of Edited Cells (%) | Key Delivery Method | Reference Year |
|---|---|---|---|---|---|---|
| Rice (Model) | CRISPR-Cas9 (NHEJ) | OsPDS | 90% | 70% | Agrobacterium | 2022 |
| Rice (Model) | Adenine Base Editor (ABE) | OsACC | 50% | 65% | Agrobacterium | 2023 |
| Apple (Recalcitrant) | CRISPR-Cas9 (HDR) | DIPM-4 | 1.2% | <5% | Particle Bombardment | 2021 |
| Apple (Recalcitrant) | Cytosine Base Editor (CBE) | Ppo | 28% | 22% | RNP Electroporation | 2023 |
| Cacao (Highly Recalcitrant) | CRISPR-Cas9 (NHEJ) | TcMLO | 15% (in callus) | 0% (no shoots) | Agrobacterium | 2020 |
| Cacao (Highly Recalcitrant) | ABE8e | TcALS | 41% (in callus) | 15% (shoot formation) | Agrobacterium + VNs | 2024 |
| Coffee Arabica (Recalcitrant) | CBE (APOBEC-nCas9) | CaMXMT1 | 3-11% (stable plants) | 1.2% | Agrobacterium | 2023 |
Key Insight: The regeneration rate of edited cells is consistently higher for base editing approaches in stubborn species, directly linking reduced DSB formation to successful tissue culture progression.
Core Experimental Protocols
Protocol 3.1: Optimized Ribonucleoprotein (RNP) Delivery for Protoplasts of Woody Species
This method avoids vector DNA integration and reduces off-target plasmid exposure.
- Protoplast Isolation: Incubate 1g of juvenile, sterile leaf tissue in enzyme solution (2% Cellulase R10, 0.5% Macerozyme R10, 0.4M mannitol, 20mM KCl, 20mM MES pH 5.7, 10mM CaCl₂, 0.1% BSA) for 16h in dark with gentle shaking.
- RNP Complex Formation: For a CBE targeting a single site, combine 10µg of purified nCas9-cytidine deaminase protein with 5µg of chemically synthesized sgRNA in a 1:2 molar ratio. Incubate in 30µL NEBuffer 3.1 at 25°C for 15 min.
- Electroporation: Wash 2x10⁵ protoplasts in electroporation buffer (0.4M mannitol, 5mM MES, 15mM MgCl₂). Resuspend in 100µL buffer with RNP complexes. Electroporate using a square-wave protocol (250V, 20ms pulse length).
- Culture & Analysis: Transfer to dark, 24°C for 48h in regeneration medium. Extract genomic DNA from microcalli (3-7 days) for PCR and high-throughput sequencing to assess base edit efficiency before initiating differentiation.
Protocol 3.2:Agrobacterium-Mediated Transformation Enhanced with Virulence Gene Inducers for Embryogenic Callus
This protocol boosts T-DNA delivery in stubborn monocots and dicots.
- Strain & Vector Preparation: Use Agrobacterium tumefaciens strain EHA105 harboring a base editor binary vector (e.g., pABE8e-U6-sgRNA) and a visual marker (e.g., GFP). Grow to OD₆₀₀ = 0.6 in induction medium (IM) containing 200µM acetosyringone (AS).
- Co-cultivation with Chemical Enhancers: Submerge embryogenic calli (200mg) in the bacterial suspension for 20 min. Blot dry and co-culture on filter paper over solid co-cultivation medium (CCM) supplemented with 200µM AS and 100µM melatonin (or 10µM lipo-oligosaccharide VNs) for 72h in dark at 22°C.
- Resting & Selection: Transfer calli to a resting medium containing 300mg/L timentin for 5 days (no selection). Subsequently, transfer to selection medium containing appropriate antibiotics/herbicides.
- Regeneration & Screening: Regenerate shoots on medium with cytokinin (e.g., 2mg/L 6-BA) and then root on auxin medium (e.g., 0.2mg/L NAA). Screen regenerated plantlets via PCR/RFLP and Sanger sequencing tracing deaminase-induced edits.
Visualizing Key Pathways and Workflows
Diagram Title: CRISPR-Cas9 vs. Base Editing Molecular Pathways and Stress Outcomes
Diagram Title: Optimized Base Editing Workflow for Stubborn Crops
The Scientist's Toolkit: Essential Research Reagents & Solutions
Table 2: Key Reagents for Delivery & Regeneration of Stubborn Crops
| Reagent/Solution | Category | Function & Rationale |
|---|---|---|
| Purified nCas9-Deaminase Protein | Enzyme | Enables RNP assembly for DNA-free, transient base editor delivery, reducing off-target integration and cellular load. |
| Acetosyringone (AS) | Chemical Inducer | A phenolic compound essential for inducing Agrobacterium vir genes, critical for efficient T-DNA transfer to plant cells. |
| Lipo-oligosaccharides (e.g., VNs) | Biological Enhancer | Specific molecules that mimic plant signals, hyper-activating Agrobacterium virulence and improving T-DNA delivery to recalcitrant tissues. |
| Melatonin | Antioxidant/Culture Additive | Reduces oxidative stress during co-cultivation and regeneration, improving cell viability and edited callus recovery. |
| Cellulase R10 / Macerozyme R10 | Enzyme Cocktail | High-purity enzymes for reliable and gentle isolation of viable protoplasts from tough plant tissues. |
| Mannitol | Osmolyte | Maintains osmotic pressure in protoplast and transformation buffers, preventing cell lysis. |
| Thidiazuron (TDZ) | Synthetic Cytokinin | Potent plant growth regulator used at low concentrations to induce shoot organogenesis in stubborn species. |
| Gelrite | Gelling Agent | Preferred over agar for regeneration media; clearer, purer, and allows better nutrient diffusion for sensitive tissues. |
| Next-Generation Sequencing (NGS) Kit | Analysis | For deep amplicon sequencing to quantify base editing efficiency and profile potential off-target effects at high resolution. |
The integration of base editing technology with the delivery and regeneration protocols detailed herein represents the most promising avenue for functional genetics and trait development in stubborn crop species. By circumventing the DSB bottleneck, base editors align with the physiological constraints of these plants. Future convergence with developmental regulator-assisted transformation (e.g., Baby boom/Wuschel overexpression) and nanoparticle-based delivery will further democratize genetic access. This approach validates the core thesis: for crop improvement beyond model species, base editing is not merely an alternative but a necessary evolution from traditional CRISPR-Cas9, fundamentally changing the equation between genetic precision and organismal regenerability.
The application of genome editing in crops research has evolved significantly from traditional CRISPR-Cas9 systems to more precise base editing platforms. While traditional Cas9 induces double-strand breaks (DSBs) repaired by error-prone non-homologous end joining (NHEJ) or homology-directed repair (HDR), base editors directly convert one target DNA base into another without creating DSBs, thereby reducing indels. However, both systems are prone to unintended edits, including off-target mutations, bystander edits, and unpredictable byproducts. This guide provides a technical framework for the systematic analysis and filtering of these unintended edits, a critical step for validating edits intended for crop improvement and therapeutic applications.
Quantitative Comparison of Unintended Edits: Base Editors vs. Cas9
Table 1: Frequency and Spectrum of Unintended Edits by System
| Edit Type | Traditional CRISPR-Cas9 (NHEJ) | Cytosine Base Editor (CBE) | Adenine Base Editor (ABE) |
|---|---|---|---|
| Primary Intended Outcome | Knockout via indels | C•G to T•A conversion | A•T to G•C conversion |
| Off-Target Frequency* | 0.1% - 50% (sgRNA-dependent) | 0.01% - 10% | 0.001% - 2% |
| Common Byproducts | Large deletions, translocations | C-to-T at bystander Cs, C-to-G/A (rarer) | A-to-G at bystander As, A-to-C/T (rarer) |
| Indel Formation | High (primary mechanism) | Low but detectable | Very Low |
| Scope of Detection | Genome-wide (WGS) | Local window (amplicon-seq) | Local window (amplicon-seq) |
*Frequencies are highly variable and depend on delivery method, cell type, and target locus. Data compiled from recent studies (2023-2024).
Table 2: Common Detection Methods and Their Resolution
| Method | Detectable Edit Types | Sensitivity | Throughput | Cost |
|---|---|---|---|---|
| Whole Genome Sequencing (WGS) | All SNVs, indels, structural variants | ~1% VAF | Low | High |
| Amplicon-Seq (NGS) | SNVs, small indels in target region | <0.1% VAF | Medium | Medium |
| Digenome-seq | Cas9 & BE off-target cleavage sites in vitro | High | High | Low |
| CIRCLE-seq | Genome-wide off-target sites for nucleases & BEs | High | High | Medium |
| RNA-seq | Transcriptomic effects, aberrant splicing | NA | Medium | Medium |
Experimental Protocols for Byproduct Analysis
Protocol 3.1: High-Throughput Amplicon Sequencing for On-Target Byproduct Characterization
- Objective: Quantify editing efficiency and identify bystander edits within the editing window.
- Materials: Edited plant tissue genomic DNA, high-fidelity PCR master mix, barcoded NGS primers.
- Steps:
- Primer Design: Design primers ~100-150 bp flanking the target site.
- PCR Amplification: Perform two-step PCR. First PCR amplifies target region. Second PCR adds Illumina adapters and sample barcodes.
- Library Purification & Pooling: Clean PCR products with magnetic beads, quantify, and pool equimolarly.
- Sequencing: Run on Illumina MiSeq or NovaSeq (2x250 bp recommended).
- Analysis: Use CRISPResso2 or BE-Analyzer to align reads to reference, calculate editing efficiency, and quantify base substitutions at each position.
Protocol 3.2: CIRCLE-seq for Genome-Wide Off-Target Discovery
- Objective: Identify potential off-target sites for a given sgRNA in the relevant genome.
- Materials: High molecular weight genomic DNA, Cas9 or BE protein, sgRNA, T4 DNA Polymerase, Circligase.
- Steps:
- Genomic DNA Fragmentation & End Repair: Shear DNA, repair ends to create 5’ phosphorylated, blunt-ended fragments.
- Circularization: Ligate fragments into circles using Circligase. Linear, un-circularized DNA is degraded by Plasmid-Safe ATP-Dependent DNase.
- In Vitro Cleavage/Deamination: Incubate circularized DNA with pre-assembled RNP (Cas9/sgRNA or BE/sgRNA). For BEs, this step creates deaminated bases at off-target sites.
- Library Preparation: Linearize cleaved/deaminated circles, add adapters, and amplify for NGS.
- Bioinformatics: Map sequencing reads to reference genome. Breaks or mis-matches indicate potential off-target sites for validation.
Visualization of Workflows and Pathways
Diagram 1: Unintended Edits Analysis & Filtering Workflow
Diagram 2: Base Editor Action & Bystander Edits
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Byproduct Analysis
| Reagent / Kit Name | Function & Application | Key Consideration |
|---|---|---|
| NEXTflex PCR-Free DNA Seq Kit (Bioo) | Prepares high-quality WGS libraries without PCR bias, critical for accurate variant calling. | Essential for detecting large structural variants and unbiased SNV/indel counts. |
| Q5 High-Fidelity DNA Polymerase (NEB) | Ultra-high-fidelity PCR for amplifying target loci for amplicon-seq with minimal introduced errors. | Low error rate is non-negotiable for detecting low-frequency (<0.5%) edits. |
| CIRCLE-seq Kit (ToolGen) | Streamlined, optimized kit for performing CIRCLE-seq to identify genome-wide off-target sites in vitro. | Reduces protocol complexity and improves reproducibility compared to in-house. |
| NEBNext Ultra II FS DNA Library Kit | Used in CIRCLE-seq and other NGS preps for fast, efficient library construction from low-input DNA. | Includes fragmentation and end-prep in a single tube. |
| CRISPResso2 (Software) | A standard bioinformatics pipeline for quantifying genome editing outcomes from NGS amplicon data. | Supports base editor analysis (BE-Analyzer mode) and pooled screening. |
| Beacon Designer (Premier Biosoft) | Designs highly specific primers and probes for target amplification and validation (e.g., for ddPCR). | Avoids secondary structures and SNP sites in primer regions. |
| Alt-R S.p. Cas9 Nuclease V3 (IDT) | High-activity, high-purity Cas9 protein for in vitro off-target assays (e.g., CIRCLE-seq) or RNP delivery. | Recombinant protein quality impacts cleavage efficiency and background noise. |
| Guide-it Long-range PCR Kit (Takara) | Amplifies large genomic fragments for validating potential large deletions or translocations. | Optimized for amplifying complex, GC-rich plant genomic regions. |
Side-by-Side Analysis: Precision, Scope, and Regulatory Pathways for Crop Innovation
This technical guide provides a quantitative framework for evaluating base editing against traditional CRISPR-Cas9 nuclease systems in crop genome engineering. Within the broader thesis that base editing offers a more precise and predictable route to crop improvement, we dissect core performance metrics—editing efficiency, product purity (indel frequency), and experimental throughput—using current data and standardized protocols.
CRISPR-Cas9 nucleases create double-strand breaks (DSBs), relying on endogenous repair pathways (non-homologous end joining, NHEJ; homology-directed repair, HDR) to generate edits. This often results in a mixture of desired mutations and stochastic indels. Base editors (BEs), fusion proteins of a catalytically impaired Cas9 and a deaminase enzyme, directly convert one base pair to another (C•G to T•A or A•T to G•C) without inducing DSBs, theoretically yielding higher product purity.
Core Metrics: Definitions and Measurement
- Editing Efficiency: The percentage of target alleles that contain any intended edit within the target window. Measured via next-generation sequencing (NGS) of PCR amplicons.
- Product Purity: The percentage of edited alleles that contain only the desired base change(s) without any concomitant insertions, deletions, or other bystander edits. A key advantage postulated for base editors.
- Throughput: The number of discrete genetic loci or plant lines that can be successfully edited within a given experimental timeframe and budget. Encompasses vector construction, delivery, and screening.
Quantitative Data Comparison
The following tables consolidate recent (2022-2024) experimental data from studies in rice (Oryza sativa), wheat (Triticum aestivum), and tomato (Solanum lycopersicum).
Table 1: Average Performance Metrics in Model Crops
| System | Crop | Avg. Editing Efficiency (%) | Avg. Product Purity (%) | Avg. Indel Frequency (%) | Primary Delivery Method |
|---|---|---|---|---|---|
| CRISPR-Cas9 Nuclease | Rice | 15-40 | 10-30* | 60-85 | Agrobacterium-mediated |
| Cytosine Base Editor (CBE) | Rice | 20-70 | 50-90 | 1-10 | Agrobacterium-mediated |
| Adenine Base Editor (ABE) | Rice | 10-50 | 40-90 | 1-5 | Agrobacterium-mediated |
| CRISPR-Cas9 Nuclease | Wheat | 5-25 | 5-20* | 70-95 | Biolistics |
| CBE | Wheat | 5-30 | 30-80 | 1-15 | Biolistics |
| ABE | Wheat | 5-20 | 30-75 | 1-10 | Biolistics |
*For Cas9, product purity refers to alleles with precise HDR-mediated edits, which are rare in plants. Most edits are imprecise NHEJ outcomes.
Table 2: Throughput and Practical Considerations
| Parameter | CRISPR-Cas9 Nuclease | Base Editing (CBE/ABE) |
|---|---|---|
| Vector Construction | Standardized, high-throughput cloning (Golden Gate). | More complex due to larger effector size; optimized toolkits available. |
| Target Sequence Requirement | Requires NGG PAM (SpCas9). Flexible. | Constrained by deaminase window position relative to PAM. |
| Mutation Types | Knock-outs, large deletions, HDR-mediated substitutions (inefficient). | Transition mutations only: C>T or A>G. |
| Screening Complexity | High: Must screen for rare HDR events amid high indel background. | Lower: Focus on identifying precise point mutations. |
| Multiplexing Potential | High for creating knock-outs. | Moderate, limited by compatible PAMs and editing windows. |
Experimental Protocols for Key Comparisons
Protocol 4.1: Side-by-Side Evaluation of Editing Outcomes
Objective: Quantify efficiency and purity of a target C-to-T change using CBE vs. Cas9+HDR.
- Vector Design: For the same target genomic locus, construct (a) a CBE plasmid (e.g., A3A-PBE) and (b) a Cas9 + ssODN HDR donor plasmid containing the desired T.
- Plant Transformation: Transform competent rice calli (indica cv. Kitake) via Agrobacterium (strain EHA105) with each construct. Include nuclease-only control.
- Regeneration & Sampling: Regenerate T0 plants. Isolate genomic DNA from leaf tissue of 20 independent lines per construct.
- Amplicon Sequencing: PCR-amplify target region with barcoded primers. Perform 300bp paired-end sequencing on an Illumina MiSeq.
- Data Analysis: Use CRISPResso2 or similar. Quantify:
- Total Editing (%) = (1 - (read count of wild-type / total read count)) x 100.
- Product Purity (%) = (read count with only the exact C>T change / total edited read count) x 100.
- Indel Frequency (%) = (read count with indels / total read count) x 100.
Protocol 4.2: High-Throughput Phenotyping Workflow
Objective: Assess the throughput from transformation to identified mutant.
- Pooled Transformation: Use a Golden Gate-assembled library of 50 sgRNAs (for Cas9 knock-out) or 50 guide RNAs targeting specific bases (for BE) in a single, high-efficiency transformation of wheat via biolistics.
- Bulk Regeneration: Regenerate plants under selection without initial molecular screening.
- Leaf-Pool DNA Extraction: At the T1 seedling stage, pool leaf punches from 100 plants per construct. Extract bulk DNA.
- Amplicon-Seq Library Prep: Perform a two-step PCR: (i) locus-specific amplification, (ii) addition of Illumina indices.
- High-Throughput Sequencing: Run on an Illumina NextSeq 2000 (P2 flow cell, 300 cycles).
- Bioinformatic Deconvolution: Map reads to reference, identify edits and their frequencies per target site. Correlate high-frequency edits with phenotypic data from the pooled plants.
Visualization of Key Concepts and Workflows
Diagram 1: Base Editing vs. CRISPR-Cas9 Core Mechanism
Diagram 2: Standard Workflow for Quantitative Comparison
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Comparative Editing Studies
| Reagent / Material | Function in Experiment | Key Consideration for Comparison |
|---|---|---|
| Modular Cloning Toolkit (e.g., MoClo Plant, Golden Gate) | Enables rapid, standardized assembly of Cas9, BE, and sgRNA expression cassettes. | Essential for constructing isogenic vectors differing only in the effector. |
| High-Efficiency Agrobacterium Strain (e.g., EHA105, LBA4404) | Delivery of T-DNA carrying editing machinery into plant cells. | Strain choice impacts transformation efficiency, a key throughput variable. |
| Chemical Selection Agents (e.g., Hygromycin, Bialaphos) | Selects for transformed plant tissue during callus phase. | Must be optimized for crop species to avoid bottlenecks. |
| NGS Amplicon-EZ Service | Provides end-to-end solution from primer design to sequencing data for target loci. | Critical for obtaining unbiased, quantitative editing data for both systems. |
| CRISPResso2 / BEBanalyzer Software | Specialized bioinformatics tools to quantify editing outcomes from NGS data. | BEBanalyzer is optimized for base editor output (e.g., analyzes deamination windows). |
| Reference Genomic DNA (Wild-type plant) | Serves as a negative control for sequencing and for standard curve generation in ddPCR assays. | Ensures accurate background subtraction and variant calling. |
| Synthetic ssODN HDR Donor | Template for precise repair with Cas9. Used in Cas9+HDR protocol arm. | Design must avoid Cas9 cleavage; efficiency is typically very low in plants. |
Quantitative comparisons substantiate the thesis that base editing provides superior product purity and predictability for installing point mutations in crops, directly addressing a major limitation of traditional CRISPR-Cas9. While editing efficiency is comparable or situationally superior, the defining advantage is the drastic reduction in indel byproducts. Throughput remains similar, constrained by transformation and regeneration. Future developments in prime editing, improved deaminases with narrower windows, and virus-induced genome editing (VIGE) will further refine these metrics, solidifying the role of precision base editing in the next generation of crop improvement.
The advent of CRISPR-Cas9 revolutionized genetic engineering, offering unprecedented precision in creating double-strand breaks (DSBs) at target genomic loci. However, the reliance on endogenous DNA repair pathways—predominantly error-prone non-homologous end joining (NHEJ)—introduces stochastic indels, limiting the predictability and scope of editable mutations. Base editing (BE) emerges as a transformative alternative, enabling direct, irreversible conversion of one target DNA base pair to another without inducing DSBs. Within crop research, the choice between traditional CRISPR-Cas9 and BE systems is dictated by the desired mutation profile, the need for precision, and the biological constraints of the plant system. This guide provides a technical assessment of the flexibility and limitations inherent to each platform, focusing on the scope of mutations achievable.
System Architectures and Core Mechanisms
Traditional CRISPR-Cas9
The canonical Streptococcus pyogenes Cas9 (SpCas9) system functions as a RNA-guided endonuclease. The guide RNA (gRNA) directs Cas9 to a target sequence adjacent to a protospacer adjacent motif (PAM, typically 5'-NGG-3'). Cas9 creates a blunt-ended DSB 3 bp upstream of the PAM. The cellular repair outcomes define the scope of editable mutations:
- NHEJ: Often results in small insertions or deletions (indels) leading to frameshifts and gene knockouts.
- Homology-Directed Repair (HDR): Can introduce precise edits using a donor DNA template, but is inefficient in most plant cells, especially in non-dividing cells.
Base Editing Systems
Base editors are fusion proteins comprising a catalytically impaired Cas9 (nCas9) or Cas9 nickase (nicking one strand) linked to a nucleobase deaminase enzyme. They operate without creating DSBs, catalyzing chemical conversion of bases within a defined "activity window."
- Cytosine Base Editors (CBEs): Convert C•G to T•A pairs. Commonly use rat APOBEC1 deaminase with uracil glycosylase inhibitor (UGI) to prevent base excision repair.
- Adenine Base Editors (ABEs): Convert A•T to G•C pairs. Evolved from TadA deaminase.
- Dual Base Editors: Emerging systems capable of both C-to-T and A-to-G conversions.
Quantitative Comparison of Mutation Scope
Table 1: Comparative Scope of Editable Mutations
| Parameter | Traditional CRISPR-Cas9 (via NHEJ) | Traditional CRISPR-Cas9 (via HDR) | Cytosine Base Editor (CBE) | Adenine Base Editor (ABE) |
|---|---|---|---|---|
| Primary Edit Type | Random insertions/deletions (indels) | User-defined sequence changes | C•G to T•A point mutation | A•T to G•C point mutation |
| Theoretical Targetable Bases | ~1 in 8 bp (near NGG PAM) | ~1 in 8 bp (near NGG PAM) | ~1 in 25 bp within activity window* | ~1 in 25 bp within activity window* |
| Typical Editing Window | Cut site (~3-4 bp range) | Defined by donor template | ~5-nt window (positions 4-8, protospacer 1-based) | ~5-nt window (positions 4-8, protospacer 1-based) |
| Precision | Low (stochastic outcomes) | High (requires donor) | High (predictable point mutation) | High (predictable point mutation) |
| Efficiency in Plants | High (10-90% indels) | Very Low (<5% typically) | Moderate to High (up to 80% in ideal cases) | Moderate to High (up to 50% in ideal cases) |
| Primary Byproducts | Unwanted indels, large deletions | Unwanted indels, off-target integration | Undesired C-to-T conversions outside window, random indels (low), bystander edits | Undesired A-to-G conversions outside window, random indels (very low) |
| Key Limitation | Unpredictable outcomes; inefficient precise sequence insertion. | Extremely low efficiency in plants; requires donor design/delivery. | Cannot create all point mutations or insertions; bystander edits. | Cannot create all point mutations or insertions; limited to A-to-G. |
*Data synthesized from recent literature (2023-2024). *Efficiency and window position vary with editor architecture and target sequence.
Experimental Protocols for Assessing Mutation Scope
Protocol: Assessing CRISPR-Cas9 NHEJ Mutation Spectrum in Protoplasts
Objective: To characterize the diversity and frequency of indel mutations induced by NHEJ at a target locus. Materials: Plant protoplasts, SpCas9 expression vector, target-specific gRNA vector, PEG transformation reagents, DNA extraction kit, PCR primers flanking target. Method:
- Co-transform protoplasts with SpCas9 and gRNA plasmids via PEG-mediated transfection.
- Incubate for 48-72 hours under optimal culture conditions.
- Harvest protoplasts and extract genomic DNA.
- Amplify target locus by PCR using high-fidelity polymerase.
- Purify PCR product and subject to next-generation sequencing (NGS) using a 2x250 bp paired-end MiSeq run.
- Analyze sequencing reads using CRISPResso2 or similar tool to quantify indel percentages, size distribution, and sequence profiles.
Protocol: Quantifying Base Editing Efficiency and Purity in Stable Transgenic Lines
Objective: To determine the on-target base conversion efficiency and identify bystander edits within the activity window. Materials: Agrobacterium strain harboring BE expression vector, plant explants for transformation, selection agents, tissue culture media. Method:
- Generate stable transgenic plants via Agrobacterium-mediated transformation.
- Select regenerated T0 plants on appropriate antibiotic/herbicide.
- Isolate genomic DNA from leaf tissue of T0 plants.
- PCR-amplify the target region from edited and control plants.
- Clone the PCR products into a blunt-end sequencing vector. Transform E. coli and pick 20-50 individual colonies per plant for Sanger sequencing. Alternatively, perform deep amplicon sequencing.
- Align sequences to the reference. Calculate the percentage of reads with intended base conversion(s). Note the frequency of any unintended "bystander" edits (other C or A bases within the activity window that were also deaminated).
Signaling Pathways and Workflow Visualizations
Title: Decision Workflow: Choosing CRISPR-Cas9 vs Base Editing
Title: Core Mutagenic Pathways: CRISPR-Cas9 vs Base Editing
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Mutation Scope Analysis
| Reagent / Material | Function in Experiment | Key Consideration for Crop Research |
|---|---|---|
| High-Efficiency CRISPR-Cas9 Vector (e.g., pRGEB32, pCambia-Cas9) | Delivers SpCas9 and gRNA expression cassettes for plant transformation. | Choose species-specific promoters (e.g., ZmUbi for maize, AtU6 for Arabidopsis). |
| Modular Base Editor Vector (e.g., pnCsBE, pH-nCas9-PBE) | Delivers nCas9-deaminase fusion and gRNA. | Select editor variant (e.g., evoFERNY-CBE, ABE8e) for optimal window & efficiency. |
| Chemically Competent Agrobacterium (Strain EHA105, GV3101) | Mediates stable genomic integration of editing constructs via transformation. | Strain choice impacts host range and transformation efficiency. |
| PEG-Ca2+ Transfection Solution | Facilitates plasmid delivery into protoplasts for rapid transient assays. | Critical for pre-screening gRNA and editor efficiency before stable transformation. |
| Next-Generation Sequencing (NGS) Kit (Illumina MiSeq, amplicon-seq) | Enables deep sequencing of target loci to quantify edits, indels, and byproducts. | Required for unbiased analysis of mutation spectrum and off-target assessment. |
| CRISPR Analysis Software (CRISPResso2, BE-Analyzer, Cas-Analyzer) | Bioinformatics tool to process NGS data and calculate editing indices. | Essential for accurately quantifying complex editing outcomes from bulk sequencing. |
| Uracil DNA Glycosylase Inhibitor (UGI) | Component of CBE vectors; suppresses base excision repair of U:G intermediate. | Enhances C-to-T editing efficiency and purity by preventing reversion. |
| HDR Donor Template (ssODN, dsDNA with homology arms) | Provides repair template for precise sequence insertion via HDR. | Very low efficiency in crops; requires optimization of delivery and design. |
Within the broader thesis comparing base editing to traditional CRISPR-Cas9 in crop research, a critical focal point is the divergent regulatory pathways and safety assessments triggered by different types of genetic modifications. The Site-Directed Nuclease (SDN) classification system—particularly SDN-1 and SDN-2—serves as a cornerstone for global regulatory frameworks, directly impacting product development timelines, data requirements, and commercial viability. This guide provides a technical analysis of how emerging base editing techniques intersect with these established categories, shaping the regulatory and safety landscape for novel crop products.
The SDN Classification Framework: Technical Definitions
Regulatory agencies categorize genome-edited products based on the nature of the edit and the presence of exogenous DNA in the final product.
- SDN-1: Application of a nuclease (e.g., Cas9) to create a targeted double-strand break (DSB) repaired by the cell's own error-prone Non-Homologous End Joining (NHEJ) pathway. This results in small insertions or deletions (indels) that can disrupt a gene's function. No template DNA is provided.
- SDN-2: A DSB is repaired using an externally supplied, short DNA template (donor) via Homology-Directed Repair (HDR). This enables precise, targeted changes of a few nucleotides (point mutations) or short insertions. The final product typically contains no exogenous DNA.
- SDN-3: Insertion of a large DNA sequence (e.g., a transgene) via HDR using an external template. The final product contains stable integration of foreign DNA.
Base editing occupies a unique space, enabling precise, single-nucleotide changes without creating a DSB and, in most implementations, without requiring a donor template. This challenges the binary SDN-1/SDN-2 distinction.
Quantitative Comparison of Regulatory Triggers
Table 1: Regulatory Classification and Data Requirements for CRISPR Techniques
| Feature | Traditional CRISPR-Cas9 (SDN-1) | Traditional CRISPR-Cas9 (SDN-2) | Base Editing (CBE/ABE) | SDN-3 (Transgenic) |
|---|---|---|---|---|
| Mechanism | NHEJ repair of DSB | HDR repair of DSB with short donor | Direct chemical conversion of base pairs (No DSB) | HDR with large donor DNA |
| Exogenous DNA in Final Plant? | Typically No | Typically No | Typically No* | Yes |
| Typical Product | Gene knockout | Precise point mutation or short insert | Precise point mutation (C•G to T•A or A•T to G•C) | Novel gene insertion |
| Global Regulatory Trend | Often exempted or streamlined (Product-based) | Case-by-case; often regulated similar to SDN-1 | Under Review / Uncertain | Regulated as GMO (Process-based) |
| Key Safety Data Points | Off-target indels, genetic stability | Off-target edits, donor DNA integration, genetic stability | Off-target deamination (not DSBs), guide-independent edits, RNA/Protein off-targets | Full molecular characterization, allergenicity, toxicity, environmental impact |
Note: *While the editing machinery is transient, the possibility of integrating plasmid DNA used for delivery requires molecular characterization.
Experimental Protocols for Key Regulatory Assessments
Protocol: Comprehensive Off-Target Analysis for Base Editors
Objective: Identify and quantify unintended edits across the genome, a critical safety endpoint for regulators. Methodology:
- In Silico Prediction: Use tools like Cas-OFFinder to predict potential off-target sites with sequence homology to the sgRNA (allow 1-6 mismatches and/or bulges).
- Cell-Free / In Vitro Testing: Perform GUIDE-seq or CIRCLE-seq on genomic DNA extracts to empirically identify cleavable or editable sites in an unbiased manner.
- Whole-Genome Sequencing (WGS): Sequence multiple, independently derived, edited plant lines (T1 or later generations) to a high depth of coverage (≥50x). Use an isogenic unedited control.
- Bioinformatic Pipeline: Align sequences to reference genome. Call variants (SNPs, indels) using tools like GATK. Filter variants present only in edited lines and absent in the control. Cross-reference with in silico and in vitro prediction lists.
- Validation: Use targeted amplicon sequencing (Sanger or NGS) to confirm high-priority off-target candidates in biological replicates.
Protocol: Molecular Characterization to Confirm Absence of Exogenous DNA
Objective: Provide evidence that the final commercial product contains only the intended edit and no integrated vector or bacterial sequences. Methodology:
- Digital PCR (dPCR) or Quantitative PCR (qPCR): Design TaqMan assays targeting common vector backbone elements (e.g., antibiotic resistance genes, bacterial origins of replication).
- Southern Blot Analysis: Digest genomic DNA with restriction enzymes that do not cut within the target genomic locus but cut within the potential integrated plasmid. Use a labeled probe specific to the plasmid sequence. This detects large integrations.
- Long-Read Sequencing (PacBio, Oxford Nanopore): Sequence the entire edited locus and flanking regions to generate a contiguous haplotype, confirming the edit's context and the absence of interspersed foreign DNA.
Visualizing Regulatory Decision Pathways
Diagram Title: Regulatory Decision Tree for Edited Crops
Product Status and Global Regulatory Snapshot
Table 2: Global Regulatory Status for SDN-1/SDN-2/Base Edited Crops (Representative)
| Country/Region | Regulatory Basis | SDN-1 Product Status | SDN-2 / Base Editing Status (as of 2023-24) |
|---|---|---|---|
| United States | Product (APHIS SECURE Rule) | Several deregulated (e.g., CRISPR'd mushrooms, waxy corn) | Case-by-case. Base edited canola (Cibus) not regulated by USDA-APHIS. FDA review ongoing. |
| Argentina | Process, with resolution for SDN-1/2 | Streamlined notification if no novel combination of genetic material. | Base edited wheat (HB4 drought tolerance) approved. SDN-2 treated similarly to SDN-1. |
| European Union | Process (ECJ Ruling) | Regulated as GMO. | Regulated as GMO. Commission studying "New Genomic Techniques" proposal for potential reform. |
| Japan | Product (Case-by-Case) | Not regulated as GMO if no foreign DNA. | High-oleic acid base edited tomato commercialized (2021). Treated similarly to SDN-1. |
| Brazil | Process, with normative resolution | SDN-1 & SDN-2 exempt if no foreign DNA in final product. | Base edited soybean (higher oleic acid) approved. Falls under exemption. |
| India | Process (Proposed) | Under review. Draft rules suggest exemption for SDN-1/2 without foreign DNA. | Pending finalization of biosafety guidelines. |
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Regulatory-Grade Genome Editing Analysis
| Reagent / Solution | Function in Regulatory Experiments | Example Vendor/Kit |
|---|---|---|
| High-Fidelity Base Editor Plasmids | Minimizes guide-independent off-target deaminase activity. Critical for safety. | BE4max, ABE8e (Addgene) |
| Hypothetical Transient Delivery System | Achieves editing without DNA integration (e.g., ribonucleoprotein complexes). Reduces regulatory burden. | Alt-R S.p. Cas9 Nuclease V3 + crRNA (IDT) |
| Whole-Genome Sequencing Service | Provides gold-standard, unbiased off-target and molecular characterization data for regulatory dossiers. | Illumina NovaSeq, PacBio HiFi |
| Digital PCR (dPCR) Master Mix | Absolute quantification of residual vector backbone or edit zygosity with high precision for molecular characterization. | QIAcuity Digital PCR System (Qiagen) |
| GUIDE-seq or CIRCLE-seq Kit | Empirically identifies potential off-target sites for Cas9 or base editors in a genome-wide manner. | GUIDE-seq Kit (Integrated DNA Technologies) |
| Sanger Sequencing & Analysis Software | Validates on-target edit efficiency and screens for common off-target sites. | CRISP-ID, TIDE, or Synthego ICE Analysis |
Diagram Title: Components of a Safety Dossier
This technical guide outlines comprehensive validation frameworks essential for assessing genome-edited crops, with a focus on differentiating outcomes from base editing versus traditional CRISPR-Cas9 systems. The precision of base editors (BEs)—comprising cytosine base editors (CBEs) and adenine base editors (ABEs)—necessitates specific, nuanced validation protocols beyond those used for double-strand break (DSB)-dependent Cas9 editing. Rigorous validation across molecular, phenotypic, and biosafety domains is critical for research reproducibility and regulatory compliance.
Molecular Validation Assays
Molecular assays confirm the intended edit, identify off-target effects, and characterize editing efficiency.
Target Site Analysis
Protocol: Amplicon Sequencing for Edit Characterization
- Genomic DNA Extraction: Use a kit (e.g., DNeasy Plant Pro Kit) to extract high-quality gDNA from edited and wild-type control tissue.
- PCR Amplification: Design primers (~150-300bp amplicon) flanking the target site. Use a high-fidelity polymerase (e.g., Q5 Hot Start).
- Library Preparation & Sequencing: Purify PCR amplicons. Prepare sequencing libraries using a ligation-based kit (e.g., NEBNext Ultra II DNA Library Prep). Sequence on an Illumina MiSeq or NovaSeq platform (minimum 10,000x depth).
- Data Analysis: Use tools like CRISPResso2, BE-Analyzer, or custom pipelines to quantify:
- Editing efficiency (% of reads with edits)
- Edit type distribution (C•G to T•A, A•T to G•C, indels)
- Unintended bystander edits within the editing window.
Key Reagent Solutions:
- High-Fidelity Polymerase: Ensures accurate amplification without introducing errors.
- Illumina Sequencing Kit: Enables high-depth, quantitative sequencing of target loci.
- CRISPResso2 Software: Open-source tool for analyzing sequencing data of genome editing outcomes.
Off-Target Analysis
Protocol: Whole Genome Sequencing (WGS) for Unbiased Screening
- Sample Prep: Extract gDNA from edited and isogenic wild-type lines.
- Library & Sequencing: Prepare paired-end WGS libraries. Sequence to a minimum coverage of 30-50x on an Illumina platform.
- Variant Calling: Align reads to the reference genome (e.g., using BWA-MEM, GATK Best Practices). Call variants (SNVs, indels) comparing edited to wild-type.
- Off-Target Filtering: Filter variants against:
- In silico predicted off-target sites (from tools like Cas-OFFinder).
- Common lab strain variants.
- Focus on variants in coding/exonic regions and potential regulatory sequences.
Table 1: Comparison of Molecular Validation Outcomes: Cas9 vs. Base Editors
| Assay Parameter | Traditional CRISPR-Cas9 (NHEJ/HDR) | Base Editors (CBEs/ABEs) | Key Implication |
|---|---|---|---|
| Primary Edit Type | Indels (NHEJ) or donor template insertions (HDR) | Targeted point mutations (C>T, A>G) within window | BEs require SNV-specific detection, not indel assays. |
| Bystander Edits | Rare, adjacent to DSB | Common within deamination window (typically ~5nt window) | Must quantify all nucleotide changes in the window. |
| Off-Target Risk Locus | DSB-dependent indels at homologous sites | DSB-independent off-target SNVs; potential gRNA-independent RNA off-targets | WGS is preferred for BE off-target profiling over CIRCLE-seq. |
| Common Detection Method | T7E1 assay, Sanger sequencing, NGS | RFLP (if edit creates/disrupts site), Sanger sequencing, NGS required | NGS (amplicon-seq) is the gold standard for BEs. |
Phenotypic Validation Assays
These assays link the genotype to the expected functional trait.
Protocol: Controlled Environment Phenotyping for Herbicide Tolerance Use Case: Validation of a novel ALS gene base edit conferring imidazolinone tolerance.
- Plant Material: T2 generation homozygous edited lines, null segregant controls, wild-type.
- Growth Conditions: Randomized block design in growth chambers (controlled light, temp, humidity).
- Treatment: Apply field-relevant dose of imidazolinone herbicide at the 3-leaf stage. Include untreated controls.
- Data Collection: At 7, 14, and 21 days after treatment (DAT):
- Digital imaging for biomass estimation.
- Chlorophyll fluorescence (Fv/Fm) to assess photosynthetic health.
- Visual injury scoring on a standardized scale (0-100%).
- Plant height and leaf count.
- Statistical Analysis: ANOVA with post-hoc tests to separate means between edited, null segregant, and wild-type lines.
Diagram Title: Phenotyping Workflow for Herbicide Tolerance
The Scientist's Toolkit: Phenotyping & Molecular Analysis
| Item | Function in Validation |
|---|---|
| Growth Chambers | Provide standardized environmental conditions for reproducible phenotyping. |
| Hyperspectral Imaging System | Captures non-visible reflectance data correlating with plant health, biochemistry, and stress. |
| Chlorophyll Fluorometer | Measures photosynthetic efficiency (Fv/Fm), a sensitive indicator of abiotic stress. |
| Digital PCR System | Enables absolute, sequence-specific quantification of edit frequency without standard curves. |
| Sanger Sequencing Kit | For rapid, cost-effective initial confirmation of edits at target locus. |
Biosafety Validation Assays
Biosafety assays evaluate unintended compositional changes and genetic stability.
Compositional Analysis
Protocol: Proximate Analysis for Major Nutrients
- Sample Preparation: Harvest mature grain/leaf tissue from edited and control lines. Dry, mill to a fine, homogeneous powder.
- Macronutrient Analysis:
- Protein: Perform Dumas combustion or Kjeldahl method (AOAC Official Methods).
- Lipids: Use Soxhlet extraction with hexane.
- Ash: Incinerate sample in a muffle furnace at 550°C.
- Carbohydrates: Calculate by difference: 100% - (%moisture + %protein + %lipid + %ash).
- Fiber: Perform enzymatic-gravimetric methods (e.g., AOAC 991.43).
- Data Comparison: Compare composition of edited line to null segregant and historical wild-type ranges using equivalence testing (e.g., OECD consensus documents).
Genetic Stability and Inheritance
Protocol: Segregation Analysis and Southern Blotting
- Segregation Analysis: Grow T1/T2 populations from a primary edited event. Genotype ~100 individuals by PCR/sequencing.
- Calculate Segregation Ratio: Determine fit to Mendelian expectations (e.g., 3:1 for a heterozygous T0) using a chi-square test. Deviation suggests genetic instability.
- Southern Blot (If Transgenic):
- Digest gDNA (10-20µg) with appropriate restriction enzymes.
- Run on a 0.8% agarose gel, depurinate, denature, neutralize, and transfer to a nylon membrane.
- Hybridize with a non-radioactive DIG-labeled probe specific to the transgene or T-DNA border.
- Detect using anti-DIG antibodies and chemiluminescence. A simple, expected banding pattern confirms simple, stable integration.
Table 2: Biosafety Assay Framework for Base Edited vs. Cas9-Edited Crops
| Assay Category | Specific Test | Relevance for CRISPR-Cas9 (DSB) | Relevance for Base Editors (No DSB) |
|---|---|---|---|
| Molecular Characterization | Southern Blot / NGS for Integration | Critical. Confirms absence of vector backbone integration. | Less critical if delivered as RNP or transient DNA; still recommended for DNA delivery. |
| Genetic Stability | Mendelian Inheritance Analysis | Essential for both. Deviations may indicate complex inserts or lethality. | Essential. Ensures edit is stable through meiosis. |
| Compositional Analysis | Proximate & Key Antinutrients | Standard requirement to assess unintended pleiotropic effects. | Standard requirement; even precise edits may alter metabolic pathways. |
| Allergenicity/Toxicity | In silico Protein Comparison (FAO/WHO) | Required if novel protein sequence created (e.g., via HDR). | Highly Relevant. A single nucleotide change could create a novel immunogenic peptide. |
| Off-Target Assessment | WGS | Important to rule out large-scale rearrangements. | Critical to identify gRNA-independent and RNA off-target SNVs. |
Diagram Title: Safety Validation Paths for Base Editors vs. Cas9
A tiered validation framework—spanning molecular precision, phenotypic function, and biosafety—is non-negotiable for advancing genome-edited crops. While traditional CRISPR-Cas9 validation focuses on indels and integration events, base editing demands assays tailored for single-nucleotide resolution, bystander effects, and distinct off-target profiles. The protocols and comparisons outlined here provide a roadmap for researchers to robustly validate their editing outcomes, ensuring scientific rigor and supporting the path toward commercialization.
Modern crop improvement is driven by the need for precise genetic modifications to enhance yield, nutritional quality, and stress resilience. Two revolutionary technologies dominate this landscape: Traditional CRISPR-Cas9 and CRISPR-based Base Editing. Within the context of crop trait development, the selection between these tools is not trivial; it is a foundational strategic decision that dictates experimental design, regulatory pathway, and ultimate success.
Traditional CRISPR-Cas9 creates double-stranded DNA breaks (DSBs), relying on the cell's endogenous repair pathways—Non-Homologous End Joining (NHEJ) or Homology-Directed Repair (HDR)—to generate edits. This is powerful for gene knock-outs but is prone to unpredictable indels and is inefficient for precise point mutations.
CRISPR Base Editing directly converts one target DNA base pair into another without inducing a DSB. By fusing a catalytically impaired Cas protein (nickase) to a deaminase enzyme, base editors enable precise chemical conversion (e.g., C•G to T•A or A•T to G•C). This is transformative for installing or correcting single-nucleotide polymorphisms (SNPs) known to confer valuable agronomic traits.
Core Technology Comparison: Decision Matrices
Table 1: Fundamental Operational Characteristics
| Feature | Traditional CRISPR-Cas9 | Base Editing (e.g., CBEs, ABEs) |
|---|---|---|
| Primary Action | Induces Double-Strand Break (DSB) | Direct chemical conversion of one base to another. |
| DSB Formation | Yes (Catalytically active Cas9) | No (Uses catalytically dead/weakened Cas nickase). |
| Primary Outcome | Gene knockouts via indels (NHEJ). Precise edits possible via HDR but with low efficiency. | Precise point mutations (transition mutations: C→T, A→G). |
| Edit Precision | Low for precise point mutations; high for disruption. | Very high for targeted transitions within the editing window. |
| Common Repair Pathway Used | NHEJ (error-prone) or HDR (requires donor template). | DNA mismatch repair (MMR) or direct replication. |
| Byproduct Frequency | High (unwanted indels, large deletions). | Low, but potential for bystander edits within the activity window. |
| Optimal Use Case | Gene knockout, large deletions, sequence insertion via HDR. | Precise SNP introduction/correction, creating missense or nonsense mutations. |
Table 2: Performance Metrics in Model Crops (Recent Data)
| Crop | Target Gene / Trait | Tool | Efficiency (Edited Plants) | Precision (Desired Edit Only) | Key Reference |
|---|---|---|---|---|---|
| Rice | ALS (Herbicide Resistance) | Cas9-HDR | ~1-5% | Moderate (with HDR donor) | Huang et al., 2021 |
| Rice | ALS (Herbicide Resistance) | CBE (rAPOBEC1-Cas9n) | ~20-50% | High (>90% no indels) | Shimatani et al., 2017 |
| Wheat | LOX2 (Reduced Lipid Peroxidation) | Cas9-NHEJ | ~5-10% (biallelic knockout) | High for knockout | Sánchez-León et al., 2018 |
| Maize | Wx1 (High-Amylose Starch) | A•T to G•C Base Editor (ABE) | ~15% | High (clean point mutation) | Li et al., 2023 |
| Tomato | SP5G (Early Yield) | Cas9-NHEJ | ~30% (mutant lines) | High for knockout | Soyk et al., 2017 |
| Potato | ALS1 & PDS (Dual Herbicide Resistance) | CBE & ABE (Dual system) | ~40% (single base edit) | High (low indels) | Veillet et al., 2019 |
Table 3: Strategic Selection Guide Based on Trait Goal
| Your Crop Trait Goal | Recommended Tool | Rationale | Critical Considerations |
|---|---|---|---|
| Loss-of-function (Gene Knockout) | Traditional CRISPR-Cas9 | NHEJ efficiently creates disruptive indels. Simple, well-established. | Potential for off-target DSBs. Screen for large deletions. |
| Gain-of-function via precise SNP (e.g., herbicide-tolerant ALS) | Base Editor (CBE or ABE) | Directly installs known SNP without donor DNA or DSB. Higher efficiency than HDR. | Check if SNP is a C→T or A→G transition within base editor window. |
| Protein domain swap or large insertion (e.g., pathogen resistance domain) | Traditional CRISPR-Cas9 + HDR | Requires donor template to insert new sequence at DSB site. | Very low efficiency in plants; requires sophisticated enrichment strategies. |
| Multiple knockouts in polyploid crops (e.g., wheat) | Traditional CRISPR-Cas9 (Multiplexed) | Single guide RNA array can target all homologs simultaneously. | Design gRNAs with perfect homology to all target gene copies. |
| Fine-tuning gene expression (e.g., promoter editing) | Base Editor | Can modify specific transcription factor binding sites precisely. | Requires deep knowledge of cis-regulatory elements. Beware bystander edits. |
| Creating a stop codon (Nonsense mutation) | Base Editor (CBE primarily) | Can create CAA (Gln), CAG (Gln), CGA (Arg) → TAA, TAG, TGA stops. | More precise and predictable than generating frameshifts via NHEJ. |
Experimental Protocols
Protocol: Targeted Gene Knockout using CRISPR-Cas9 in Rice Protoplasts (Validation Assay)
Objective: To rapidly validate gRNA efficiency and generate knockout lines. Materials: See "The Scientist's Toolkit" below.
Methodology:
- gRNA Design & Cloning: Design 20-nt spacer sequences targeting the 5' exons of your gene of interest using tools like CHOPCHOP. Clone two complementary oligonucleotides into a U6-promoter driven gRNA expression vector via BsaI Golden Gate assembly.
- Vector Assembly: Assemble the final binary vector combining a plant codon-optimized Cas9 gene (driven by a ZmUbi promoter) and the validated gRNA expression cassette.
- Transformation & Delivery: Transform the vector into Agrobacterium tumefaciens strain EHA105. For rapid validation, perform PEG-mediated transfection of rice protoplasts (isolated from etiolated seedlings) with the purified plasmid. Incubate for 48h.
- Efficiency Analysis (PCR/RE Assay): a. Extract genomic DNA from protoplasts. b. Amplify a 300-500bp region flanking the target site. c. Perform a Restriction Enzyme (RE) digest if the edit destroys a natural site, or use T7 Endonuclease I (T7EI) assay: Denature and reanneal PCR products to form heteroduplexes if indels are present. T7EI cleaves mismatches. d. Analyze fragment patterns on gel electrophoresis. Editing efficiency is calculated as the intensity ratio of cleaved to uncleaved bands.
- Regeneration: For stable lines, perform Agrobacterium-mediated transformation of rice callus. Regenerate plants on selective media and genotype by sequencing.
Protocol: Installing a Herbicide-Resistance SNP using a Cytosine Base Editor (CBE) in Wheat
Objective: To convert a specific C•G base pair to T•A in the Acetolactate Synthase (ALS) gene, conferring resistance to imidazolinone herbicides.
Methodology:
- Target Site Selection: Identify the known resistance-conferring SNP (e.g., Pro-174-Ser). Ensure the target C is within the optimal "activity window" (positions 4-8, counting from the PAM distal end) of your CBE (e.g., rAPOBEC1-nCas9-UGI).
- Vector Construction: Clone the specific gRNA into the CBE system vector (containing the fusion protein: nCas9-deaminase-UGI). Use a wheat U6 promoter for gRNA and a strong constitutive promoter (e.g., TaUbi) for the base editor.
- Wheat Transformation: Isolate immature embryos from a model wheat cultivar. Co-cultivate with the Agrobacterium strain carrying the CBE vector.
- Selection & Screening: a. Primary Screening (Phenotypic): Grow T0 callus or plantlets on media containing a low dose of the imidazolinone herbicide. Resistant tissues indicate potential successful editing. b. Genotypic Validation: Perform Sanger sequencing of the target region from resistant tissues. Use chromatogram decomposition software (e.g., EditR, BEAT) to quantify the base conversion percentage. Look for a clean C→T peak at the target position. c. Off-Target Analysis: Use whole-genome sequencing or targeted sequencing of predicted off-target sites (based on sequence similarity) to check for unwanted edits.
- Segregation: Grow T1 plants. Sequence to identify lines where the edited allele is stably inherited without the transgene (segregated away).
Diagrams & Visual Workflows
Title: Decision Workflow: Base Editor vs. CRISPR-Cas9 Selection
Title: Molecular Mechanism Comparison: Base Editor vs. CRISPR-Cas9
The Scientist's Toolkit: Essential Reagents & Materials
| Item | Function & Description | Example Product/Catalog |
|---|---|---|
| Base Editor Plasmid Kit | All-in-one vector systems for plant expression of BE components (nCas9-deaminase-UGI). Critical for rapid deployment. | Addgene #138489 (pCBE-AtU6gRNA), #140375 (pABE8e). |
| Traditional Cas9 Vector | Binary vector with plant codon-optimized SpCas9, selection markers, and gRNA cloning scaffold. | pRGEB32 (Cas9-OsU6gRNA for rice), pBUN411 (for wheat/maize). |
| gRNA Cloning Kit | Modular system for assembling multiple gRNAs into a single transcriptional unit (tRNA or polycistronic). | Golden Gate MoClo Plant Toolkit (e.g., ToolBox I). |
| T7 Endonuclease I | Enzyme for detecting indels by cleaving heteroduplex DNA formed from wild-type and mutant PCR amplicons. | NEB, Cat# M0302S. |
| High-Efficiency Agrobacterium | Engineered strains for plant transformation. Essential for stable integration. | A. tumefaciens EHA105, AGL1, GV3101. |
| Plant DNA Isolation Kit | Reliable, PCR-ready genomic DNA extraction from tough plant tissues (leaf, callus). | DNeasy Plant Pro Kit (Qiagen), CTAB method reagents. |
| Sanger Sequencing Primers | Primers flanking the target site for sequencing validation of edits. | Custom-designed, HPLC purified. |
| Edit Analysis Software | Tools for quantifying base editing efficiency from Sanger chromatograms. | EditR (IDT), BEAT, DECODR. |
| Herbicide/Selective Agent | For phenotypic screening of edited lines (e.g., Imazethapyr for ALS edits). | Commercial grade, tissue culture tested. |
| Protoplast Isolation Enzymes | Cellulase and macerozyme mixtures for liberating protoplasts for rapid transient assays. | Cellulase R10, Macerozyme R10 (Yakult). |
Conclusion
Base editing and traditional CRISPR-Cas9 represent complementary, powerful arsenals for crop genome engineering, each with distinct advantages. While CRISPR-Cas9 remains unparalleled for gene knock-outs and large insertions via DSBs, base editing offers superior precision and simplicity for installing targeted point mutations without the complications of double-strand breaks. The choice between them hinges on the specific trait goal, the nature of the required genetic change, and the regulatory landscape. Future directions involve expanding the editable sequence space through novel base editor variants, improving delivery to elite crop varieties, and integrating these tools with other technologies like prime editing. For researchers, a deep understanding of both platforms is essential for designing robust breeding strategies that accelerate the development of climate-resilient, high-yielding, and nutritious crops to meet global food security challenges.