Rapid NBS-LRR Functional Screening: A Comprehensive Guide to Agrobacterium Transient Expression
This article provides a comprehensive guide for researchers and drug development professionals on utilizing Agrobacterium-mediated transient expression (AMTE) for the rapid functional screening of Nucleotide-Binding Site-Leucine-Rich Repeat (NBS-LRR) immune receptors.
Rapid NBS-LRR Functional Screening: A Comprehensive Guide to Agrobacterium Transient Expression
Abstract
This article provides a comprehensive guide for researchers and drug development professionals on utilizing Agrobacterium-mediated transient expression (AMTE) for the rapid functional screening of Nucleotide-Binding Site-Leucine-Rich Repeat (NBS-LRR) immune receptors. We first establish the foundational principles, comparing NBS-LRRs in plant and animal innate immunity and detailing the core Agrobacterium transformation mechanism. We then present a detailed, step-by-step methodological pipeline from vector design to phenotype scoring. To address common experimental hurdles, we dedicate a section to troubleshooting and optimizing key parameters like bacterial strain selection, silencing suppression, and protein expression levels. Finally, we discuss critical validation strategies, including comparisons to stable transformation and alternative transient systems, and highlight how data from this platform can inform biomedical research on mammalian NLR proteins and inflammasome biology. This guide synthesizes current best practices to enable robust, high-throughput screening of NBS-LRR function.
Understanding the Framework: NBS-LRR Immunity and Agrobacterium Biology
Nucleotide-binding site leucine-rich repeat (NBS-LRR) proteins constitute the largest family of intracellular immune receptors in plants. They detect pathogen effector proteins directly or indirectly, initiating robust defense signaling known as effector-triggered immunity (ETI). This response often includes a hypersensitive response (HR) characterized by programmed cell death at the infection site.
Table 1: Major Classes of Plant NBS-LRR Proteins
| Class | Domain Architecture (N- to C-terminus) | Representative Subfamilies | Key Features & Detection Mechanism |
|---|---|---|---|
| TNL | TIR - NBS - LRR | Arabidopsis RPS4, RPP1 | Contains a Toll/Interleukin-1 Receptor (TIR) domain. Often requires helper proteins (e.g., EDS1, NRG1). Common in dicots. |
| CNL | CC - NBS - LRR | Arabidopsis RPS2, RPS5 | Contains a coiled-coil (CC) domain. Often requires helper proteins (e.g., NDR1). Found in both dicots and monocots. |
| RNL | RPW8 CC - NBS - LRR | Arabidopsis ADR1, NRG1 | Helper NBS-LRRs (executors) that transduce signals from sensor TNLs/CNLs. Often function as "helper" or "executor" nodes. |
Table 2: Quantitative Genomic Distribution of NBS-LRR Genes
| Plant Species | Approx. Total NBS-LRR Count | TNL (%) | CNL/RNL (%) | Reference (Year) |
|---|---|---|---|---|
| Arabidopsis thaliana | ~150 | ~50% | ~50% (Majority CNL) | (Meyers et al., 2003) |
| Oryza sativa (Rice) | ~500 | <1% | ~99% (Majority CNL) | (Zhou et al., 2004) |
| Zea mays (Maize) | ~150 | 0% | 100% (CNL + RNL) | (Xiao et al., 2007) |
| Solanum lycopersicum (Tomato) | ~300 | ~30% | ~70% | (Andolfo et al., 2014) |
| Nicotiana benthamiana | ~400 | ~40% | ~60% | (Seong et al., 2020) |
The Role of Agrobacterium-Mediated Transient Expression in NBS-LRR Research
Agrobacterium tumefaciens-mediated transient expression in leaves (agroinfiltration) is a cornerstone technique for rapid in planta functional analysis of NBS-LRR proteins. Its utility within a thesis on functional screening includes:
- High-Throughput Screening: Co-expression of candidate NBS-LRR genes with known or putative effectors to identify novel immune pairs.
- Structure-Function Studies: Deploying domain swaps, point mutations, and truncations to dissect signaling domains.
- Pathway Elucidation: Co-infiltration with silencing suppressors or reporters to map downstream signaling components.
- Protein Localization and Interactions: Expressing fluorescently tagged versions for confocal microscopy or co-immunoprecipitation (Co-IP).
Core Protocols for NBS-LRR Functional Screening
Protocol 3.1: Agroinfiltration for Hypersensitive Response (HR) Assay
Objective: To test if a candidate NBS-LRR protein triggers cell death upon recognition of a co-expressed effector. Materials: See "Research Reagent Solutions" table. Method:
- Clone Generation: Clone your NBS-LRR gene (without stop codon if tagging) and putative effector gene into binary vectors (e.g., pEAQ-HT-DEST1 for effector, pBIN-GFP for NBS-LRR). Transform into Agrobacterium strain GV3101.
- Culture Preparation:
- Inoculate single colonies in 5 mL LB with appropriate antibiotics (Kanamycin, Rifampicin, Gentamicin). Grow overnight at 28°C, 200 rpm.
- Pellet cultures at 3000 x g for 10 min. Resuspend in infiltration buffer (10 mM MES, 10 mM MgCl₂, 150 µM acetosyringone, pH 5.6) to an OD₆₀₀ of 0.5.
- Incubate resuspended cultures at room temperature for 2-4 hours.
- Infiltration Mix:
- For co-expression, mix equal volumes of the NBS-LRR and effector Agrobacterium suspensions.
- Include controls: NBS-LRR alone, effector alone, empty vector.
- Infiltration: Using a 1 mL needleless syringe, press the tip against the abaxial side of a 4-5 week old N. benthamiana leaf and gently infiltrate the bacterial suspension. Mark infiltration zones.
- Phenotyping: Monitor leaves daily for 3-7 days for collapse and bleaching (HR). Document under white light. Quantify ion leakage (see Protocol 3.2) or conduct trypan blue staining for cell death validation.
Protocol 3.2: Quantitative Ion Leakage Assay for HR Quantification
Objective: To provide a quantitative measure of cell death by measuring electrolyte leakage from infiltrated leaf discs. Method:
- At 24-48 hours post-infiltration (hpi), harvest three 8-mm leaf discs from each infiltration zone.
- Rinse discs in 10 mL distilled water for 30 min to remove surface ions.
- Transfer discs to a tube with 10 mL fresh distilled water.
- Measure initial conductivity (C_initial) of the water using a conductivity meter.
- Incubate tubes at room temperature with gentle shaking for 3-6 hours.
- Measure conductivity again (C_sample).
- Autoclave the tubes to release all ions, cool, and measure final conductivity (C_total).
- Calculate Ion Leakage:
% Ion Leakage = [(C_sample - C_initial) / (C_total - C_initial)] * 100. Plot mean ± SD for triplicate samples.
Visualizing NBS-LRR Signaling and Experimental Workflows
Diagram 1: NBS-LRR Mediated Immune Signaling
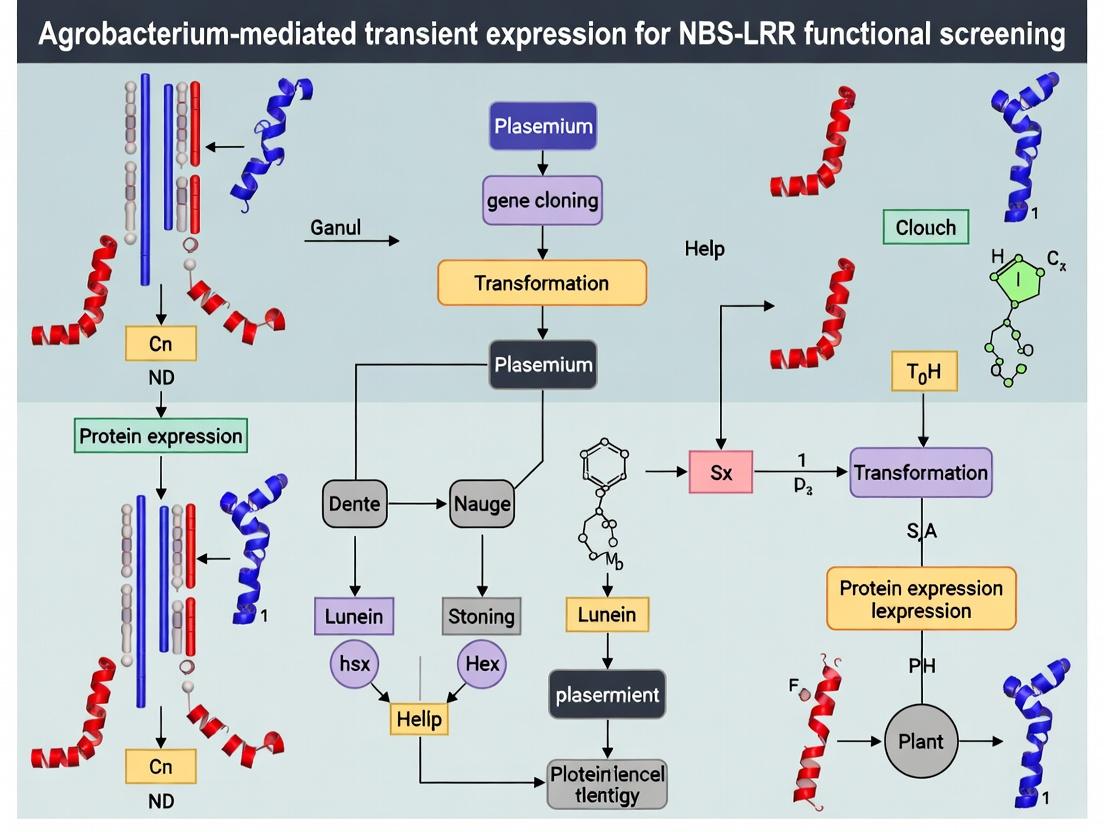
Diagram 2: Transient Expression Screening Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Agroinfiltration-Based NBS-LRR Screening
| Item | Function/Description | Example Product/Catalog |
|---|---|---|
| Binary Vector (High Expression) | For high-level transient expression of NBS-LRR or effector gene. Often includes 5' and 3' UTRs for stability. | pEAQ-HT series, pGWB vectors |
| Agrobacterium Strain | Disarmed strain optimized for plant transformation and transient expression. | GV3101 (pMP90), AGL-1, EHA105 |
| Acetosyringone | Phenolic compound that induces Agrobacterium vir genes, essential for T-DNA transfer. | Sigma-Aldrich, D134406 |
| Infiltration Buffer | Provides correct pH and cations for bacterial viability and vir gene induction during infiltration. | 10 mM MES, 10 mM MgCl₂, pH 5.6 |
| Plant Material | Model plant with well-characterized genetics and high susceptibility to agroinfiltration. | Nicotiana benthamiana (4-5 weeks old) |
| Cell Death Stain | Histological stain that visualizes dead (blue) cells. Validates HR phenotype. | Trypan Blue Solution (0.4%) |
| Conductivity Meter | Device to measure ion leakage from leaf discs as a quantitative HR readout. | Horiba B-173, Orion Star A212 |
| Confocal Microscopy | For subcellular localization studies of fluorescently tagged NBS-LRR proteins. | Zeiss LSM 900, Leica SP8 |
| Co-Immunoprecipitation Kits | For validating protein-protein interactions between NBS-LRRs, effectors, or signaling components. | μMACS Epitope Tag Protein Isolation Kits (Miltenyi) |
| Gateway Cloning System | Enables rapid, high-throughput transfer of NBS-LRR ORFs into multiple destination vectors. | Thermo Fisher, 12535-027 |
Plant Nucleotide-Binding Site Leucine-Rich Repeat proteins (NBS-LRRs) and mammalian NOD-like receptors (NLRs) are central cytosolic sentinels in innate immunity. They share a tripartite domain architecture, enabling pathogen perception, nucleotide-dependent activation, and downstream signaling. This structural conservation allows for cross-kingdom functional insights and screening methodologies.
Table 1: Core Structural and Functional Parallels between NBS-LRRs and NLRs
| Feature | Plant NBS-LRRs | Mammalian NLRs (Inflammasome-forming) |
|---|---|---|
| Domains | N-terminal TIR/CC, NB-ARC, LRR | N-terminal PYD/CARD, NACHT, LRR |
| Sensor Module | LRR domain (direct/indirect ligand binding) | LRR domain (often for PAMP/DAMP sensing) |
| Oligomerization & Switch | NB-ARC (ADP/ATP binding; "on/off" switch) | NACHT (ATP hydrolysis; oligomerization platform) |
| Signal Adapter/Effector | TIR/CC domains (self-association) | PYD/CARD (recruit ASC, caspase-1) |
| Activation Outcome | HR, SA/JA/ET pathways, Transcriptional Reprogramming | Inflammasome assembly, Caspase-1 activation, IL-1β/IL-18 maturation, Pyroptosis |
| Regulatory Mechanisms | SGT1, HSP90, RIN4, miRNAs | SGT1, HSP90, CARD-only proteins, miRNAs |
Application Notes for Functional Screening via Agrobacterium Transient Expression
Within the thesis context of Agrobacterium-mediated transient expression for NBS-LRR functional screening, these parallels are exploited to:
- Screen for Autoactive Mutants: Expression of wild-type and mutant NBS-LRRs in Nicotiana benthamiana to identify constitutive activators (phenocopying NLR gain-of-function mutations causing autoinflammatory disease).
- Reconstitute Signaling Pathways: Co-expression of sensor, adapter, and effector components from both plant and mammalian systems to study compatibility and signaling logic.
- Identify and Validate Inhibitors: Use the rapid cell death readout from autoactive NBS-LRR/NLRs to screen for pharmacological or protein-based suppressors.
Table 2: Quantitative Parameters for Transient Expression Assays
| Parameter | Typical Range/Value | Notes for NBS-LRR/NLR Studies |
|---|---|---|
| Agrobacterium Strain | GV3101, LBA4404, AGL1 | GV3101 (pMP90) is most common for N. benthamiana. |
| OD600 for Infiltration | 0.2 - 0.8 (final, resuspended) | High OD (>0.5) often needed for strong NBS-LRR expression. |
| Incubation Time Post-Infiltration | 24 - 72 hours | Cell death from autoactive proteins typically appears 36-48 hpi. |
| Optimal Leaf Age | 3-4 weeks old | Fully expanded, robust leaves. |
| Co-infiltration Mix Ratios (Sensor:Effector) | 1:1 to 1:5 (OD-based) | Requires optimization for each pair (e.g., NLRP3:ASC). |
| Positive Control Cell Death Onset | 24-36 hpi | e.g., AvrPto/Pto, Bax, ZAR1 resistosome components. |
Detailed Experimental Protocols
Protocol 3.1: Agrobacterium-Mediated Transient Expression inN. benthamianafor NBS-LRR/NLR Activation Assay
Objective: To express and assess the activity of wild-type, mutant, or chimeric NBS-LRR/NLR proteins.
Materials: See "The Scientist's Toolkit" below. Procedure:
- Clone gene of interest into a binary vector with a strong plant promoter (e.g., 35S) and an epitope tag (HA, FLAG, GFP).
- Transform the plasmid into Agrobacterium tumefaciens strain GV3101 via electroporation or freeze-thaw.
- Select single colonies on LB plates with appropriate antibiotics (e.g., rifampicin, gentamicin, kanamycin). Incubate at 28°C for 2 days.
- Start a 5 mL liquid culture in LB + antibiotics. Shake at 28°C, 200 rpm, for 24-48 hrs.
- Pellet cells at 4000 x g for 10 min. Resuspend in Infiltration Buffer (10 mM MES, 10 mM MgCl2, 150 µM acetosyringone, pH 5.6).
- Adjust the OD600 to the desired final density (e.g., 0.5 for the gene of interest, 0.3 for a suppressor screen). Let the suspension sit at room temp for 1-4 hrs.
- Infiltrate the suspension into the abaxial side of 3-4 week old N. benthamiana leaves using a needleless syringe.
- Monitor plants daily for phenotypes (e.g., hypersensitive response cell death, ion leakage). Document photographically.
- Harvest leaf discs at specified time points (e.g., 36 hpi) for protein extraction (using standard Laemmli buffer or RIPA buffer) or ion leakage measurement.
Protocol 3.2: Ion Leakage Assay for Quantitative Cell Death Measurement
Objective: To quantify membrane integrity loss, a hallmark of NBS-LRR/NLR-induced hypersensitive response/pyroptosis.
Procedure:
- At the monitoring time point, harvest uniform leaf discs (e.g., 8 mm diameter) from infiltrated zones using a cork borer.
- Place 4-6 discs in a tube with 10 mL of distilled, deionized water. Rinse briefly (20 sec) to remove initial electrolytes from wounding.
- Transfer discs to a new tube containing 10 mL of fresh distilled water.
- Place tubes on a shaker at low speed for the duration of the experiment.
- Measure conductivity (Cinitial) of the bathing solution immediately using a conductivity meter.
- Incubate samples at room temperature for 4-8 hours.
- Measure conductivity again (Ctime).
- Autoclave or boil the tubes to kill all tissue and release total electrolytes. Cool to room temperature and measure final conductivity (Ctotal).
- Calculate ion leakage: [(Ctime - Cinitial) / (Ctotal - Cinitial)] * 100%.
- Plot % ion leakage vs. time for different constructs.
Visualization: Signaling Pathways and Workflows
Title: Core Immune Activation Pathways in Plants and Mammals
Title: Transient Expression Screening Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in NBS-LRR/NLR Screening | Example/Notes |
|---|---|---|
| Binary Vector (High Expression) | Drives transgene expression in plant cells. | pEAQ-HT, pBIN61, pB7WG2. 35S or UBQ promoter preferred. |
| Agrobacterium Strain | Delivers T-DNA containing gene of interest into plant cells. | GV3101 (pMP90) is standard; offers good virulence, lacks oncogenes. |
| Acetosyringone | Phenolic inducer of Agrobacterium virulence genes. Critical for efficient transformation. | Use at 150-200 µM in infiltration buffer. Add fresh before use. |
| Silencing Suppressor | Co-expressed to boost transgene expression levels by inhibiting RNAi. | p19 protein from Tomato bushy stunt virus. Often co-infiltrated. |
| Epitope Tags | Enables protein detection, localization, and co-IP validation. | C-terminal HA, FLAG, GFP, or RFP tags. Avoid N-terminal tags for some NLRs. |
| Conductivity Meter | Essential for quantifying ion leakage as a measure of cell death. | Requires high sensitivity for low-conductivity solutions (µS/cm range). |
| Anti-Tag Antibodies | For immunoblot or co-IP to confirm protein expression and complexes. | Anti-HA, anti-FLAG (mouse monoclonal), anti-GFP (rabbit polyclonal). |
| Positive Control Constructs | Validates infiltration efficiency and cell death readout. | Pro-apoptotic murine Bax, Autoactive N (TIR-NBS-LRR), NLRP3 mutants. |
Agrobacterium tumefaciens, a soil-borne pathogen, is a cornerstone of plant biotechnology due to its natural ability to transfer DNA (T-DNA) into plant genomes. Within the broader thesis on "Agrobacterium-mediated transient expression for NBS-LRR functional screening," this bacterium serves as the primary delivery vector. Its utility lies in enabling rapid, high-throughput functional characterization of Nucleotide-Binding Site Leucine-Rich Repeat (NBS-LRR) immune receptors in planta without genomic integration. This protocol set details the optimized use of the Agrobacterium toolbox for such screenings.
Key Signaling Pathways in Agrobacterium-Mediated Transformation
Diagram 1: Agrobacterium vir Gene Induction and T-DNA Transfer
Application Notes & Protocols
Application Note 1: High-Throughput NBS-LRR Screening Workflow This workflow utilizes Agrobacterium-mediated transient expression (agroinfiltration) in Nicotiana benthamiana to rapidly assess NBS-LRR-mediated cell death responses upon co-expression with candidate effector proteins.
Diagram 2: NBS-LRR Functional Screening Workflow
Protocol 1: Agroinfiltration of Nicotiana benthamiana for Transient Co-expression
- Objective: Deliver NBS-LRR and candidate effector genes into plant leaf cells for functional interaction assays.
- Materials: See "Research Reagent Solutions" (Section 5).
- Method:
- Strain Preparation: Inoculate single colonies of Agrobacterium carrying your genes of interest in 5 mL LB with appropriate antibiotics. Grow overnight at 28°C, 220 rpm.
- Induction: Sub-culture to an OD600 of 0.1 in fresh LB (+antibiotics, 10 mM MES pH 5.6, 20 μM acetosyringone). Grow to OD600 0.5-1.0 (approx. 6-8 hrs).
- Harvest & Resuspension: Pellet cells at 4000 x g for 10 min. Resuspend in infiltration buffer (10 mM MgCl₂, 10 mM MES pH 5.6, 150 μM acetosyringone). Adjust final OD600 to 0.5 for each construct. For co-infiltration, mix equal volumes of NBS-LRR and effector strains.
- Infiltration: Incubate resuspensions at room temp for 1-3 hrs. Using a needleless syringe, press the tip against the abaxial side of a 4-5 week old N. benthamiana leaf and gently infiltrate the bacterial suspension.
- Post-Infiltration: Maintain plants under normal growth conditions (22-24°C, 16/8 hr light/dark).
- Phenotyping: Visually monitor infiltrated zones for HR cell death (collapse, bleaching) starting at 24 hours post-infiltration (hpi). Document with photography.
Protocol 2: Ion Leakage Assay for Quantitative HR Cell Death Measurement
- Objective: Quantify NBS-LRR/effector-induced cell death by measuring electrolyte leakage from leaf discs.
- Method:
- At 24-48 hpi, harvest leaf discs (e.g., 8 mm diameter) from infiltrated zones using a cork borer. Include control discs (empty vector, effector alone).
- Rinse discs briefly in distilled water to remove surface ions.
- Place 4-6 discs in a 50 mL tube containing 25 mL of distilled water. Shake gently (50 rpm) at room temperature.
- Measure conductivity of the bathing solution at time 0 (C0) and at regular intervals (e.g., 1, 2, 4, 8, 24 h) using a conductivity meter.
- After final measurement, autoclave the tubes to release all electrolytes, cool, and measure total conductivity (Ctotal).
- Calculate ion leakage as a percentage: % Ion Leakage = (Ct / Ctotal) * 100. Plot % leakage over time.
Table 1: Optimization Parameters for Transient NBS-LRR Screening
| Parameter | Optimal Range for N. benthamiana | Effect on Transient Expression | Recommended for NBS-LRR Screening |
|---|---|---|---|
| Agrobacterium Strain | GV3101 (pMP90), AGL-1 | High virulence, suppressed silencing | GV3101 (pSoup) - Superior for co-expression |
| Infiltration OD600 | 0.2 - 1.0 | Higher OD increases protein but can cause non-specific HR | 0.4 - 0.6 (per strain in mix) - Balances expression & specificity |
| Acetosyringone (Induction) | 100 - 200 μM | Essential for vir gene induction | 150 - 200 μM in final resuspension |
| Plant Age | 4 - 5 weeks post-sowing | Younger plants more susceptible | 4 - 5 weeks - Fully expanded leaves, robust response |
| Time to HR Phenotype | 20 - 96 hours post-infiltration | Varies by NBS-LRR/Effector pair | Score at 24, 48, 72 hpi - Kinetic assessment is critical |
| Co-infiltration Ratio (NBS-LRR:Effector) | 1:1 to 1:5 (v/v) | Ratios can modulate interaction sensitivity | Start with 1:1; titrate effector if background exists |
Table 2: Advantages of Transient vs. Stable Expression for NBS-LRR Screening
| Feature | Agrobacterium-Mediated Transient Expression (N. benthamiana) | Stable Transformation (e.g., Arabidopsis) |
|---|---|---|
| Time Scale | 3-7 days from infiltration to result | 3-6 months to generate T1 plants |
| Throughput | Very High - Dozens of constructs tested per plant | Low - Labor-intensive per line |
| Gene Silencing | Minimal before assay completion | Can occur in later generations |
| Lethality Tolerance | High - Can assay lethal immune responses | Low - Lethality prevents plant regeneration |
| Cost per Assay | Low | High |
| Best For | Rapid functional screening, mutagenesis studies, effector identification | Detailed phenotypic analysis, genetic crosses, inheritance studies |
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in NBS-LRR Screening | Example/Notes |
|---|---|---|
| Binary Vector (e.g., pEAQ-HT, pBIN61) | Carries gene of interest between T-DNA borders for transfer. | pEAQ-HT offers very high, silencing-suppressed expression. |
| Agrobacterium Strain GV3101 (pMP90RK) | Disarmed, helper plasmid-containing strain for efficient transformation. | Contains pSoup plasmid for vir gene complementation. Superior for co-delivery. |
| Acetosyringone | Phenolic compound that activates the VirA/VirG two-component system, inducing vir genes. | Critical for efficient T-DNA transfer in non-wounded plants. |
| Infiltration Buffer (MgCl₂/MES) | Provides optimal ionic and pH conditions for bacterial viability and plant cell interaction. | 10 mM MgCl₂, 10 mM MES pH 5.6, with fresh acetosyringone. |
| Nicotiana benthamiana Plants | Model plant host with high susceptibility to agroinfiltration and low endogenous NBS-LRR interference. | 4-5 weeks old, grown under controlled conditions for consistency. |
| Silencing Suppressor (e.g., p19) | Co-expressed to inhibit post-transcriptional gene silencing, boosting protein yield. | Optional but recommended for weak interactions; use a third Agrobacterium strain in the mix. |
| Conductivity Meter | Essential tool for quantifying the Hypersensitive Response (HR) via the ion leakage assay. | Provides quantitative, reproducible data on cell death progression. |
| Syringe (1 mL, needleless) | Tool for physically introducing the Agrobacterium suspension into the leaf apoplast. | Apply gentle, even pressure to avoid damaging the leaf. |
Core Mechanism of T-DNA Transfer and Transient Expression
1. Introduction: Context within NBS-LRR Functional Screening Within a thesis exploring Agrobacterium-mediated transient expression for functional screening of Nucleotide-Binding Site Leucine-Rich Repeat (NBS-LRR) immune receptors, understanding the core T-DNA transfer mechanism is fundamental. Transient expression via Agrobacterium tumefaciens enables rapid, high-throughput in planta assessment of NBS-LRR gene function, auto-activation, and signaling interactions. The efficiency and timing of this screening platform are directly governed by the molecular machinery of T-DNA transfer.
2. Core Mechanism: From Bacterial Cell to Plant Nucleus The transfer of T-DNA (Transferred-DNA) from Agrobacterium to the plant cell is a Vir (Virulence) protein-mediated process. The following table summarizes the key quantitative parameters of this process under optimal laboratory conditions.
Table 1: Key Quantitative Parameters of T-DNA Transfer for Transient Expression
| Parameter | Typical Value / Range | Notes / Conditions |
|---|---|---|
| Time to Initial Protein Detection | 24-48 hours post-infiltration (hpi) | For NBS-LRR-GFP fusions, earliest detection via confocal microscopy. |
| Peak Transient Expression Window | 48-96 hpi | Optimal time for harvesting tissue for assays (e.g., cell death scoring, protein extraction). |
| Optimal Agrobacterium OD₆₀₀ for Infiltration | 0.2 - 0.8 | Depends on plant species, target tissue, and vector. Higher ODs can trigger phytotoxicity. |
| Acetosyringone Induction Concentration | 100 - 200 µM | Essential for inducing the vir gene region. |
| Co-cultivation Period Post-Inoculation | 24 - 72 hours | Critical for T-DNA transfer; requires high humidity and appropriate temperature (22-25°C). |
| Estimated T-DNA Copy Number per Cell | 1 - 10+ (transient) | Variable; T-DNA remains unintegrated for transient expression. |
3. Experimental Protocols
Protocol 3.1: Agrobacterium Preparation for Leaf Infiltration (Syringe Method) Objective: To prepare Agrobacterium tumefaciens (strain GV3101 or LBA4404) harboring a binary vector with an NBS-LRR gene construct for transient expression in Nicotiana benthamiana leaves.
- Culture Initiation: Inoculate a single colony of the transgenic Agrobacterium into 2-5 mL of LB medium with appropriate antibiotics (e.g., rifampicin, gentamicin, kanamycin). Grow overnight at 28°C with shaking (220 rpm).
- Secondary Culture: Dilute the primary culture 1:50 into fresh LB with antibiotics and 10 mM MES (pH 5.6). Add acetosyringone to a final concentration of 200 µM. Grow overnight to late-log phase (OD₆₀₀ ≈ 0.8-1.2).
- Harvesting: Pellet bacteria by centrifugation at 3000-5000 x g for 15 minutes at room temperature.
- Resuspension/Induction: Resuspend the pellet in MMA infiltration buffer (10 mM MgCl₂, 10 mM MES pH 5.6, 200 µM acetosyringone). Adjust the final OD₆₀₀ to 0.2-0.5 for most assays. For strong cell death assays with NBS-LRRs, 0.4 is a typical starting point.
- Induction Incubation: Let the resuspended culture sit at room temperature, protected from light, for 1-3 hours before infiltration.
- Infiltration: Using a 1 mL needleless syringe, press the tip against the abaxial side of a 4-5 week-old N. benthamiana leaf and gently infiltrate the bacterial suspension. Mark the infiltrated area.
- Post-Infiltration: Maintain plants under standard growth conditions (22-25°C, 16h light/8h dark) until analysis.
Protocol 3.2: Monitoring NBS-LRR-Induced Cell Death Response Objective: To quantitatively assess the hypersensitive response (HR) triggered by a transiently expressed autoactive NBS-LRR or an effector-recognizing NBS-LRR.
- Time-Course Documentation: Photograph infiltrated leaf patches at 24, 48, 72, and 96 hpi under consistent lighting.
- Trypan Blue Staining (for Cell Death Visualization): a. Staining Solution: Prepare 1:1:1:1 mixture of phenol, glycerol, lactic acid, and water. Add 1 mg/mL Trypan Blue. Filter before use. b. Procedure: Submerge leaf discs in boiling staining solution for 1 minute. Incubate at room temperature overnight with gentle shaking. c. Destaining: Transfer tissue to a 2.5 g/mL chloral hydrate solution for 24-48 hours, changing solution until background is clear. d. Imaging: Mount discs in 50% glycerol and image under a bright-field microscope. Dead cells stain dark blue.
- Ion Conductance Measurement (Quantitative HR): a. Use a conductivity meter. b. Excise a uniform leaf disc (e.g., 8 mm diameter) from the infiltrated zone. c. Place the disc in a tube with 5 mL of distilled, deionized water. d. Shake gently for 2-3 hours at room temperature. e. Measure the conductivity of the solution (C1). f. Boil the sample for 15 minutes, cool to room temperature, and measure total conductivity (C2). g. Calculate relative ion leakage: (C1 / C2) * 100%.
4. Diagrams: Signaling Pathways and Workflows
Diagram 1: Agrobacterium T-DNA Transfer Signaling Pathway (100 chars)
Diagram 2: Transient NBS-LRR Expression Screening Workflow (99 chars)
5. The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Agrobacterium-Mediated Transient NBS-LRR Assays
| Reagent / Material | Function & Rationale |
|---|---|
| Binary Vector (e.g., pCambia, pEAQ) | Carries NBS-LRR gene between T-DNA borders and plant selection marker. Gateway-compatible versions enable high-throughput cloning. |
| A. tumefaciens Strain (GV3101) | Disarmed, widely used strain with high transformation efficiency in N. benthamiana. Contains a rifampicin resistance marker. |
| Acetosyringone | Phenolic compound that activates the bacterial VirA/VirG system, inducing vir gene expression essential for T-DNA transfer. |
| MMA Buffer (MgCl₂, MES, AS) | Optimized low-salt, acidic buffer for bacterial resuspension that promotes virulence and supports plant tissue health during infiltration. |
| Nicotiana benthamiana Plants | Model plant for transient assays due to susceptibility to Agrobacterium, low silencing background, and well-characterized immune system. |
| Trypan Blue Stain | Vital dye excluded by live cells; selectively stains dead cells, allowing visualization of the NBS-LRR-triggered hypersensitive response (HR). |
| Conductivity Meter | Provides quantitative, reproducible measurement of electrolyte leakage from damaged plant cells, a key metric for HR cell death intensity. |
| Needleless Syringes (1 mL) | Standard tool for manually infiltrating bacterial suspension into the leaf intercellular space (apoplast). |
Why Transient Expression? Advantages for High-Throughput NBS-LRR Screening
Application Notes
Within the context of Agrobacterium-mediated transient expression for NBS-LRR functional screening research, transient expression offers a rapid, scalable, and flexible alternative to stable transformation. This is critical for dissecting the functions of large NBS-LRR gene families in plant immunity, particularly for high-throughput screening of effector recognition and cell death responses.
Core Advantages:
- Speed: Functional data can be obtained within 2-4 days post-infiltration, bypassing the months required for stable plant transformation and regeneration.
- Scalability: Enables parallel testing of dozens of NBS-LRR / effector combinations in a single experiment.
- Flexibility: Allows co-expression of multiple constructs (e.g., NBS-LRR, effector, fluorescent markers) in varying ratios.
- Overcome Lethality: Facilitates the study of NBS-LRR genes whose constitutive expression would be lethal in stable lines.
- In Planta Context: The assay occurs within the living plant cell, preserving essential components for proper folding, subcellular localization, and signaling.
Table 1: Comparison of Expression Methods for NBS-LRR Screening
| Parameter | Stable Transformation | Agrobacterium-Mediated Transient Expression |
|---|---|---|
| Time to Result | 3-9 months | 2-4 days |
| Throughput | Low (construct/line) | High (multiple constructs/leaf) |
| Expression Level | Consistent, often low | High, variable |
| Multiplexing Potential | Difficult | Straightforward (co-infiltration) |
| Suitability for HR-based Screen | Low (lethality issues) | High |
| Cost per Construct Tested | High | Low |
Table 2: Typical Metrics for Transient NBS-LRR/Effector Screening
| Metric | Typical Range/Observation | Key Influencing Factor |
|---|---|---|
| Peak Protein Expression | 24-72 hours post-infiltration (hpi) | Plant species, Agrobacterium strain, temperature |
| Hypersensitive Response (HR) Onset | 20-48 hpi | NBS-LRR/Effector pair specificity, strength |
| Assay Throughput (leaves/week) | 50-200 | Lab scale optimization |
| Coefficient of Variation (HR assays) | 15-25% | Infiltration uniformity, plant health |
Experimental Protocols
Protocol 1: High-ThroughputAgrobacteriumTransient Expression inN. benthamianafor NBS-LRR Screening
Research Reagent Solutions & Essential Materials:
| Item | Function/Explanation |
|---|---|
| N. benthamiana Plants (4-5 week old) | Model plant with low background of NBS-LRRs, highly susceptible to Agrobacterium infiltration. |
| GV3101 pSoup Agrobacterium Strain | Disarmed strain with modified Ti plasmid for high-level transient T-DNA expression. |
| Binary Vector (e.g., pEAQ-HT, pCambia) | Carries gene of interest (NBS-LRR or effector) between T-DNA borders for transfer. |
| Induction Buffer (10 mM MES, 10 mM MgCl₂, 150 µM Acetosyringone, pH 5.6) | Buffer for final Agrobacterium resuspension; acetosyringone induces Vir genes. |
| Syringe (1 mL needleless) | For manual leaf infiltration. |
| Optical Density (OD600) Spectrophotometer | For standardizing bacterial culture density. |
| Silwet L-77 (or similar surfactant) | For vacuum infiltration of whole seedlings (high-throughput method). |
Methodology:
- Construct Preparation: Clone NBS-LRR candidates and known/predicted effector genes into binary expression vectors (e.g., C-terminal tags for detection).
- Agrobacterium Transformation: Introduce constructs into Agrobacterium strain GV3101 via electroporation or freeze-thaw.
- Culture Initiation: Inoculate single colonies into 2-5 mL LB with appropriate antibiotics (e.g., Kanamycin, Rifampicin, Gentamicin). Incubate at 28°C, 200 rpm for 24-48h.
- Culture Scaling: Subculture 1:50 into fresh LB with antibiotics and 10 mM MES (pH 5.6), 20 µM acetosyringone. Grow to OD600 ~0.8-1.2 (28°C, ~24h).
- Harvest & Induction: Pellet bacteria (3000 x g, 10 min). Resuspend in Induction Buffer to a final OD600 of 0.4-0.6 for single constructs. For co-expression (NBS-LRR + effector), mix suspensions to final OD600 of 0.2-0.3 each.
- Incubation: Let suspensions sit at room temperature, in the dark, for 1-3 hours.
- Leaf Infiltration: Using a needleless syringe, press tip against abaxial side of N. benthamiana leaf and gently infiltrate. Mark infiltration zones. Include controls (empty vector, effector alone, NBS-LRR alone).
- Plant Maintenance: Keep plants at 21-25°C, with high humidity for first 24h, then normal conditions.
- Phenotyping & Analysis (24-72 hpi):
- HR Scoring: Visually document cell death (collapsed tissue, bleaching). Use standardized scoring scales (e.g., 0=no HR, 5=confluent tissue collapse).
- Ion Leakage Assay: Quantify HR by measuring electrolyte leakage from leaf discs.
- Protein Extraction & Immunoblot: Verify expression levels of NBS-LRR and effector proteins.
- Sample Collection: Freeze tissue in liquid N₂ for subsequent biochemical assays.
Protocol 2: Quantitative Ion Leakage Assay for Hypersensitive Response
Methodology:
- Sample Collection: At desired time point (e.g., 24 hpi), take leaf discs from infiltrated zones using a cork borer (e.g., 6 mm diameter).
- Washing: Rinse discs in 10 mL of distilled water for 30 min to remove initial ions from wounding.
- Incubation: Transfer 4-6 discs to a tube containing 10 mL of fresh, sterile distilled water.
- Measurement: Use a conductivity meter to measure initial conductivity (C_initial). Shake tubes gently at room temperature for 3-6 hours.
- Final Measurement: Measure conductivity again (Cfinal). Then autoclave or boil the tube to release all ions, cool, and measure total conductivity (Ctotal).
- Calculation: Calculate ion leakage as: % Ion Leakage = [(Cfinal - Cinitial) / C_total] x 100. Plot values over time or as a final endpoint.
Visualizations
Title: Transient NBS-LRR Screening Workflow
Title: NBS-LRR Activation & Signaling Output
Within the broader thesis on Agrobacterium-mediated transient expression for NBS-LRR (Nucleotide-Binding Site Leucine-Rich Repeat) functional screening, these application notes detail three critical, interconnected assays. The transient expression system (e.g., in Nicotiana benthamiana) allows for rapid, high-throughput functional characterization of plant immune receptors and pathogen effector proteins. These assays—Effector Recognition, Autoactivity, and Cell Death—are fundamental for identifying and validating immune receptor-effector pairs, elucidating signaling pathways, and characterizing gain-of-function mutations.
Application Notes & Protocols
Effector Recognition Assay
Purpose: To determine if a co-expressed candidate NBS-LRR receptor can specifically recognize a pathogen effector protein, triggering a measurable Hypersensitive Response (HR).
Detailed Protocol:
- Clone Construction: Clone the candidate NBS-LRR gene into a binary expression vector (e.g., pEAQ-HT or pBINplus) under a strong constitutive promoter (e.g., 35S). Clone the candidate effector gene into a compatible vector (e.g., pGREENII 0029 with a different antibiotic resistance).
- Agrobacterium Transformation: Transform separate Agrobacterium tumefaciens strains (e.g., GV3101) with each construct. Select on appropriate antibiotics (rifampicin, gentamicin, plus vector-specific antibiotics).
- Culture Preparation: Grow individual Agrobacterium cultures overnight at 28°C in LB with antibiotics. Pellet cultures and resuspend in infiltration buffer (10 mM MES, 10 mM MgCl₂, 150 µM acetosyringone, pH 5.6) to a final OD₆₀₀ of 0.5 for the receptor and 0.3 for the effector.
- Mixed Infiltration: Combine the resuspended bacterial cultures in a 1:1 ratio. Using a needleless syringe, infiltrate the mixture into the abaxial side of 4-6 week-old N. benthamiana leaves. Include controls: effector + empty vector (EV), receptor + EV, and EV + EV.
- Incubation & Monitoring: Place plants in controlled conditions (22-25°C, 16h light). Monitor infiltrated areas for cell death symptoms (collapsed, water-soaked, or desiccated tissue) at 24, 48, 72, and 96 hours post-infiltration (hpi).
- Scoring & Documentation: Visually score HR strength (0-5 scale) and photograph under consistent lighting. For quantification, conduct ion leakage assays or stain with trypan blue.
Table 1: Typical HR Scoring Scale for Effector Recognition
| Score | Phenotype Description | Interpretation |
|---|---|---|
| 0 | No visible symptoms | No recognition |
| 1 | Mild chlorosis/bleaching | Very weak recognition |
| 2 | Focal, confluent chlorosis | Weak recognition |
| 3 | Patchy tissue collapse | Clear HR/recognition |
| 4 | Strong, confluent collapse within zone | Strong recognition |
| 5 | Rapid, full collapse spreading beyond zone | Very strong recognition |
Autoactivity Assay
Purpose: To identify gain-of-function mutations or allelic variants in NBS-LRR receptors that trigger constitutive, effector-independent immune signaling and cell death.
Detailed Protocol:
- Mutant/Variant Library: Generate a site-directed mutant or allelic variant library of the target NBS-LRR gene. Clone into a binary expression vector as in 2.1.
- Agrobacterium Preparation: Transform and prepare Agrobacterium cultures as per steps 2-3 in 2.1, adjusting the final OD₆₀₀ to 0.4.
- Single-Strain Infiltration: Infiltrate each NBS-LRR variant individually into separate leaf panels of N. benthamiana.
- Phenotypic Screening: Monitor infiltrated areas daily for 5-7 days. Autoactivity is indicated by spontaneous cell death in the absence of co-infiltrated effector.
- Confirmation & Titration: For positive hits, repeat assay with a dilution series (OD₆₀₀ 0.05, 0.1, 0.2, 0.4) to assess the strength of autoactivity. Co-infiltration with generic silencing suppressors (e.g., P19) may be used to enhance protein expression and confirm phenotype.
- Quantification: Use electrolyte leakage assays over a 48-hour time course to quantify cell death kinetics.
Table 2: Autoactivity Assay Quantification via Ion Leakage
| Time (hpi) | Wild-Type NBS-LRR (μS/cm) | Autoactive Mutant M1 (μS/cm) | Autoactive Mutant M2 (μS/cm) |
|---|---|---|---|
| 0 | 10 ± 2 | 12 ± 3 | 11 ± 2 |
| 24 | 15 ± 4 | 85 ± 12 | 210 ± 25 |
| 48 | 25 ± 5 | 320 ± 45 | 450 ± 60 |
Cell Death Assay
Purpose: A broader assay to characterize the cell death phenotype triggered either by effector recognition, autoactive receptors, or directly cytotoxic effectors.
Detailed Protocol:
- Experimental Setup: Depending on the goal, prepare Agrobacterium cultures for single or co-infiltration as described in 2.1 and 2.2.
- Infiltration: Perform infiltrations in a minimum of three biological replicates (different plants).
- Phenotypic Documentation:
- Visual Scoring: As per Table 1.
- Trypan Blue Staining: Harvest leaf discs at defined time points. Boil in trypan blue staining solution (10 mL lactic acid, 10 mL glycerol, 10 mL phenol, 10 mg trypan blue, dissolved in 10 mL water and made up to 40 mL with ethanol) for 2 minutes. Incubate overnight at room temperature. Destain in chloral hydrate solution (2.5 g/mL). Image; dead cells stain dark blue.
- Electrolyte Leakage: Place six 4-mm leaf discs in 5 mL of distilled water. Measure conductivity of the bathing solution (C1). Shake for 3 hours and measure again (C2). Boil samples for 15 minutes, cool, and measure final conductivity (C3). Calculate ion leakage as [(C2 - C1) / C3] * 100%.
- Statistical Analysis: Perform ANOVA or t-tests on quantified data (e.g., ion leakage percentages) from at least three independent experiments.
Diagrams
Diagram 1: Immune receptor activation leading to HR.
Diagram 2: Transient expression screening workflow.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Transient NBS-LRR Screening
| Item | Function & Application | Example/Details |
|---|---|---|
| Binary Vectors | High-level protein expression in plants. | pEAQ-HT (high yield), pBINplus (standard), pGREENII (effector co-expression). |
| Agrobacterium Strain | Delivery vehicle for T-DNA into plant cells. | GV3101 (pMP90), AGL-1. Rifampicin and gentamicin resistant. |
| Infiltration Buffer | Suspension medium inducing virulence. | 10 mM MES, 10 mM MgCl₂, 150 µM acetosyringone, pH 5.6. |
| N. benthamiana Plants | Model plant for transient assays. | Grown at 22-25°C, 16h light, for 4-6 weeks. |
| Silencing Suppressor | Enhances transgene expression. | p19 protein of Tomato bushy stunt virus, co-infiltrated or transgenic line. |
| Trypan Blue Solution | Histochemical stain for dead cells. | 0.05% Trypan blue in lactophenol-ethanol. |
| Conductivity Meter | Quantifies ion leakage from dying cells. | Essential for generating quantitative cell death data. |
| Acetosyringone | Phenolic compound inducing Agrobacterium vir genes. | Critical for high transformation efficiency. Stock in DMSO. |
A Step-by-Step Protocol for Transient NBS-LRR Expression and Screening
1. Introduction Within the context of a broader thesis focused on Agrobacterium-mediated transient expression for the functional screening of plant NBS-LRR (Nucleotide-Binding Site-Leucine-Rich Repeat) immune receptors, precise vector design is paramount. This application note details the rationale for selecting genetic components and provides protocols for constructing vectors optimized for high-level, rapid protein expression in plant systems, such as Nicotiana benthamiana.
2. Promoter Selection for Transient Expression The choice of promoter dictates the timing, tissue specificity, and magnitude of transgene expression. For transient assays, strong constitutive promoters are preferred to achieve rapid, high-level protein accumulation to trigger or assess NBS-LRR-mediated immune responses.
Table 1: Comparison of Common Promoters for Transient Expression
| Promoter | Origin | Expression Profile | Relative Strength in N. benthamiana | Key Consideration for NBS-LRR Studies |
|---|---|---|---|---|
| CaMV 35S | Cauliflower mosaic virus | Constitutive, strong | 1.0 (Reference) | May be too strong, potentially causing autoactivity; often used with enhancer duplicates. |
| pNOS | Agrobacterium tumefaciens | Constitutive, weak to moderate | ~0.3-0.5 | Lower expression may help study weakly autoactive NBS-LRR variants. |
| pUBQ10 | Arabidopsis thaliana | Constitutive, strong | ~0.8-1.2 | Plant-derived, may offer more consistent expression across tissues. |
| pRPS5a | A. thaliana | Constitutive, strong | ~1.5-2.0 | Exceptionally strong, useful for expressing low-abundance interactors or reporters. |
3. Protein Tags and Their Applications Epitope and fluorescent tags are essential for detecting protein expression, localizing NBS-LRRs, and purifying complexes. Bipartite tags (e.g., split-YFP) are crucial for studying protein-protein interactions, such as NBS-LRR self-association or interactions with pathogen effectors.
Table 2: Common Protein Tags for NBS-LRR Functional Analysis
| Tag | Size (kDa) | Primary Application | Recommended Position | Protocol Reference |
|---|---|---|---|---|
| 3xFLAG | ~3.0 | Immunodetection, co-immunoprecipitation (Co-IP) | C-terminus | See Protocol 4.1 |
| eGFP/mCherry | ~27/28 | Subcellular localization, confocal microscopy | C-terminus | See Protocol 4.2 |
| Split-YFP (nYFP/cYFP) | ~17/10 | Bimolecular Fluorescence Complementation (BiFC) for interaction studies | C-terminus (both fragments) | See Protocol 4.3 |
| HA, Myc | ~1-2 | Immunodetection, Co-IP | N- or C-terminus | Standard Immunoblotting |
4. Detailed Experimental Protocols
Protocol 4.1: Co-Immunoprecipitation (Co-IP) for NBS-LRR Complex Analysis Objective: To isolate protein complexes containing a tagged NBS-LRR protein expressed transiently in N. benthamiana. Materials: See "The Scientist's Toolkit" below. Method:
- Infiltration: Infiltrate 4-week-old N. benthamiana leaves with Agrobacterium strains (OD600=0.5) harboring your NBS-LRR and candidate interactor constructs. Include empty vector controls.
- Harvest: Harvest leaf discs at 36-48 hours post-infiltration (hpi), flash-freeze in liquid N2, and store at -80°C.
- Extraction: Grind tissue to a fine powder. Homogenize in 2 mL/g of ice-cold Co-IP Buffer (50 mM Tris-HCl pH 7.5, 150 mM NaCl, 10% glycerol, 0.5% NP-40, 1x protease inhibitor cocktail). Centrifuge at 14,000xg for 20 min at 4°C.
- Pre-clearing: Incubate supernatant with 20 μL of pre-washed Protein A/G Agarose beads for 1 hour at 4°C. Centrifuge, collect supernatant.
- Immunoprecipitation: Add 2 μg of anti-FLAG antibody. Incubate for 2 hours. Add 30 μL of pre-washed Anti-FLAG M2 Magnetic Beads. Incubate for 1 hour.
- Washing: Pellet beads magnetically. Wash 4 times with 1 mL of Co-IP Buffer (without inhibitors).
- Elution: Elute proteins by boiling beads in 40 μL 2x Laemmli buffer for 5 min.
- Analysis: Analyze input (1% of extract) and IP samples by SDS-PAGE and immunoblotting with relevant antibodies.
Protocol 4.2: Confocal Microscopy for NBS-LRR Localization Objective: To determine the subcellular localization of a fluorescently tagged NBS-LRR protein. Method:
- Infiltration & Sampling: Infiltrate N. benthamiana leaves as in 4.1. At 24-48 hpi, excise small leaf sections.
- Mounting: Mount leaf sections in water on a glass slide, adaxial side down, with a coverslip.
- Imaging: Use a confocal laser scanning microscope. For eGFP: Ex 488 nm, Em 500-530 nm. For mCherry: Ex 587 nm, Em 610-650 nm. Include controls (empty vector, free fluorescent protein).
- Staining (Optional): For nuclear co-localization, infiltrate with DAPI (1 μg/mL) 30 min before imaging (Ex 405 nm).
Protocol 4.3: Bimolecular Fluorescence Complementation (BiFC) Assay Objective: To visualize in vivo protein-protein interactions in plant cells. Method:
- Vector Construction: Clone your NBS-LRR gene in-frame with the N-terminal half of YFP (nYFP) and your putative interacting partner with the C-terminal half (cYFP) in appropriate binary vectors.
- Co-infiltration: Co-infiltrate N. benthamiana leaves with Agrobacterium strains (OD600=0.3 each) carrying the nYFP and cYFP constructs. Include negative controls (partner + empty half).
- Imaging: At 48-72 hpi, image leaf sections using confocal microscopy (YFP channel: Ex 514 nm, Em 525-550 nm). Reconstituted YFP signal indicates interaction.
5. Visualization: Pathways and Workflows
Diagram Title: Vector Design to Functional Screening Workflow
Diagram Title: NBS-LRR Immune Signaling & Tagging Strategy
6. The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for Transient NBS-LRR Studies
| Reagent/Material | Supplier Examples | Function in Research |
|---|---|---|
| pEAQ-HT or pCAMBIA Binary Vectors | Addgene, CAMBIA | High-expression, modular T-DNA vectors for Agrobacterium transformation. |
| Gateway Cloning Kits (BP/LR Clonase) | Thermo Fisher | Enables rapid, recombination-based transfer of GOI into multiple destination vectors. |
| Anti-FLAG M2 Magnetic Beads | Sigma-Aldrich | For high-specificity, low-background immunoprecipitation of FLAG-tagged NBS-LRR proteins. |
| cOmplete Protease Inhibitor Cocktail | Roche | Protects expressed NBS-LRR proteins from degradation during extraction. |
| Agrobacterium tumefaciens Strain GV3101 | Lab Stock, CICC | Disarmed, virulent strain optimized for plant transformation and transient expression. |
| Silwet L-77 Surfactant | Lehle Seeds | Enhances Agrobacterium infiltration efficiency into leaf mesophyll. |
| DAPI Stain | Thermo Fisher | Nuclear counterstain for confocal microscopy co-localization studies. |
| Syringe (1 mL without needle) | Various | For precise manual infiltration of Agrobacterium suspension into leaves. |
1. Introduction Within the context of a thesis on Agrobacterium-mediated transient expression for NBS-LRR functional screening, the preparation of high-viability, virulent Agrobacterium tumefaciens cultures is the foundational step. This protocol details the selection of appropriate strains, culture media, and induction conditions to optimize T-DNA delivery for transient expression in plant leaves, a critical prerequisite for assessing NBS-LRR immune receptor function and its applications in plant disease resistance and drug discovery.
2. Key Agrobacterium Strains for Transient Expression The choice of strain is dictated by the vector system and desired transformation efficiency. The following table summarizes commonly used strains in contemporary research.
Table 1: Common Agrobacterium tumefaciens Strains for Transient Expression Studies
| Strain | Genetic Background | Key Features & Applications |
|---|---|---|
| GV3101 (pMP90) | C58 chromosomal background; Ti plasmid pMP90 (Gent⁺). | Disarmed, helper plasmid provides vir genes in trans. Excellent for transient expression in Nicotiana benthamiana; low polysaccharide production. |
| LBA4404 | Ach5 chromosomal background; Ti plasmid pAL4404 (Str⁺). | A classical disarmed strain. Slightly lower transformation efficiency than C58 derivatives but highly reliable for many plant species. |
| AGL-1 | C58 chromosomal background; Ti plasmid pTiBo542DT-DNA (Carb⁺). | Contains the "supervirulent" pTiBo542 Ti plasmid, providing enhanced vir gene activity. Often used for recalcitrant plant species. |
| EHA105 | C58 chromosomal background; Ti plasmid pEHA105 (Kan⁺). | Derived from the supervirulent strain EHA101. Provides high vir gene induction, leading to robust transient expression levels. |
3. Culture Media and Induction Conditions Optimal growth and virulence induction require specific media. Acetosyringone (AS) is the key phenolic compound used to induce the vir genes on the Ti plasmid.
Table 2: Standard Media and Induction Parameters
| Component/Condition | Standard Recipe/Condition | Purpose & Notes |
|---|---|---|
| Liquid Culture Media | LB, YEB, or YEP broth with appropriate antibiotics for the strain and binary vector (e.g., Gentamicin, Rifampicin, Kanamycin). | Supports robust bacterial growth. Maintains selection for both the helper Ti plasmid and the binary vector. |
| Induction Medium | MMA (MS salts, MES buffer, sucrose). pH adjusted to 5.6 prior to autoclaving. | Low-pH, sugar-rich environment that mimics wounded plant tissue. Serves as the base for acetosyringone addition. |
| Inducing Agent | Acetosyringone (AS), typically used at 150-200 µM final concentration. | The key phenolic signal that activates the VirA/VirG two-component system, inducing expression of all other vir genes. |
| Induction Duration | 2-6 hours at room temperature (22-28°C) with gentle agitation (e.g., 50-100 rpm). | Allows for full activation of the vir region and assembly of the T-pilus without excessive bacterial overgrowth. |
| Optical Density (OD₆₀₀) | Cells harvested at OD₆₀₀ ~0.5-1.0 for induction, then resuspended in MMA+AS to a final OD₆₀₀ of ~0.5-2.0 for infiltration. | Ensures bacteria are in log phase (high viability). Final OD is plant species and experiment-dependent. |
4. Detailed Protocol: Culture Preparation for Leaf Infiltration
Materials Required:
- Agrobacterium tumefaciens strain (e.g., GV3101) harboring the binary vector of interest.
- Appropriate liquid media (LB, YEB) and solid agar plates with antibiotics.
- Antibiotics stock solutions.
- Acetosyringone stock solution (100 mM in DMSO, stored at -20°C).
- MMA Induction Medium (10 mM MgCl₂, 10 mM MES, pH 5.6 with KOH, 100 µM acetosyringone).
- Sterile centrifuge tubes, shaking incubator, spectrophotometer.
Methodology: Day 1: Starter Culture
- Streak the Agrobacterium strain from a -80°C glycerol stock onto a selective agar plate. Incubate for 2 days at 28°C. Day 3: Primary Liquid Culture
- Pick a single colony and inoculate 2-5 mL of liquid media with the correct antibiotics. Grow overnight (16-20 hrs) at 28°C with vigorous shaking (220 rpm). Day 4: Secondary Culture & Induction
- Dilute the overnight culture 1:50 to 1:100 into fresh, antibiotic-containing liquid medium (e.g., 50 mL in a 250 mL flask). Grow at 28°C, 220 rpm, until the OD₆₀₀ reaches 0.5-1.0 (typically 4-6 hours).
- Harvest cells by centrifugation at room temperature (e.g., 3000 x g for 10-15 minutes).
- Gently resuspend the bacterial pellet in freshly prepared, room-temperature MMA induction medium (with 100-200 µM acetosyringone). Adjust the final OD₆₀₀ to the desired value (typically 0.5 for strong expression, up to 2.0 for effector-NBS-LRR interaction studies).
- Incubate the cell suspension at room temperature with gentle agitation (50-100 rpm) for 3-6 hours to induce the vir genes.
- The induced culture is now ready for infiltration into plant leaves (e.g., N. benthamiana) using a needleless syringe or vacuum infiltration. Perform infiltration within 24 hours.
5. The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for Agrobacterium Culture Preparation
| Reagent/Material | Function & Role in the Protocol |
|---|---|
| Acetosyringone (AS) | Phenolic plant signal molecule; binds to VirA sensor kinase to activate the vir gene regulon, essential for T-DNA transfer. |
| MES Buffer | Biological buffer used in MMA induction medium to maintain a stable acidic pH (5.6), mimicking the plant apoplast and enhancing vir induction. |
| Binary Vector System | Engineered plasmid containing the gene of interest (e.g., NBS-LRR or effector) between T-DNA borders and a plant selection marker. |
| Disarmed Helper Strain | Agrobacterium strain (e.g., GV3101) with a modified Ti plasmid providing vir genes in trans but lacking oncogenes, enabling transient expression without tumor formation. |
| Antibiotic Stocks (Rif, Gent, Kan) | Selective agents to maintain the helper Ti plasmid and binary vector in the bacterial culture, preventing plasmid loss. |
| MMA Induction Medium | A low-nutrient, acidic, and sugar-containing medium that stresses the bacteria and enhances their competence for T-DNA transfer post vir induction. |
6. Visualizing Key Processes
Title: Agrobacterium Culture and Infiltration Workflow
Title: Acetosyringone-Induced Vir Gene Activation Pathway
Within the broader thesis on Agrobacterium-mediated transient expression for functional screening of NBS-LRR (Nucleotide-Binding Site Leucine-Rich Repeat) immune receptors, selecting the optimal infiltration technique is critical. The method must efficiently deliver Agrobacterium tumefaciens (harboring the NBS-LRR construct of interest) into the appropriate plant tissue to elicit a robust, measurable phenotype without causing excessive damage. This application note compares the two primary techniques—syringe and vacuum infiltration—detailing their suitability for different plant tissues, with a focus on applications in NBS-LRR signaling research.
Comparison of Techniques: Key Parameters
The choice between syringe and vacuum infiltration depends on tissue type, scalability, and experimental objectives. The quantitative data below summarizes the core differences.
Table 1: Comparative Analysis of Syringe vs. Vacuum Infiltration
| Parameter | Syringe Infiltration | Vacuum Infiltration |
|---|---|---|
| Primary Tissue Target | Leaves (esp. Nicotiana benthamiana), Petioles | Seedlings, Whole Leaves, Root Systems, Thin Tissues |
| Infiltration Scale | Localized, spot-based (few cm²) | Whole-tissue or whole-seedling saturation |
| Typical Efficiency (Transformation) | 70-90% in infiltrated spot | 95-100% in susceptible tissues |
| Throughput | Low to medium (manual) | High (batch processing) |
| Tissue Damage Risk | Moderate (physical puncture) | Low (if vacuum/pressure optimized) |
| Bacterial Volume Used | Low (50-100 µl per spot) | High (50-200 ml per sample batch) |
| Ideal for NBS-LRR Screening | Hypersensitive Response (HR) localization, paired effector/ receptor tests | High-throughput protein expression, whole-tissue immune response assays, screening root tissues |
Detailed Protocols
Protocol 3.1: Syringe Infiltration for Leaf Transient Expression
Application: Localized delivery for HR cell death assays or protein-protein interaction studies in leaves.
Materials: See "The Scientist's Toolkit" (Section 5). Procedure:
- Grow Agrobacterium strain (e.g., GV3101 pSoup) carrying your NBS-LRR construct in selective media to late log phase (OD₆₀₀ = 0.8-1.2).
- Pellet cells at 4,000 x g for 10 min and resuspend in fresh infiltration medium (10 mM MES, 10 mM MgCl₂, 200 µM acetosyringone) to a final OD₆₀₀ of 0.4-0.8.
- Incubate the suspension at room temperature for 1-3 hours.
- Select fully expanded leaves of 4-5 week-old N. benthamiana plants.
- Using a 1 mL needleless syringe, gently press the tip against the abaxial (lower) leaf surface while supporting the leaf from the opposite side.
- Slowly infiltrate the bacterial suspension, watching the liquid front spread to form a water-soaked area (~1-3 cm diameter).
- Mark the infiltrated zone. Maintain plants under standard conditions (22-24°C, 16h light/8h dark) for 24-96 hours before analysis.
Protocol 3.2: Vacuum Infiltration for Whole Seedlings or Leaf Discs
Application: High-efficiency transformation of Arabidopsis seedlings for uniform NBS-LRR expression or immune signaling studies.
Materials: See "The Scientist's Toolkit" (Section 5). Procedure:
- Prepare Agrobacterium culture as in Protocol 3.1, steps 1-3.
- For Arabidopsis seedlings, grow vertically on agar plates for 5-7 days. For leaf discs, excise 1 cm² discs.
- Place seedlings or leaf discs into a beaker containing the Agrobacterium suspension.
- Transfer the beaker to a vacuum desiccator. Apply a gentle vacuum (10-25 in. Hg) for 2-5 minutes. The tissue should appear water-soaked as air is displaced.
- Rapidly release the vacuum. The sudden pressure change drives the suspension into intercellular spaces.
- Briefly blot the tissue on sterile paper and transfer to co-cultivation media plates (with acetosyringone).
- Co-cultivate in the dark at 22°C for 48-72 hours before transferring to analysis or selection plates.
Visualization of Workflows
Syringe Infiltration Protocol Flow
Vacuum Infiltration Protocol Flow
Technique Selection for NBS-LRR Screening
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Agrobacterium-Mediated Transient Assays
| Item | Function/Application |
|---|---|
| Agrobacterium Strain GV3101 (pMP90) | Disarmed, helper plasmid; provides virulence genes, low background in plants. |
| pSoup Plasmid (or pCH32) | Provides trans replication functions for binary vectors (e.g., pGreen, pCAMBIA). |
| Binary Vector (e.g., pGreenII, pEAQ-HT) | Carries gene of interest (NBS-LRR) between T-DNA borders for transfer into plant cells. |
| Acetosyringone | Phenolic compound that induces Agrobacterium vir gene expression, critical for T-DNA transfer. |
| Infiltration Buffer (10 mM MES, 10 mM MgCl₂) | Maintains bacterial viability and optimizes T-DNA transfer during infiltration. |
| Silwet L-77 (for vacuum) | Surfactant that lowers surface tension, improving wetting and infiltration efficiency. |
| Nicotiana benthamiana Seeds | Model plant with high susceptibility to Agrobacterium and suppressed RNAi (for high expression). |
| Luciferase Reporter Construct (e.g., pFRK1::LUC) | Immune-responsive reporter to quantify NBS-LRR activation post-infiltration. |
| Sterile Syringes (1 mL, needleless) | For applying localized pressure in syringe infiltration without damaging leaf tissue. |
| Vacuum Desiccator & Pump | For applying and rapidly releasing vacuum to drive bacterial suspension into tissues. |
Application Notes
Within the framework of a thesis on Agrobacterium-mediated transient expression for high-throughput NBS-LRR functional screening, co-infiltration is a critical, rapid, and powerful methodology. It enables the in planta characterization of immune receptor activation and the identification of pathogen virulence mechanisms. This approach bypasses the need for stable transformation, allowing for the parallel testing of dozens of NBS-LRR/effector pairs in a matter of days. The core principle involves the simultaneous delivery of two or more Agrobacterium tumefaciens strains into plant leaf tissue: one carrying the gene for the NBS-LRR receptor of interest and another carrying the gene for a putative pathogen effector or host suppressor protein. A visible hypersensitive response (HR), typically tissue collapse or necrosis, indicates specific recognition and receptor activation. The absence of HR suggests either no recognition or successful suppression of immunity by the effector.
This system is instrumental for:
- Validating Effector Recognition: Confirming direct or indirect physical interaction between an NBS-LRR and a pathogen effector.
- Effector Screenings: Screening candidate effectors from pathogen genomes for those recognized by specific NBS-LRRs.
- Functional Analysis of NBS-LRR Domains: Co-expressing effector proteins with NBS-LRR mutants to map functional domains required for recognition/activation.
- Identifying Suppressors of Cell Death: Screening pathogen effectors or host proteins that can suppress the HR triggered by a known NBS-LRR/effector pair, revealing virulence strategies.
Table 1: Example Quantitative Outcomes from a Co-infiltration Assay
| NBS-LRR Expressed | Putative Effector/Suppressor Expressed | HR Phenotype (Incidence) | Average HR Onset (hours post-infiltration) | Interpretation |
|---|---|---|---|---|
| Rx (Potato) | PVX CP (Positive control) | Strong Necrosis (95%) | 48 | Validated Recognition |
| Rx (Potato) | AvrPik (Negative control) | No HR (0%) | N/A | No Recognition |
| RPP1 (Arabidopsis) | ATR1Emoy2 | Strong Necrosis (98%) | 36 | Specific Recognition |
| RPP1 (Arabidopsis) | ATR1Noks1 | No HR (2%) | N/A | Effector Allele Not Recognized |
| NLRP3 (Chimeric) | EffectorA | Strong Necrosis (90%) | 60 | EffectorA Recognized |
| NLRP3 + EffectorA | SuppressorX | Suppressed HR (10%) | N/A | SuppressorX Inhibits Immunity |
Protocols
Protocol 1: Cloning Genes into Binary Vectors for Transient Expression
- Objective: Insert NBS-LRR and effector/suppressor genes into compatible binary expression vectors (e.g., pEAQ-HT, pBIN61, pGWB vectors).
- Methodology:
- Amplify coding sequences (without stop codon for tags) using high-fidelity PCR with appropriate restriction enzyme sites or Gateway attB sites.
- Purify PCR products and perform restriction digest/ligation or Gateway BP/LR recombination reactions according to manufacturer protocols.
- Transform resulting plasmids into E. coli for propagation, isolate plasmid DNA, and verify by sequencing.
- Transform verified plasmids into electrocompetent Agrobacterium tumefaciens strain GV3101 (pMP90) or LBA4404.
Protocol 2: Agrobacterium Culture Preparation for Co-infiltration
- Objective: Prepare adjusted cultures of Agrobacterium strains for infiltration.
- Methodology:
- Inoculate single colonies of each Agrobacterium strain (harboring NBS-LRR, effector, or empty vector control) into 5 mL of LB medium with appropriate antibiotics (e.g., kanamycin, rifampicin, gentamycin). Grow overnight at 28°C, 250 rpm.
- Sub-culture 1 mL of the overnight culture into 50 mL of fresh LB with antibiotics and 10 mM MES (pH 5.6). Grow to an OD600 of 0.8-1.0.
- Pellet cells at 4000 x g for 10 min at room temperature.
- Resuspend pellets in infiltration buffer (10 mM MgCl2, 10 mM MES pH 5.6, 150 µM acetosyringone) to a final OD600 of 0.5.
- Incubate resuspended cultures at room temperature in the dark for 2-4 hours.
- For co-infiltration, mix the adjusted bacterial suspensions in a 1:1 volume ratio. For effector-suppressor assays, a 1:1:1 ratio (NBS-LRR:Effector:Suppressor) is typical.
Protocol 3: Transient Co-infiltration inNicotiana benthamianaand Phenotyping
- Objective: Deliver mixed Agrobacterium cultures into plant leaves and monitor for HR.
- Methodology:
- Use 4-5 week-old N. benthamiana plants grown under controlled conditions.
- Using a 1 mL needleless syringe, press the tip against the abaxial side of a leaf and gently infiltrate the bacterial mixture. Mark infiltration zones.
- Maintain plants under normal growth conditions (22-24°C, 16h light/8h dark).
- Monitor infiltrated areas daily for 3-7 days. Document the presence/absence of HR (necrosis, tissue collapse) using photography.
- Key Control Infiltrations: a) NBS-LRR + Empty Vector, b) Effector + Empty Vector, c) Empty Vector + Empty Vector, d) Positive Control Pair (e.g., Rx/PVX CP).
- For quantification, HR incidence can be scored across multiple leaves (e.g., n=12 infiltrated spots per combination). Advanced phenotyping can include electrolyte leakage assays or trypan blue staining for dead cells.
Diagrams
Title: Co-infiltration Assay Workflow for NBS-LRR/Effector Recognition
Title: Effector Recognition and Suppression Pathways
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions for Co-infiltration Assays
| Item | Function & Rationale |
|---|---|
| Binary Vectors (pEAQ-HT, pBIN61) | High-expression, modular vectors for easy cloning of NBS-LRR/effector genes via Gibson Assembly or Gateway. |
| A. tumefaciens Strain GV3101 | Standard, disarmed strain with high transfection efficiency in N. benthamiana. Contains pMP90 (pTiC58) helper plasmid. |
| Acetosyringone | Phenolic compound that induces the Agrobacterium Vir genes, essential for efficient T-DNA transfer into plant cells. |
| Infiltration Buffer (10 mM MgCl₂, MES) | Maintains bacterial viability and optimizes pH for the Vir gene induction during the infiltration process. |
| Nicotiana benthamiana Plants | Model plant with a silenced RNAi defense system, enabling high-level transient protein expression and clear HR readouts. |
| Needleless Syringes (1 mL) | Standard tool for manual, low-pressure infiltration of bacterial suspensions into leaf intercellular spaces. |
| Electrolyte Leakage Conductivity Meter | Quantitative tool to measure ion leakage from damaged tissues, providing a numerical HR strength metric. |
| Trypan Blue Stain | Histochemical stain that selectively colors dead plant cells blue, visually confirming cell death during HR. |
Within the broader thesis on Agrobacterium-mediated transient expression for NBS-LRR (Nucleotide-Binding Site Leucine-Rich Repeat) functional screening research, optimizing the post-infiltration incubation period is a critical determinant of success. This application note details protocols and data for establishing the optimal timeline of expression to maximize the accumulation of functional NBS-LRR proteins and their signaling partners in plant tissues, such as Nicotiana benthamiana. Precise temporal control is essential for capturing dynamic immune responses and for high-throughput screening of effector-triggered immunity.
Table 1: Typical Protein Accumulation Dynamics in N. benthamiana Transient Expression
| Protein Class/Type | Onset of Detection (dpi*) | Peak Accumulation (dpi) | Notes & Key References |
|---|---|---|---|
| Reporter (e.g., GFP) | 1-2 | 3-4 | Standard benchmark for system health and infiltration efficiency. |
| Simple Agonists (e.g., Avr Proteins) | 2 | 3-5 | Effector proteins often stable; optimal for co-expression with NBS-LRRs at later time points. |
| NBS-LRR Receptors (Full-length) | 2-3 | 4-6 | Large, complex proteins requiring proper folding and subcellular localization. |
| Activated Defense Markers (e.g., PR1, ion leakage) | 3-4 | 4-7 | Downstream phenotypic readout; timing depends on receptor and effector pair. |
| Autoactive NBS-LRR Mutants | 2-3 | 3-5 | Often induce rapid hypersensitive response (HR), necessitating earlier harvest. |
*dpi: days post-infiltration
Table 2: Optimized Incubation Periods for Common Screening Objectives
| Research Objective | Recommended Incubation (dpi) | Rationale & Protocol Adjustment |
|---|---|---|
| Maximal Protein Yield (Biochemistry) | 4-5 | Balance between peak accumulation and onset of tissue senescence. |
| HR-Based Cell Death Scoring | 3-4 | Score before secondary necrosis or systemic symptoms obscure results. |
| Effector Recognition Screening | Co-infiltration: 4-5 | Express effector 1-2 d before NBS-LRR (staggered) or co-express at 4 dpi for simultaneous peak. |
| Downstream Signaling Analysis | 3-4 & 5-6 | Two time-point harvest recommended to capture early and late signaling events. |
| Protein-Protein Interaction (Co-IP) | 4 | Target peak co-accumulation of bait and prey proteins. |
Detailed Experimental Protocols
Protocol 1: Time-Course Optimization for a Novel NBS-LRR
Objective: Determine the optimal harvest window for accumulation of a transiently expressed NBS-LRR protein.
Materials: See "The Scientist's Toolkit" below. Procedure:
- Agrobacterium Preparation: Transform your NBS-LRR construct (in a binary vector, e.g., pEAQ-HT) into Agrobacterium tumefaciens strain GV3101 (pSoup). Select single colonies on appropriate antibiotics (e.g., Rifampicin, Kanamycin, Gentamicin).
- Starter Culture: Inoculate 5 mL of LB medium with antibiotics. Grow at 28°C, 200 rpm for 24-48 hrs.
- Induction Culture: Dilute starter 1:100 into 50 mL of fresh LB with antibiotics and 10 mM MES pH 5.6. Add Acetosyringone to a final concentration of 200 µM. Induce at 28°C, 200 rpm for 18-24 hrs.
- Harvest & Resuspension: Pellet cells at 4000 x g for 10 min. Resuspend to a final OD₆₀₀ of 0.5-1.0 in infiltration buffer (10 mM MgCl₂, 10 mM MES pH 5.6, 200 µM Acetosyringone). Incubate at room temperature for 2-4 hrs.
- Plant Infiltration: Infiltrate the abaxial side of 4-6 week-old N. benthamiana leaves using a needleless syringe. Infiltrate multiple leaves/plants to allow for destructive sampling over time.
- Time-Course Harvest: Label plants/leaves. Harvest leaf discs from the infiltrated zones at the following dpi: 1, 2, 3, 4, 5, 6, 7. Flash-freeze immediately in liquid N₂. Store at -80°C.
- Analysis: Analyze samples by:
- Immunoblotting: Detect protein accumulation using tag-specific (e.g., anti-His, anti-HA) or protein-specific antibodies.
- Phenotypic Scoring: Visually document HR or other symptoms daily.
- Functional Assay: e.g., Ion leakage assay from leaf discs at each time point.
Protocol 2: Staggered Co-Infiltration for Effector Recognition Studies
Objective: To co-express an NBS-LRR and a candidate effector, ensuring both proteins peak simultaneously to maximize detection of recognition.
Procedure:
- Prepare Agrobacterium cultures for the NBS-LRR construct and the effector construct separately, as in Protocol 1, Steps 1-4.
- Day 0: Infiltrate the effector construct (OD₆₀₀=0.4) alone into one leaf sector.
- Day 2: Infiltrate the NBS-LRR construct (OD₆₀₀=0.6) into the same leaf sector. This "booster" infiltration will take up the NBS-LRR strain into tissues already primed with the effector.
- Alternative (Single Time Point): Mix both induced cultures in a 1:1 ratio (adjusting ODs as needed) and co-infiltrate simultaneously at day 0. This is less precise but suitable for high-throughput initial screens.
- Harvest: Monitor daily for HR. Harvest tissue for analysis at 4-5 days after the second infiltration (i.e., 6-7 dpi from Day 0).
Signaling Pathways & Workflow Visualizations
Diagram Title: NBS-LRR Screening Workflow & Signaling
Diagram Title: Protein Accumulation Timeline & Harvest Windows
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Transient Expression Optimization
| Item | Function & Rationale | Example/Supplier |
|---|---|---|
| Agrobacterium Strain GV3101 (pSoup) | Standard disarmed strain for transient expression; pSoup provides vir genes in trans for efficient T-DNA transfer. | Common lab strain, available from biological resource centers. |
| Binary Expression Vector (e.g., pEAQ-HT) | High-expression vector utilizing CPMV-HT system, yields very high levels of recombinant protein. | (Lomonossoff et al., 2009) |
| Acetosyringone | Phenolic compound that induces the Agrobacterium vir genes, essential for T-DNA transfer competence. | Sigma-Aldrich, D134406. Prepare 200 mM stock in DMSO. |
| Infiltration Buffer (10 mM MgCl₂, 10 mM MES, pH 5.6) | Provides optimal ionic and pH conditions for Agrobacterium attachment and gene transfer. | Prepare fresh, filter sterilize. |
| Nicotiana benthamiana Plants | Model plant with high susceptibility to Agrobacterium, lacks silencing machinery for some viruses (e.g., RNAi mutants). | Grow at 22-25°C, 16-hr light, for 4-6 weeks. |
| Tag-Specific Antibodies (Anti-His, HA, GFP) | Critical for detecting and quantifying transiently expressed proteins via immunoblot or ELISA. | Commercial sources: Thermo Fisher, Roche, Abcam. |
| Conductivity Meter | For quantifying ion leakage, a quantitative and sensitive measure of the hypersensitive response (HR) cell death. | Essential equipment for functional screening. |
| RNase Inhibitors & Protease Inhibitor Cocktails | Preserve RNA and protein integrity during extraction for downstream signaling analysis. | e.g., RNasin, Complete Tablets (Roche). |
In the functional screening of Nucleotide-Binding Site Leucine-Rich Repeat (NBS-LRR) immune receptors, Agrobacterium-mediated transient expression (agroinfiltration) in plants like Nicotiana benthamiana is a cornerstone technique. It enables rapid, high-throughput assessment of receptor autoactivity or specific recognition of pathogen effectors. A primary phenotypic readout for successful immune activation is the Hypersensitive Response (HR), a form of programmed cell death (PCD) localized to the site of pathogen recognition. Quantitative and qualitative measurement of HR and cell death is therefore critical for validating NBS-LRR function, mapping effector recognition, and studying signaling pathways.
Key Quantitative Metrics for HR/Cell Death Assessment
The following table summarizes the primary quantitative methods used to measure HR and cell death, detailing their outputs, advantages, and limitations.
Table 1: Comparative Analysis of HR and Cell Death Measurement Assays
| Method | Measured Parameter | Output Data Type | Typical Time Post-Infiltration | Throughput | Key Advantages | Key Limitations |
|---|---|---|---|---|---|---|
| Visual Scoring | Area & Intensity of Necrosis | Ordinal Scale (e.g., 0-5) | 24-72 hours | Very High | Simple, fast, scalable for initial screening. | Subjective, low resolution, requires experience. |
| Ion Leakage (Conductivity) | Loss of Membrane Integrity | Quantitative (µS/cm) | 16-48 hours | Medium | Objective, quantitative, kinetic data possible. | Destructive, background from wounding, medium-throughput. |
| Evans Blue Staining | Loss of Membrane Integrity | Quantitative (Absorbance/Image Analysis) | 20-48 hours | Medium-High | Visual confirmation, can be quantified. | Destructive, multiple steps, dye can be toxic. |
| Trypan Blue Staining | Dead Cell Penetration | Qualitative/ Semi-Quant. (Image Analysis) | 24-72 hours | Medium | Classic for microscopy, stains dead cells blue. | Destructive, not easily scalable to 96-well format. |
| Fluorescein Diacetate (FDA) Staining | Esterase Activity in Live Cells | Quantitative (Fluorescence) | 20-48 hours | Medium-High | Measures viability inversely; live cells fluoresce. | Signal can be transient, requires optimization. |
| Luciferase Imaging | Suppression of Cytoplasmic Luciferase Activity | Quantitative (Light Units) | 24-48 hours | High | Non-destructive, in planta kinetics, high sensitivity. | Requires co-expression, equipment cost, substrate needed. |
Detailed Experimental Protocols
Protocol 3.1: Agroinfiltration for HR Assay inN. benthamiana
Objective: Transient expression of NBS-LRR constructs or effector pairs to elicit HR.
Materials:
- Agrobacterium tumefaciens strain GV3101 (pSoup) carrying expression vector (e.g., pEAQ, pBIN61).
- N. benthamiana plants (4-5 weeks old).
- Infiltration buffer (10 mM MES, 10 mM MgCl₂, 150 µM acetosyringone, pH 5.6).
- 1 mL needleless syringe.
Method:
- Grow Agrobacterium overnight in appropriate antibiotics at 28°C.
- Pellet cultures, resuspend in infiltration buffer to a final OD₆₀₀ of 0.4-0.8 (adjust based on construct strength).
- Incubate resuspension at room temperature for 1-3 hours.
- For co-infiltration (e.g., R-Avr), mix bacterial suspensions 1:1.
- Using a syringe, gently press the tip against the abaxial side of a leaf and infiltrate the suspension. Mark infiltration zones.
- Maintain plants under standard conditions (22-24°C, 16h light).
- Monitor for HR symptoms starting at 18-24 hours post-infiltration (hpi).
Protocol 3.2: Quantitative Ion Leakage Assay
Objective: Objectively quantify cell death by measuring electrolyte leakage from leaf discs.
Materials:
- Leaf tissue from infiltration zones.
- Cork borer or sharp punch (e.g., 6-8 mm diameter).
- 50 mL conical tubes or 24-well plates with 0.9% NaCl or distilled H₂O.
- Conductivity meter.
Method:
- At designated timepoints (e.g., 24 hpi), harvest leaf discs from the center of infiltration zones using a cork borer. Include control (e.g., empty vector) discs.
- Rinse discs briefly in distilled water to remove surface ions.
- Place 4-6 discs in a tube containing 20 mL of 0.9% NaCl. Ensure discs are fully submerged.
- Incubate with gentle shaking (50 rpm) at room temperature.
- Measure the initial conductivity of the solution (C₀).
- Continue incubation, measuring conductivity at regular intervals (e.g., 2, 4, 6, 8, 24 hours).
- After final measurement, autoclave or boil samples for 10 minutes to kill all cells, cool, and measure total conductivity (Cₜ).
- Calculate relative ion leakage:
(Cₓ - C₀) / (Cₜ - C₀) * 100%, where Cₓ is conductivity at time x.
Protocol 3.3: Evans Blue Uptake Quantification Assay
Objective: Quantify dead cells based on uptake and retention of Evans Blue dye.
Materials:
- Evans Blue dye stock (0.5% w/v in water).
- Leaf tissue or discs.
- 1 mL microcentrifuge tubes.
- Incubator at 37°C.
- Spectrophotometer or plate reader.
- Washing solution: 1% SDS in 50% methanol.
Method:
- Harvest leaf discs (as in 3.2) and place in a microcentrifuge tube.
- Add 1 mL of 0.1% Evans Blue (diluted from stock in water). Incubate for 30 minutes at room temperature with gentle agitation.
- Remove dye solution. Wash discs extensively with distilled water (4-5 times, 5 min each) until no blue color is seen in the wash from control (healthy) discs.
- Add 1 mL of washing solution (1% SDS in 50% methanol). Homogenize discs or incubate at 37°C for 30 minutes with vortexing to extract the dye.
- Centrifuge at 12,000 x g for 5 minutes.
- Measure absorbance of the supernatant at 600 nm. Higher absorbance correlates with greater cell death.
Visualizing Pathways and Workflows
Title: HR Screening Workflow for NBS-LRR Function
Title: Key Signaling Events in Hypersensitive Response
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for HR/ Cell Death Assays
| Item | Function & Application in HR Research | Example/Notes |
|---|---|---|
| pEAQ-HT Expression Vector | High-level transient protein expression in plants via agroinfiltration. Minimizes gene silencing. | Critical for strong NBS-LRR or effector expression to elicit clear HR. |
| GV3101 (pSoup) A. tumefaciens | Standard disarmed strain for agroinfiltration. pSoup provides trans replication functions. | Compatible with binary vectors containing ColE1 origin. |
| Acetosyringone | Phenolic inducer of Agrobacterium vir genes. Enhances T-DNA transfer efficiency. | Added to infiltration buffer (150-200 µM). |
| Syringe (1mL, needleless) | Tool for pressure-based infiltration of bacterial suspension into leaf mesophyll. | Creates a defined, reproducible infiltration zone. |
| Conductivity Meter | Measures electrolyte concentration in solution for ion leakage assays. | Provides quantitative, kinetic data on membrane integrity loss. |
| Evans Blue Dye | Diazo dye excluded by intact plasma membranes; taken up and retained by dead cells. | Quantifiable via spectrophotometry after extraction. |
| Fluorescein Diacetate (FDA) | Cell-permeant esterase substrate. Live cells cleave it to fluorescent fluorescein. | Measures viability inversely; loss of fluorescence indicates cell death. |
| Luciferase Reporter (e.g., FLuc) | Co-expressed cytoplasmic reporter. Cell death/translation shutoff reduces luminescence. | Enables non-destructive, kinetic imaging of HR progression in planta. |
| cOmplete Protease Inhibitor | Inhibits proteases released during cell death, preserving proteins for subsequent analysis (e.g., immunoblot). | Added during tissue homogenization if protein analysis follows. |
Within the broader thesis investigating Agrobacterium-mediated transient expression for high-throughput functional screening of plant NBS-LRR immune receptors, biochemical validation is a critical step. This document details the application notes and protocols for protein extraction and immunodetection following agroinfiltration of Nicotiana benthamiana leaves. Successful validation confirms the expression, stability, and often the expected modification (e.g., phosphorylation, cleavage) of the transfected NBS-LRR protein, correlating phenotypic observations (e.g., hypersensitive response) with molecular data.
Key Research Reagent Solutions
| Reagent / Material | Function in Protocol |
|---|---|
| Agrobacterium tumefaciens (Strain GV3101) | Vector for transient gene delivery into plant cells. |
| Silwet L-77 | Surfactant that enhances leaf wetting and infiltration efficiency. |
| c-Myc or FLAG Epitope Tag | Engineered peptide sequence fused to the NBS-LRR gene for universal immunodetection. |
| Anti-c-Myc/FLAG Primary Antibody | Binds specifically to the expressed epitope-tagged protein of interest. |
| HRP-conjugated Secondary Antibody | Binds to the primary antibody for chemiluminescent detection. |
| Enhanced Chemiluminescence (ECL) Substrate | Enzymatic reaction produces light signal for blot imaging. |
| Plant Protease Inhibitor Cocktail | Prevents degradation of extracted proteins during sample preparation. |
| Phosphatase Inhibitors (e.g., NaF, β-glycerophosphate) | Preserves phosphorylation status of signaling proteins. |
| PVDF or Nitrocellulose Membrane | Solid support for immobilized proteins during immunoblotting. |
| Loading Control Antibody (e.g., anti-Rubisco, anti-Actin) | Verifies equal protein loading across gel lanes. |
Detailed Protocols
Protocol A: Total Protein Extraction from InfiltratedN. benthamianaLeaf Discs
This protocol is optimized for extracting soluble proteins, including NBS-LRRs, 2-4 days post-infiltration.
Materials:
- Liquid Nitrogen
- Extraction Buffer (freshly added): 50 mM Tris-HCl (pH 7.5), 150 mM NaCl, 1 mM EDTA, 10% Glycerol, 1% (v/v) IGEPAL CA-630, 1x Plant Protease Inhibitor Cocktail, 1 mM PMSF, 5 mM DTT, 10 mM NaF, 1 mM Na3VO4.
- Mortar and pestle (pre-chilled)
- Microcentrifuge (4°C)
Method:
- Harvesting: Excise leaf discs from infiltrated zones using a cork borer (e.g., 1 cm diameter). Immediately flash-freeze in liquid nitrogen. Store at -80°C if not processing immediately.
- Grinding: Using a pre-chilled mortar and pestle, grind tissue to a fine powder under liquid nitrogen.
- Extraction: Transfer ~100 mg of powder to a pre-chilled 1.5 mL microcentrifuge tube. Add 300 µL of ice-cold Extraction Buffer. Vortex vigorously for 10 seconds.
- Incubation: Incubate on ice for 15 minutes, vortexing briefly every 5 minutes.
- Clarification: Centrifuge at 16,000 × g for 20 minutes at 4°C.
- Collection: Carefully transfer the supernatant (total protein extract) to a new pre-chilled tube. Place on ice.
- Quantification: Determine protein concentration using a Bradford or BCA assay. Proceed to SDS-PAGE or store at -80°C.
Protocol B: Immunoblotting (Western Blot) for Tagged NBS-LRR Detection
Materials:
- SDS-PAGE Gel (4-20% gradient recommended)
- Wet or Semi-Dry Transfer System
- Blocking Buffer: 5% (w/v) non-fat dry milk in TBST (Tris-Buffered Saline with 0.1% Tween-20).
- Primary Antibody Dilution Buffer: 1% BSA in TBST.
- Secondary Antibody Dilution Buffer: 1% non-fat dry milk in TBST.
Method:
- Electrophoresis: Dilute 20-40 µg of total protein extract with Laemmli buffer. Denature at 95°C for 5 min. Load samples and appropriate molecular weight markers on the gel. Run at constant voltage until dye front reaches bottom.
- Transfer: Using a wet transfer system, transfer proteins to a PVDF membrane (pre-activated in methanol) at 100V for 60-90 minutes on ice.
- Blocking: Block membrane with 10 mL Blocking Buffer for 1 hour at room temperature with gentle agitation.
- Primary Antibody Incubation: Dilute anti-tag primary antibody (e.g., anti-c-Myc, 1:3000) in Primary Antibody Dilution Buffer. Incubate membrane with antibody solution overnight at 4°C with gentle agitation.
- Washing: Wash membrane 3 times for 10 minutes each with 15 mL TBST.
- Secondary Antibody Incubation: Dilute HRP-conjugated secondary antibody (e.g., anti-mouse, 1:5000) in Secondary Antibody Dilution Buffer. Incubate membrane for 1 hour at room temperature.
- Washing: Repeat step 5.
- Detection: Apply ECL substrate to the membrane according to manufacturer's instructions. Image using a chemiluminescence documentation system.
Data Presentation: Representative Quantitative Analysis
Table 1: Comparative Protein Expression Levels of Mutant NBS-LRR Variants Data from a representative experiment 3 days post-infiltration (n=4 biological replicates). Signal intensity normalized to Actin control and empty vector (EV) set to 1. HR = Hypersensitive Response phenotype observed.
| Construct (c-Myc tagged) | Relative Protein Expression (Mean ± SD) | HR Phenotype (Y/N) | Notes |
|---|---|---|---|
| EV (pGD) | 1.0 ± 0.2 | N | Background control |
| WT NBS-LRR | 8.5 ± 1.3 | Y | Strong accumulation |
| Kinase-dead (K->A) | 7.9 ± 1.1 | N | Stable, non-functional |
| P-loop Mutant (G->A) | 2.1 ± 0.5 | N | Severely reduced accumulation |
| Autoactive ΔLRR | 12.4 ± 2.0 | Y | Enhanced accumulation |
Table 2: Effect of Proteasome Inhibitor (MG132) on NBS-LRR Stability Leaf discs harvested 40 hours post-infiltration, treated with 50 µM MG132 or DMSO for 8 hours prior to extraction (n=3).
| Construct | Treatment | Relative Protein Expression (Mean ± SD) | Fold Change vs. DMSO |
|---|---|---|---|
| WT NBS-LRR | DMSO | 7.2 ± 0.9 | 1.0 |
| WT NBS-LRR | MG132 | 14.1 ± 1.7 | 2.0 |
| P-loop Mutant | DMSO | 1.8 ± 0.4 | 1.0 |
| P-loop Mutant | MG132 | 5.5 ± 0.8 | 3.1 |
Visualization: Workflow and Pathway Diagrams
Title: Protein Extraction Workflow Post-Agroinfiltration
Title: Biochemical Validation in NBS-LRR Signaling Pathway
Solving Common Problems: Maximizing Signal and Reproducibility
Within the context of a broader thesis on Agrobacterium-mediated transient expression for NBS-LRR functional screening in plants, the frequent challenge of low or no transgene expression can halt research progress. This Application Note details a systematic troubleshooting framework to diagnose failures in T-DNA delivery and subsequent transcriptional events, enabling researchers to rescue experiments and obtain reliable data for plant immunity studies and drug discovery pipelines.
Troubleshooting Framework: A Diagnostic Roadmap
The following flowchart outlines a logical, step-by-step approach to isolate the cause of expression failure.
Title: Troubleshooting Workflow for Failed Transient Expression
Key Experimental Protocols for Diagnosis
Protocol 1: Rapid Agrobacterium Viability and Plasmid Integrity Check
Purpose: To confirm the Agrobacterium tumefaciens strain (e.g., GV3101, AGL1) is healthy and harbors the correct, intact binary vector before plant infiltration.
Materials: LB broth/plates with appropriate antibiotics (e.g., rifampicin, gentamicin, spectinomycin), PCR reagents, plasmid isolation kit, primers for vector backbone and insert.
Procedure:
- Re-streak & Grow: Streak glycerol stock onto selective LB agar plates. Incubate at 28°C for 2 days.
- Liquid Culture: Inoculate a single colony into 5 mL selective LB broth. Shake at 28°C, 200 rpm for 24-48h.
- Plasmid Isolation: Isolate plasmid from 2 mL of culture using a standard mini-prep kit (e.g., alkaline lysis).
- Diagnostic PCR: Perform PCR using:
- Backbone check: Primers flanking the T-DNA border or within the vector backbone.
- Insert check: Gene-specific primers for your NBS-LRR gene of interest.
- Restriction Digest: Digest the isolated plasmid with 1-2 enzymes that release the insert. Analyze fragment sizes on an agarose gel against expected sizes.
Expected Results: A single band of correct size in both PCR and digest confirms plasmid integrity.
Protocol 2: Histochemical GUS Assay for T-DNA Delivery Verification
Purpose: To visually confirm successful T-DNA transfer and nuclear import in infiltrated leaf tissue using the β-glucuronidase (GUS) reporter.
Materials: Infiltrated leaf tissue, GUS staining solution (1 mM X-Gluc, 50 mM sodium phosphate buffer pH 7.0, 0.1% Triton X-100, 0.5 mM potassium ferricyanide/ferrocyanide), ethanol series (70%-100%), sterile water.
Procedure:
- Infiltrate: Co-infiltrate your experimental construct with a constitutive promoter (e.g., 35S)-driven GUS reporter construct, or use a binary vector where GUS is part of the T-DNA.
- Incubate: Allow expression for 24-48 hours post-infiltration (hpi).
- Stain: Submerge leaf discs in GUS staining solution. Vacuum infiltrate for 5-10 minutes to ensure solution penetration.
- Incubate: Place samples at 37°C in the dark for 4-24 hours.
- Destain: Replace staining solution with 70% ethanol to remove chlorophyll. Change ethanol until tissue is clear.
- Image: Observe under a dissecting microscope for blue precipitate indicating GUS activity.
Expected Results: Uniform blue staining across the infiltrated area confirms successful T-DNA delivery.
Protocol 3: RT-qPCR for Transgene Transcript Quantification
Purpose: To quantitatively assess if the delivered T-DNA is being transcribed, distinguishing delivery failure from transcriptional silencing.
Materials: RNA extraction kit (e.g., TRIzol), DNase I, reverse transcription kit, qPCR master mix, gene-specific primers for transgene and endogenous control (e.g., EF1α, UBQ).
Procedure:
- Sample Harvest: Harvest infiltrated leaf tissue at expected peak expression (e.g., 48-72 hpi). Flash-freeze in liquid N₂.
- RNA Extraction: Grind tissue and extract total RNA. Treat with DNase I to remove genomic DNA contamination.
- cDNA Synthesis: Perform reverse transcription on 1 µg of total RNA using an oligo(dT) or random hexamer primer.
- qPCR Setup: Prepare reactions in triplicate containing cDNA template, gene-specific primers for:
- Target: Your NBS-LRR transgene (design primers spanning an intron or the terminator to avoid genomic DNA amplification).
- Reference: A stable endogenous plant gene.
- Control: A known highly expressed transgene (e.g., 35S::GFP) to confirm system functionality.
- Run & Analyze: Perform qPCR using a standard cycling protocol. Calculate relative expression (∆∆Cq) normalized to the endogenous reference.
Expected Results: High Cq values (>35) or undetectable signal for the transgene, alongside a strong control signal, indicate a transcriptional block.
Table 1: Common Causes and Diagnostic Results for Low/No Expression
| Problem Category | Specific Issue | Diagnostic Test (See Protocol) | Expected Diagnostic Result |
|---|---|---|---|
| Bacterial/Vector | Agrobacterium culture senescence | Protocol 1 (Re-streak) | Poor or no bacterial growth |
| Plasmid rearrangement/loss | Protocol 1 (PCR/Digest) | Incorrect PCR product or restriction pattern | |
| T-DNA Delivery | Inefficient infiltration | Protocol 2 (GUS) | Patchy or absent GUS staining |
| Suppressed vir gene induction | Co-infiltration with GUS reporter | GUS positive, experimental construct negative | |
| Transcription | Promoter incompatibility | Protocol 3 (RT-qPCR) | Low transgene mRNA, high control mRNA |
| Gene Silencing (PTGS/TGS) | Protocol 3 (RT-qPCR + time course) | mRNA high initially, then declines sharply | |
| Post-Transcriptional | Toxic protein effect | Protocol 3 & Western Blot | mRNA present, protein absent |
| Codon bias/improper folding | Protein assay with epitope tag | Protein detected but non-functional |
Table 2: Optimization Parameters for Enhanced Transient Expression
| Parameter | Typical Range for NBS-LRR Screening | Optimization Recommendation |
|---|---|---|
| OD₆₀₀ of Infiltration Culture | 0.2 - 1.0 | Test 0.4, 0.6, 0.8 for each new construct |
| Acetosyringone Concentration | 100 - 500 µM | Include 200 µM in both induction medium and infiltration buffer |
| Plant Age/Growth Condition | 3-5 week-old leaves | Use consistent, optimal light/temperature |
| Co-cultivation Time | 48 - 72 hours | Harvest multiple time points to determine peak |
| Suppressor of Silencing (p19, etc.) | Co-infiltration ratio 1:1 | Always include as a standard practice to boost expression |
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Troubleshooting | Example/Brand |
|---|---|---|
| Binary Vector with Reporter | Visual confirmation of T-DNA delivery. | pBIN-GFP, pCAMBIA1301 (GUS) |
| Virulence Gene Inducer | Enhances vir gene expression, critical for T-DNA transfer. | Acetosyringone (Sigma-Aldrich) |
| RNA Silencing Suppressor | Co-expressed to inhibit post-transcriptional gene silencing (PTGS). | Tomato Bushy Stunt Virus p19 protein |
| Epitope Tag Antibodies | Detect low-abundance or unstable proteins via Western blot. | Anti-HA, Anti-FLAG, Anti-Myc |
| Protease Inhibitor Cocktail | Prevents protein degradation during extraction. | cOmplete (Roche) |
| Positive Control Plasmid | Construct with known high expression to validate entire system. | 35S::GFP or 35S::Luciferase |
| High-Efficiency Agrobacterium Strain | Optimized for transient expression in specific hosts (e.g., Nicotiana). | GV3101 (pMP90), AGL-1 |
Signaling and Transcriptional Silencing Pathways
Understanding host pathways that limit transgene expression is critical. Below is a simplified diagram of key plant responses to Agrobacterium infection and transgene expression that can lead to silencing.
Title: Plant Pathways Limiting Transient Transgene Expression
The Critical Role of Silencing Suppressors (e.g., p19, HC-Pro)
Within the framework of Agrobacterium-mediated transient expression for functional screening of plant Nucleotide-Binding Site Leucine-Rich Repeat (NBS-LRR) immune receptors, achieving high-level, rapid protein expression is paramount. A major bottleneck is Post-Transcriptional Gene Silencing (PTGS), an antiviral defense mechanism plants deploy against transgene mRNA. Viral silencing suppressors (VSRs) are proteins that inhibit PTGS, thereby stabilizing transgene mRNA and dramatically enhancing recombinant protein accumulation.
The integration of VSRs, notably Tombusvirus p19 and Potyvirus HC-Pro, is now a standard co-expression strategy in transient assays. This is critical for NBS-LRR screening, as these receptors often require precise stoichiometry and high accumulation levels to trigger measurable immune responses (e.g., hypersensitive response, HR) upon recognition of their cognate effectors. Suppressing silencing ensures consistent, robust expression necessary for reliable phenotype scoring in high-throughput formats.
Key Quantitative Benefits of Silencing Suppressors: Research consistently shows that co-expression of VSRs can increase recombinant protein yield by 10- to 50-fold in Nicotiana benthamiana leaf transient expression systems, directly translating to more sensitive and reproducible functional assays.
Comparative Data on Common Silencing Suppressors
Table 1: Characteristics and Efficacy of Key Viral Silencing Suppressors
| Suppressor | Viral Origin | Primary Mechanism | Typical Yield Increase* | Key Advantage | Consideration for NBS-LRR Screening |
|---|---|---|---|---|---|
| p19 | Tomato bushy stunt virus | Sequesters 21-nt siRNA duplexes, preventing RISC assembly. | 25- to 50-fold | Potent, well-characterized, minimal off-target effects on plant physiology. | Ideal for most screens; provides clean, high-level expression of the NBS-LRR/effector pair. |
| HC-Pro | Tobacco etch virus | Binds and inhibits RISC, may also inhibit siRNA methylation. | 10- to 25-fold | Broad-spectrum inhibition, can enhance synergistic expression. | May slightly alter plant defense signaling; use when p19 is less effective. |
| p25 | Potato virus X | Inhibits systemic silencing, less potent locally. | 5- to 10-fold | Useful for systemic spread of expression. | Not recommended for localized, high-intensity transient assays. |
| 2b | Cucumber mosaic virus | Binds AGO1 to inhibit slicing, moves systemically. | 10- to 20-fold | Nuclear localization may affect specific pathways. | Potential for stronger perturbation of endogenous miRNA activity. |
Yield increase is for recombinant protein expression relative to no suppressor control in *N. benthamiana. Actual fold-change depends on the protein of interest.
Experimental Protocols
Protocol 1: Co-infiltration ofAgrobacteriumHarboring Suppressor and Gene-of-Interest Constructs for NBS-LRR Screening
Objective: To transiently express an NBS-LRR receptor and its putative cognate effector in the presence of silencing suppressor p19 for functional phenotype assessment.
Materials: See "The Scientist's Toolkit" below.
Method:
- Strain Preparation: Inoculate separate cultures of A. tumefaciens strain GV3101 (pMP90) harboring the following binary vectors:
- Strain A: NBS-LRR gene under a strong plant promoter (e.g., 35S).
- Strain B: Candidate effector gene (35S promoter).
- Strain C: p19 silencing suppressor gene (35S promoter).
- Culture & Induction: Grow cultures at 28°C in LB with appropriate antibiotics to OD600 ~1.5. Pellet cells (4000 x g, 10 min) and resuspend in infiltration buffer (10 mM MES, 10 mM MgCl2, 150 µM acetosyringone, pH 5.6). Adjust final OD600 to 0.5 for each strain.
- Mix Preparation: Combine the bacterial suspensions in a 1:1:1 ratio (NBS-LRR : Effector : p19). For controls, prepare mixtures:
- Test: NBS-LRR + Effector + p19.
- Receptor Control: NBS-LRR + Empty Vector + p19.
- Effector Control: Empty Vector + Effector + p19.
- Infiltration: Incubate mixes at room temperature for 1-3 hours. Using a needleless syringe, infiltrate the mixtures into the abaxial side of leaves of 4-5 week-old N. benthamiana plants. Mark infiltration zones.
- Phenotyping: Monitor infiltrated areas daily for 3-7 days for HR development (tissue collapse, bleaching). Document phenotypes visually and by measuring ion leakage or via fluorescent dyes (e.g., Evans Blue).
Protocol 2: Quantifying Suppressor Efficacy via Co-expression with a Reporter Gene
Objective: To empirically determine the optimal suppressor for your system by measuring its effect on reporter protein accumulation.
Method:
- Prepare Agrobacterium strains carrying GUS (β-glucuronidase) or GFP reporter and candidate suppressor (p19, HC-Pro, etc.) as in Protocol 1.
- Co-infiltrate reporter strain (OD600 0.5) with individual suppressor strains (OD600 0.5) or empty vector control.
- Harvest leaf discs at 3 days post-infiltration (dpi).
- Quantification:
- For GUS: Homogenize tissue in GUS extraction buffer. Perform a fluorometric assay using 4-MUG as substrate. Measure fluorescence (excitation 365 nm, emission 455 nm). Calculate relative activity.
- For GFP: Extract total soluble protein, separate by SDS-PAGE, and perform immunoblotting with anti-GFP antibodies, comparing band intensity.
Signaling Pathways and Workflow Visualizations
Title: PTGS Pathway & Suppressor Mechanism
Title: Transient NBS-LRR Screening Workflow
The Scientist's Toolkit
Table 2: Essential Research Reagents and Materials
| Item | Function in Experiment | Example/Details |
|---|---|---|
| Agrobacterium tumefaciens GV3101 | Disarmed vector for T-DNA delivery into plant cells. | Often carries helper plasmid pMP90 (rifampicin, gentamicin resistant). |
| Binary Vector (e.g., pEAQ, pBIN) | Plant expression vector containing gene of interest between T-DNA borders. | Includes strong promoter (35S), terminator, and bacterial selectable marker. |
| p19 Expression Vector | Standardized source of silencing suppressor. | e.g., pBIN61-p19 or pEAQ-HT-p19. |
| Acetosyringone | Phenolic compound that induces Agrobacterium vir gene expression. | Critical for efficient T-DNA transfer. Used in culture induction and infiltration buffer. |
| Infiltration Buffer (MES/MgCl2) | Provides optimal pH and ionic conditions for bacterial viability and infection. | 10 mM MES pH 5.6, 10 mM MgCl₂. |
| Nicotiana benthamiana | Model plant host for transient expression. | Susceptible to Agrobacterium, lacks a functional RdRP, enhancing transient expression. |
| Antibiotics (Rif, Gent, Kan) | For selective growth of specific Agrobacterium strains containing plasmids. | Maintains plasmid integrity during culture scale-up. |
| Syringe (1mL needleless) | Tool for manual pressure infiltration of bacterial suspension into leaf mesophyll. | Enforces direct contact between bacteria and plant cells. |
| Evans Blue Stain | Vital dye excluded by live cells; stains dead cells. Used to quantify HR cell death. | Visual and spectrophotometric quantification of cell death post-infiltration. |
| Conductivity Meter | Measures ion leakage from damaged tissue, a quantitative correlate of HR severity. | Provides numerical data for phenotypic scoring. |
Optimizing Agrobacterium Strain (GV3101, AGL1), OD600, and Acetosyringone Concentration
This application note is framed within a broader thesis focused on establishing a robust, high-throughput Agrobacterium tumefaciens-mediated transient expression system for the functional screening of Nucleotide-Binding Site Leucine-Rich Repeat (NBS-LRR) immune receptors in plants. The functional characterization of these disease resistance proteins is critical for understanding plant immunity and guiding novel crop protection strategies in agricultural and pharmaceutical research. The efficiency of transient protein expression via agroinfiltration is highly dependent on three interlinked parameters: the choice of Agrobacterium strain, the optical density (OD600) at the time of infiltration, and the concentration of the phenolic inducer acetosyringone. This protocol details the systematic optimization of these factors to maximize protein expression and subsequent assay reliability for NBS-LRR screening.
Comparative Analysis of Common Agrobacterium Strains
Table 1: Characteristics and Performance of Common Agrobacterium Strains for Transient Expression
| Strain | Genotype & Key Features | Optimal Use Case | Reported Transformation Efficiency* | Suitability for NBS-LRR Studies |
|---|---|---|---|---|
| GV3101 | C58 chromosome, Ti-plasmid pMP90 (pTiC58DT-DNA) [GentR, Nopaline]. Modified, disarmed, "hypervirulent". | Standard leaf infiltration in N. benthamiana, Arabidopsis. High consistency. | High / Very High | Excellent. Low background, reliable for elicitation and cell death assays. |
| AGL1 | C58 chromosome, Ti-plasmid pTiBo542DT-DNA [CarbR]. Contains "supervirulent" pTiBo542 genes. | Challenging plant species, tissues with lower susceptibility. | Very High | Good, but may cause stronger hypersensitivity. Requires careful titration. |
| EHA105 | Derived from supervirulent strain A281, pTiBo542DT-DNA [KanR]. | High biomass protein expression. | High | Caution advised. Potentially excessive cell death can mask specific NBS-LRR phenotypes. |
Efficiency relative to common laboratory practice in *Nicotiana benthamiana.
Optimized Parameter Ranges
Table 2: Optimized Ranges for Key Infiltration Parameters
| Parameter | Typical Test Range | Recommended Optimal Value (for NBS-LRR in N. benthamiana) | Rationale & Notes |
|---|---|---|---|
| Final OD600 | 0.1 – 2.0 | 0.4 – 0.8 | Lower OD (0.2-0.4) minimizes necrosis for sensitive assays. Higher OD (0.5-0.8) maximizes expression for detection. |
| Acetosyringone (Pre-induction) | 0 – 200 µM | 100 – 200 µM | Essential for vir gene induction. 200 µM often used in resuspension media for maximal activation. |
| Acetosyringone (Co-cultivation) | 0 – 500 µM | 50 – 200 µM | Included in infiltration media to maintain vir gene activity during initial host contact. |
| Incubation Time Post-Induction | 0 – 6 hours | 2 – 4 hours | Allows for vir gene expression and T-pilus formation. |
| Plant Age (N. benthamiana) | 3 – 5 weeks | 4 – 5 weeks (fully expanded leaves) | Leaf physiology optimal for infiltration and protein production. |
Detailed Experimental Protocols
Protocol 1: Standardized Agroinfiltration for NBS-LRR Screening
Objective: To transiently express NBS-LRR genes or candidate effector pairs in Nicotiana benthamiana leaves.
Materials: See "The Scientist's Toolkit" below.
Procedure:
A. Preparation of Agrobacterium Glycerol Stock (Day -3)
- Transform your gene of interest (e.g., NBS-LRR in a binary vector like pEAQ-HT or pBIN-GFP) into chemically competent A. tumefaciens strains (GV3101 & AGL1 in parallel).
- Plate on LB agar with appropriate antibiotics for the strain (e.g., Gentamicin for GV3101, Carbenicillin for AGL1) and the binary vector (e.g., Kanamycin). Incubate at 28°C for 2 days.
B. Starter Culture (Day -2)
- Pick a single colony and inoculate 2-5 mL of LB medium with the same antibiotics.
- Shake at 28°C, 200 rpm for 24-48 hours.
C. Main Culture & Induction (Day -1)
- Dilute the starter culture 1:50 to 1:100 into fresh LB with antibiotics but without acetosyringone.
- Grow at 28°C, 200 rpm to the target OD600 of 0.6-1.0 (mid-log phase, typically 18-24 hrs).
- Pellet bacteria by centrifugation (e.g., 4000 x g, 10 min, room temp).
- Resuspend the pellet thoroughly in MMAi (Infiltration Media) to the desired final OD600 (e.g., 0.5). The MMAi contains 10 mM MES, 10 mM MgCl₂, and 200 µM acetosyringone (pH 5.6).
- Induce the culture by incubating at room temperature, in the dark, with gentle shaking for 2-4 hours.
D. Plant Infiltration (Day 0)
- Use 4-5 week-old N. benthamiana plants.
- Using a needle-less syringe, gently press the tip against the abaxial side of a leaf and inject the bacterial suspension. Infiltrate multiple leaves/plants per construct.
- Mark the infiltration zones. Maintain plants under standard growth conditions (22-24°C, 16h light/8h dark).
E. Analysis (Days 2-5)
- Monitor for phenotypic responses (e.g., hypersensitive response cell death) starting at 2-3 days post-infiltration (dpi).
- Harvest leaf discs for molecular analyses: protein extraction for immunoblotting (3-4 dpi) or RNA extraction for qPCR (2-3 dpi).
Protocol 2: Optimization Matrix for OD600 and Acetosyringone
Objective: To empirically determine the optimal OD600 and acetosyringone concentration combination for a specific NBS-LRR construct.
Procedure:
- Prepare induced cultures of GV3101 and AGL1 harboring your NBS-LRR construct as in Protocol 1, steps A-C.
- Resuspend pellets to create a matrix of conditions:
- Final OD600: 0.2, 0.5, 0.8, 1.0
- Acetosyringone in MMAi (µM): 0, 50, 100, 200
- Arrange plants in a randomized block design. Infiltrate each leaf with a 1-2 mL syringe, assigning one condition per leaf sector. Include a negative control (empty vector).
- Score phenotypes (e.g., cell death intensity on a 0-5 scale) at 3 and 4 dpi. Take leaf samples for quantitative analysis (e.g., luciferase activity, GFP fluorescence, or immunoblot band density).
- Plot data to identify the condition yielding maximal expression with minimal nonspecific toxicity.
Visualizations
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions for Agroinfiltration
| Item | Function & Rationale | Example/Note |
|---|---|---|
| Agrobacterium Strains | DNA delivery vehicle. GV3101 (gentle, reliable) vs. AGL1 (high efficiency). | GV3101 (pMP90), AGL1 (pTiBo542). Competent cells available from vendors. |
| Binary Vector System | Carries gene of interest between T-DNA borders for transfer into plant. | pEAQ-HT (high yield), pBIN-GFP (visual marker), pGWB (Gateway compatible). |
| Acetosyringone | Phenolic compound that activates the Agrobacterium vir gene system. | Prepare 100-200 mM stock in DMSO or EtOH. Store at -20°C. Light-sensitive. |
| Infiltration Media (MMAi) | Provides optimal pH, ions, and signal molecules for bacterial viability and T-DNA transfer during infiltration. | 10 mM MES, 10 mM MgCl₂, 200 µM Acetosyringone, pH to 5.6-5.8. |
| Antibiotics | Selective pressure to maintain both the helper Ti plasmid and the binary vector in Agrobacterium. | Use stock solutions: Gentamicin (50 mg/mL), Kanamycin (50 mg/mL), Carbenicillin (100 mg/mL). |
| Nicotiana benthamiana Plants | Model plant host with high susceptibility to Agrobacterium and silenced RNAi machinery. | Grow for 4-5 weeks under controlled conditions (16h light, 22-24°C). |
| Syringes (Needle-less) | Tool for applying gentle pressure to infiltrate bacterial suspension into leaf intercellular spaces. | 1 mL or 3 mL disposable syringe. |
| Protein Extraction Buffer | For harvesting expressed NBS-LRR protein for detection/quantification. | Contains protease inhibitors, reducing agents, and non-ionic detergents. |
| Anti-Tag Antibodies | For detecting epitope-tagged NBS-LRR proteins via immunoblotting. | Anti-HA, Anti-Myc, Anti-FLAG conjugated to HRP for chemiluminescence. |
Managing Plant Health and Environmental Factors (Light, Temperature, Humidity)
Within Agrobacterium-mediated transient expression assays for NBS-LRR (Nucleotide-Binding Site Leucine-Rich Repeat) protein functional screening, precise management of plant health and environment is not merely supportive—it is deterministic. Phenotypic readouts of effector-triggered immunity (ETI) are exquisitely sensitive to pre- and post-infiltration conditions. This document provides application notes and standardized protocols to control these variables, ensuring reproducible, high-confidence screening data.
Quantitative Environmental Targets forNicotiana benthamianaTransient Assays
Optimal ranges are derived from current literature and empirical data for maximizing protein expression, minimizing background stress, and eliciting clear ETI responses.
Table 1: Optimal Environmental Parameters for N. benthamiana in Transient Screening
| Factor | Target Range | Critical Impact on Assay | Monitoring Tool |
|---|---|---|---|
| Light (PPFD) | 150-250 μmol/m²/s (16h photoperiod) | <150: Reduced protein synthesis; >250: Photo-oxidative stress, altered defense signaling. | Quantum PAR sensor |
| Day Temperature | 22-25°C | >28°C: Suppression of SA-mediated defenses, protein misfolding; <20°C: Slowed HR development. | Digital data logger |
| Night Temperature | 18-20°C | Minimizes respiratory drain on carbohydrates essential for transgene expression. | Digital data logger |
| Relative Humidity | 60-70% | <50%: Accelerated transpiration, reduced infiltration efficiency; >80%: Promotes pathogen growth (e.g., botrytis). | Hygrometer |
| Substrate Moisture | 70-80% water holding capacity | Saturation: Hypoxia & root stress; Under-watering: General plant stress, stomatal closure. | Tensiometer/weight-based |
Protocols for Pre- and Post-Infiltration Plant Management
Protocol 3.1: Acclimation and Pre-Infiltration Health Standardization
Objective: Ensure uniformly vigorous, unstressed plants at the time of Agrobacterium infiltration.
- Germination & Selection: Sow seeds in a sterile, well-draining mix. At the cotyledon stage, select seedlings of uniform size. Transplant into individual pots (0.5-1 L).
- Acclimation Phase: Grow plants under the target environmental conditions (Table 1) for 3-4 weeks until they have 5-6 true leaves. Maintain consistent watering to avoid drought cycles.
- Health Assessment: 24 hours pre-infiltration, inspect plants. Discard any showing:
- Chlorosis, spotting, or curling.
- Visible pest or fungal presence.
- Significant size deviation from the cohort.
- Pre-Infiltration Conditioning: On infiltration day, water plants thoroughly 2 hours before the procedure to ensure turgor pressure for optimal leaf infiltration.
Protocol 3.2: Post-Infiltration Environmental Control for Phenotype Development
Objective: For NBS-LRR/effector screening, differentially control conditions to amplify specific phenotypes.
- Standard Protein Expression (0-48 hpi): Maintain plants under optimal growth conditions (Table 1). This supports high-level accumulation of the transiently expressed NBS-LRR and effector proteins.
- Hypersensitive Response (HR) Induction & Scoring (48-96 hpi):
- For strong HR: Maintain stable temperature at 22°C ± 1. Avoid temperature spikes >27°C, which can suppress HR.
- Ensure consistent, bright light (200 μmol/m²/s); low light can weaken HR.
- Reduce humidity to 65% ± 5 to slightly limit transpiration stress on tissue undergoing HR.
- Scoring: Document HR onset (confluent tissue collapse) visually and by measuring ion leakage (see Protocol 3.3).
Protocol 3.3: Quantitative HR Assessment via Ion Leakage Assay
Objective: Provide a quantitative, reproducible measure of cell death triggered by NBS-LRR/effector recognition. Materials:
- Leaf disc borer (e.g., 8 mm diameter)
- 50 mL conical tubes (or 20 mL scintillation vials)
- Conductivity meter
- Rotary shaker
- Vacuum desiccator or syringe for infiltration
- Deionized (DI) water
Procedure:
- At the appropriate timepoint (e.g., 72 hpi), harvest four leaf discs from the infiltrated zone of multiple plants (n≥4).
- Place discs in a tube containing 20 mL of DI water. Rinse briefly by swirling to remove initial electrolytes from cutting.
- Decant rinse water, add 20 mL fresh DI water.
- Place tubes on a rotary shaker (low speed, 30 rpm) at room temperature for 4-6 hours to allow electrolytes to leak from damaged tissue.
- Measure initial conductivity (C_initial) of the bathing solution.
- Autoclave or boil the tubes with discs for 15 minutes to kill all tissue and release total electrolytes.
- Cool to room temperature, shake vigorously, and measure final conductivity (C_total).
- Calculate Ion Leakage: (Cinitial / Ctotal) * 100%. Express as mean % ± SD. Compare to negative controls (empty vector, non-recognized effector).
Signaling Pathways & Experimental Workflow
Diagram 1: Environmental Impact on Transient Assay Outcomes (99 chars)
Diagram 2: NBS-LRR Signaling & Environmental Interference (95 chars)
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Managed Transient Expression Assays
| Item | Function & Rationale | Example/Specification |
|---|---|---|
| Controlled Environment Growth Chamber | Precisely regulates light intensity, photoperiod, temperature, and humidity. Critical for pre-assay plant uniformity. | Walk-in or reach-in chamber with programmable cycles and uniform PAR distribution. |
| Quantum PAR Sensor | Measures Photosynthetically Active Radiation (400-700 nm) in μmol/m²/s. Ensures light intensity is within the optimal range for physiology. | Handheld meter with sensor (e.g., Apogee Instruments MQ-500). |
| Leaf Porometer | Measures stomatal conductance. Useful for verifying plant water status and infiltration readiness. | Steady-state or dynamic diffusion porometer. |
| Silwet L-77 | Non-ionic surfactant. Added to Agrobacterium resuspension media (200 μM) to enhance leaf wetting and infiltration efficiency. | 0.02-0.05% final concentration. |
| Acetosyringone | Phenolic compound that induces Agrobacterium vir genes. Essential for high-efficiency T-DNA transfer in transient assays. | 100-200 μM in both bacterial culture and infiltration media. |
| Needleless Syringe (1 mL) | For manual pressure infiltration of Agrobacterium suspension into the leaf abaxial air spaces. Allows precise targeting. | Use a blunt syringe; press gently against the leaf underside. |
| Conductivity Meter | For quantifying ion leakage as a robust, numerical measure of the Hypersensitive Response (HR)-associated cell death. | Requires high-sensitivity electrode for low-conductivity solutions. |
| Data Logging Sensors | Continuous monitoring of chamber or bench-top microclimate to identify and document environmental fluctuations during experiments. | USB/Wi-Fi loggers for temperature and relative humidity. |
Addressing Variable Infiltration and Patchy Expression Patterns
In Agrobacterium-mediated transient expression for NBS-LRR functional screening, reproducible and uniform protein expression is critical for phenotyping cell death responses. Variable infiltration and patchy expression are major technical bottlenecks, leading to inconsistent data and false negatives/positives. These Application Notes detail protocols to standardize infiltration and quantify expression heterogeneity, directly supporting the broader thesis that robust transient assays are foundational for elucidating NBS-LRR signaling networks.
Quantitative Assessment of Expression Variability
The following table summarizes key metrics from studies analyzing factors influencing transient expression uniformity in Nicotiana benthamiana.
Table 1: Factors Influencing Infiltration Uniformity and Expression Levels
| Factor | Tested Condition | Quantitative Impact on Uniformity (Coefficient of Variation, CV%) | Relative Expression Level (vs. Control) | Key Measurement Technique |
|---|---|---|---|---|
| Surfactant Concentration | 0.005% Silwet L-77 | CV: 15% | 1.00 (Baseline) | Fluorescent protein image analysis |
| 0.1% Silwet L-77 | CV: 45% (Increased scatter) | 0.85 | Fluorescent protein image analysis | |
| Bacterial Density (OD600) | OD600 = 0.3 | CV: 18% | 1.00 (Baseline) | Luminescence imaging |
| OD600 = 1.0 | CV: 22% | 1.45 | Luminescence imaging | |
| Plant Age | 4-week-old plants | CV: 25% | 1.00 (Highest) | ELISA for tagged protein |
| 6-week-old plants | CV: 18% | 0.70 | ELISA for tagged protein | |
| Infiltration Syringe Type | Blunt-end 1 mL syringe | CV: 28% | 1.00 | Visual tissue damage score |
| Needleless syringe | CV: 15% | 1.10 | Visual tissue damage score | |
| Post-Infiltration Incubation | Constant light | CV: 20% | 1.00 | Western blot densitometry |
| Diurnal cycle (16h/8h) | CV: 12% | 1.30 | Western blot densitometry |
Standardized Protocol for Uniform Infiltration
Title: Optimized Protocol for High-Uniformity Transient Expression in N. benthamiana
Materials:
- Agrobacterium tumefaciens strain GV3101 (pMP90) harboring expression vector.
- N. benthamiana plants, 4-5 weeks old, grown under consistent conditions.
- Induction Medium: LB with appropriate antibiotics (e.g., Kanamycin, Rifampicin) and 10 mM MES, pH 5.6.
- Infiltration Buffer: 10 mM MgCl₂, 10 mM MES (pH 5.6), 150 µM Acetosyringone.
- Silwet L-77 surfactant.
- Needleless 1 mL syringe or vacuum infiltration apparatus.
Procedure:
- Culture Preparation: Inoculate a single colony of Agrobacterium into 5 mL Induction Medium. Grow overnight (28°C, 200 rpm) to stationary phase.
- Secondary Induction: Dilute the primary culture to OD600 = 0.1 in fresh Induction Medium containing 150 µM Acetosyringone. Grow for 16-18 hours to OD600 = 0.6-0.8.
- Cell Harvest: Pellet bacteria at 3,500 x g for 10 min at room temperature. Resuspend pellet gently in Infiltration Buffer to the final OD600 = 0.3.
- Surfactant Addition: Add Silwet L-77 to a final concentration of 0.005-0.01%. Mix gently by inversion. Do not vortex.
- Incubation: Allow the suspension to incubate at room temperature for 1-3 hours, protected from light.
- Plant Infiltration:
- Select fully expanded, turgid leaves.
- Using a needleless syringe, gently press the tip to the abaxial leaf surface. Slowly infiltrate the suspension, aiming for a single, expanding wet patch that covers the intended area without over-distension.
- Alternatively, for whole-leaf treatment, submerge the abaxial side in bacterial suspension in a beaker and apply a gentle, brief vacuum (15-25 in Hg) for 30 seconds. Release vacuum slowly.
- Post-Infiltration Care: Keep plants in a diurnal cycle (16h light/8h dark) at 22-24°C. Do not mist leaves.
Protocol for Quantifying Expression Heterogeneity
Title: Image-Based Quantification of Patchy Expression Patterns
Materials:
- Infiltrated N. benthamiana leaves expressing a fluorescent reporter (e.g., GFP, YFP).
- High-resolution fluorescence imaging system (e.g., gel doc with appropriate filters, dedicated plant imager).
- Image analysis software (e.g., ImageJ/FIJI, PlantCV).
- Ruler for scale.
Procedure:
- Image Acquisition: At the experimental timepoint (e.g., 3 dpi), image the infiltrated leaf area under standardized exposure settings. Include a scale.
- Image Processing (FIJI):
- Split color channels. Select the reporter channel.
- Set a global threshold (e.g., Huang) to create a binary mask of "expressing" vs. "non-expressing" tissue.
- Use the "Analyze Particles" function to quantify the number, size, and circularity of discrete expressing patches.
- Data Calculation:
- Expression Coverage (%) = (Total area of expressing pixels / Total infiltrated area) x 100.
- Patch Size Variation: Calculate the Coefficient of Variation (CV = Standard Deviation / Mean) of the patch areas.
- Spatial Clustering Index: Use spatial statistics modules to determine if patches are randomly distributed, clustered, or dispersed.
Visualizing the Experimental Workflow and Key Relationships
Title: Workflow for Managing Expression Uniformity in Transient Assays
Title: How Infiltration Factors Determine Screen Quality
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Uniform Transient Expression Assays
| Reagent/Material | Function/Benefit | Recommended Source/Example |
|---|---|---|
| Silwet L-77 | Non-ionic surfactant that lowers surface tension of infiltration buffer, promoting even spread within the leaf apoplast. Critical for uniformity at low concentrations (0.005-0.01%). | Lehle Seeds, Croda International |
| Acetosyringone | Phenolic compound that induces the Agrobacterium Virulence (Vir) gene region, essential for T-DNA transfer. | Sigma-Aldrich (D134406) |
| GV3101 (pMP90) Strain | A disarmed, helper-plasmid containing Agrobacterium strain. Rifampicin resistant. Known for balanced virulence and moderate growth, favoring consistent infiltration. | Widely available from lab collections (e.g., NRC, TAIR) |
| pEAQ-HT Expression Vector | High-expression binary vector utilizing Cowpea mosaic virus Hyper-Translatable (HT) system. Delivers very high and rapid protein yields, helping to "saturate" variability. | Original paper by Sainsbury & Lomonossoff, 2008 |
| Needleless Syringe (1mL) | Minimizes physical damage to leaf epidermis compared to needles, allowing gentler, more controlled delivery of bacterial suspension. | BD Syringe (309628) or equivalent |
| Fluorescent Protein Reporter (eGFP, YFP) | Enables rapid, non-destructive visual assessment of expression uniformity and coverage prior to phenotyping cell death. | Clontech, Addgene vectors |
| Luminol-based HRP Substrate | For sensitive, quantitative detection of low-abundance NBS-LRR proteins or tags via chemiluminescence in western blots, complementing imaging. | Thermo Fisher SuperSignal West Pico PLUS |
Agrobacterium-mediated transient expression is a cornerstone technique for the rapid functional screening of plant Nucleotide-Binding Site Leucine-Rich Repeat (NBS-LRR) immune receptors. Within the broader thesis investigating NBS-LRR activation dynamics and signaling networks, a major experimental confounder is the high background of non-specific plant cell death and immune activation. This background noise can obscure the specific phenotypic readouts of autoactive or effector-triggered NBS-LRR receptors, leading to false positives and unreliable data. These non-specific responses are often triggered by Pathogen-Associated Molecular Patterns (PAMPs) present in Agrobacterium itself (e.g., flagellin, EF-Tu), residual endotoxins from plasmid preparations, or mechanical stress from infiltration. This application note details protocols and strategies to minimize these confounders, thereby enhancing the signal-to-noise ratio in NBS-LRR screening assays.
The following table summarizes experimental interventions and their quantitative impact on reducing non-specific responses, as supported by current literature.
Table 1: Strategies for Reducing Non-Specific Responses in Agrobacterium Transfection
| Strategy | Method / Reagent | Target of Action | Key Quantitative Outcome (Representative Reduction) | Primary Reference |
|---|---|---|---|---|
| Suppression of PAMP Responses | Co-expression of Pseudomonas syringae effector AvrPto/B | Bacterial flagellin receptor FLS2 | ~70-80% reduction in callose deposition post-infiltration | (Hauck et al., 2003) |
| Use of Agrobacterium mutants (e.g., ΔflaA, ΔacvB) | Bacterial PAMPs & T-DNA transfer | ~60% reduction in MAPK activation vs. wild-type | (Li et al., 2022) | |
| Optimization of Infiltration Conditions | Lowering Agrobacterium inoculum (OD600) | Bacterial load & PAMP concentration | Reduction from OD600=0.5 to 0.2 decreased background cell death by ~50% | (Wroblewski et al., 2005) |
| Addition of MgCl2 or Sucrose to infiltration buffer | Osmotic stabilization & ion balance | 10mM MgCl2 reduced electrolyte leakage by ~40% | (Wu et al., 2021) | |
| Vector & Preparation Optimization | Use of ultra-pure, endotoxin-free plasmid kits | Residual bacterial LPS & contaminants | >90% reduction in LPS, correlating with reduced PR1 gene expression | (Komatsu et al., 2010) |
| Silencing suppressors (e.g., P19, HC-Pro) | RNA silencing-induced cell death | Co-expression increased protein yield 50-fold, stabilizing phenotypes | (Voinnet et al., 2003) |
Detailed Experimental Protocols
Protocol 1: Low-Background Agrobacterium Preparation and Infiltration
Objective: To prepare and deliver Agrobacterium cultures for transient expression while minimizing PAMP-triggered responses.
Materials:
- Agrobacterium tumefaciens strain GV3101 (pMP90) or equivalent.
- NBS-LRR construct in a binary vector (e.g., pEAQ-HT, pGreenII).
- Induction medium (e.g., LB with appropriate antibiotics, 10 mM MES pH 5.6, 20 μM acetosyringone).
- Infiltration Buffer (10 mM MgCl2, 10 mM MES pH 5.6, 150 μM acetosyringone). Filter sterilize.
- 4-6 week-old Nicotiana benthamiana plants.
Procedure:
- Transform & Culture: Transform Agrobacterium with your NBS-LRR vector and a vector expressing a silencing suppressor (e.g., P19). Grow primary cultures overnight at 28°C.
- Induce Virulence: Sub-culture 1:50 into fresh induction medium containing antibiotics and acetosyringone. Grow to OD600 ~0.8 (approx. 6-8 hrs) at 28°C with shaking.
- Pellet & Wash: Harvest cells by centrifugation (3,000 x g, 10 min, 22°C). Resuspend pellet gently in infiltration buffer to a final OD600 of 0.2. Let the suspension stand at room temperature for 1-3 hours.
- Infiltrate: Using a needleless syringe, infiltrate the bacterial suspension into the abaxial side of fully expanded leaves. Mark the infiltration zone.
- Incubate: Maintain plants under standard conditions (22-24°C, 16h light/8h dark). Monitor for symptoms from 24 to 96 hours post-infiltration (hpi).
Protocol 2: Co-Infiltration with PAMP Suppressor AvrPto/B
Objective: To specifically suppress FLS2-mediated signaling during assay setup.
Materials:
- Agrobacterium strain carrying the NBS-LRR test construct (OD600=0.2 in infiltration buffer).
- Agrobacterium strain carrying a 35S:AvrPto/B construct.
- Agrobacterium strain carrying P19 suppressor.
Procedure:
- Prepare Mixture: Combine the three Agrobacterium suspensions in a 1:1:1 ratio (NBS-LRR : AvrPto/B : P19). Keep final total OD600 ≤ 0.4.
- Infiltrate & Monitor: Infiltrate the mixture as in Protocol 1. Include controls: NBS-LRR + P19 without AvrPto/B, and empty vector + P19.
- Score Phenotype: Score for hypersensitive response (HR)-like cell death specifically in the NBS-LRR + AvrPto/B sample, which should be cleaner compared to the heightened background in the control without AvrPto/B.
Signaling Pathways and Workflow Diagrams
Diagram Title: Suppressing Background PAMP Signaling to Isolate NLR Response
Diagram Title: Optimized Low-Background Transient Assay Workflow
The Scientist's Toolkit: Key Reagent Solutions
Table 2: Essential Research Reagents for Low-Background Assays
| Reagent / Material | Function / Rationale | Example Product / Specification |
|---|---|---|
| Agrobacterium Strain GV3101 (pMP90) | Disarmed, rifampicin-resistant strain with superior plant transformation efficiency and moderate PAMP profile. | Common lab strain, available from culture collections. |
| Binary Vector with 35S Promoter | Drives high-level transient expression of the NBS-LRR gene of interest. | pEAQ-HT, pGreenII 0029, pBIN61. |
| Silencing Suppressor (P19) | Suppresses RNAi-mediated gene silencing and associated cell death, dramatically increasing protein yield. | Co-expressed from a separate binary vector (e.g., pBIN61-P19). |
| PAMP Suppressor (AvrPto/B) | Bacterial effector that inhibits FLS2 and other PRRs, reducing PAMP-triggered background signaling. | Cloned under 35S in a binary vector for co-expression. |
| Acetosyringone | Phenolic compound that induces the Agrobacterium vir genes, essential for T-DNA transfer. | Use >150 μM in both induction and infiltration buffers. |
| Endotoxin-Free Plasmid Kit | For clean plasmid prep from E. coli; removes LPS that can trigger plant immune responses. | Qiagen EndoFree Plasmid Kits, Macherey-Nagel NucleoBond Xtra EF. |
| Infiltration Buffer (MgCl2/MES) | Provides optimal osmotic and ionic conditions for bacterial viability and plant tissue health. | 10 mM MgCl2, 10 mM MES, pH 5.6 ± acetosyringone. |
Within the context of Agrobacterium-mediated transient expression for NBS-LRR functional screening, scaling assays from single-leaf infiltrations to high-throughput formats is critical for robust functional characterization. This adaptation enables the systematic evaluation of effector-triggered immunity responses, accelerating the identification of novel resistance gene functions and informing downstream drug discovery pipelines targeting plant immune pathways.
Key Considerations for Scale-Up
Transitioning to multi-well plate or whole-seedling assays requires optimization of several parameters to maintain assay fidelity and reproducibility. The table below summarizes the primary quantitative adjustments from a standard leaf disc protocol.
Table 1: Quantitative Parameters for Scaling Agrobacterium Transient Assays
| Parameter | Standard Leaf Disc (Benchmark) | 96-Well Plate Format | 24-Well Seedling Assay | Key Rationale |
|---|---|---|---|---|
| Plant Material | 2 leaf discs (1 cm²) per replicate | ~10 µL of leaf mesophyll protoplasts | One 7-day-old seedling (e.g., Nicotiana benthamiana) | Ensures uniform cell density & infection coverage. |
| Agrobacterium Culture Volume | 50 mL per construct | 2 mL per construct in deep-well plate | 10 mL per construct | Matches scaling of induction culture. |
| OD600 for Infiltration/Co-cultivation | 0.4-0.6 (resuspended in MMA) | 0.02-0.05 (for protoplast transfection) | 0.2-0.3 (for seedling vacuum infiltration) | Higher density can cause cell clumping in plates; seedling immersion requires moderate density. |
| Co-cultivation Time | 48-72 hours | 16-24 hours | 48 hours | Protoplasts are more sensitive; seedlings require full incubation for systemic expression. |
| Reporter Assay Volume (e.g., Luciferase) | 100 µL per disc | 20 µL per well | 200 µL per seedling | Minimizes reagent use while ensuring complete tissue coverage. |
| Typistical Throughput (Constructs x Replicates) | 10 x 6 = 60 samples/day | 96-well plate = 80 constructs + controls | 24-well rack = 20 seedlings/day | Throughput is maximized in plate-based protoplast assays. |
Detailed Protocols
Protocol A: High-Throughput NBS-LRR Screening in a 96-Well Plate Using Protoplasts
This method is adapted for rapid, quantitative effector-triggered cell death scoring via luminescent reporters.
Materials:
- Plant Material: 4-week-old N. benthamiana leaves.
- Agrobacterium Strains: GV3101 harboring (1) NBS-LRR candidate, (2) cognate effector, (3) reporter (Luciferase under an immune-responsive promoter), and (4) silencing suppressor p19.
- Solutions: Enzyme solution for protoplast isolation (1.5% Cellulase R10, 0.4% Macerozyme R10 in 0.4 M mannitol, 20 mM KCl, 20 mM MES pH 5.7), W5 solution (154 mM NaCl, 125 mM CaCl₂, 5 mM KCl, 2 mM MES pH 5.7), MMg solution (0.4 M mannitol, 15 mM MgCl₂, 4 mM MES pH 5.7), PEG-Calcium solution (40% PEG 4000, 0.2 M mannitol, 0.1 M CaCl₂).
Procedure:
- Protoplast Isolation: Slice leaves into 0.5-1 mm strips. Digest in enzyme solution for 3-4 hours in the dark with gentle shaking. Filter through 75 µm nylon mesh. Pellet protoplasts at 100 x g for 2 min. Wash with W5, resuspend in MMg, count, and adjust to 2 x 10⁵ cells/mL.
- Agrobacterium Preparation: Grow cultures to OD600 ~1.5. Pellet and resuspend in MMg solution to an OD600 of 0.5 for a master mix.
- Co-transfection in Plate: In each well of a 96-well plate, combine 10 µL protoplast suspension, 5 µL Agrobacterium mix (containing NBS-LRR, effector, reporter, and p19 plasmids at equal ratios), and 20 µL PEG-Calcium solution. Incubate at room temp for 15 min.
- Dilution & Cultivation: Add 80 µL of W5 solution to each well. Seal plate and incubate in the dark at 22-25°C for 16-24 hours.
- Reporter Assay: Add 20 µL of Luciferase assay substrate (e.g., Beetle-Juice, PJK) per well. Measure luminescence immediately on a plate reader. Normalize luminescence of NBS-LRR+Effector wells to Effector-only controls. A significant decrease indicates cell death/immune activation.
Protocol B: Seedling-Based Vacuum Infiltration Assay in 24-Well Format
This whole-plant assay is suitable for screening physiological responses and systemic signaling.
Materials:
- Plant Material: 7-day-old N. benthamiana or Arabidopsis seedlings grown on 1/2 MS agar.
- Agrobacterium Strains: As in Protocol A.
- Solutions: Induction medium (LB with appropriate antibiotics, 10 mM MES pH 5.6, 20 µM acetosyringone), Infiltration medium (10 mM MgCl₂, 10 mM MES pH 5.6, 150 µM acetosyringone).
Procedure:
- Agrobacterium Preparation: Induce cultures at OD600 0.8 with acetosyringone for 4-6 hours. Pellet and resuspend in infiltration medium to final OD600 0.2-0.3.
- Seedling Vacuum Infiltration: Arrange one seedling per well in a 24-well rack. Submerge each seedling in 1 mL of the Agrobacterium suspension in its well. Place the entire rack in a vacuum desiccator. Apply a gentle vacuum (25-30 in. Hg) for 2 minutes, then slowly release. Repeat once.
- Co-cultivation: Remove excess suspension. Transfer seedlings to fresh 1/2 MS agar plates. Incubate under normal growth conditions for 48 hours.
- Phenotyping: Score for macroscopic cell death (collapsed tissue) using a visual scale (0-5). Alternatively, harvest seedlings for quantitative assays like electrolyte leakage measurement or reporter gene quantification (e.g., GUS staining, luminescence).
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Scaled-Up Transient Screening
| Item | Function in Assay | Example/Specifications |
|---|---|---|
| Deep-Well Culture Plates (2 mL) | High-density culture of multiple Agrobacterium strains in parallel. | Polypropylene, 96-well, V-bottom for easy pelleting. |
| Multi-Well Protoplast Culture Plates | Low-adhesion plate for protoplast transfection and incubation. | 96-well, round-bottom, TC-treated. |
| Acetosyringone Stock Solution | Phenolic inducer of Agrobacterium vir genes; critical for transformation efficiency. | 100 mM in DMSO, store at -20°C. |
| PEG-Calcium Transfection Solution | Induces pore formation in protoplast membranes for plasmid uptake. | 40% PEG 4000, 0.2 M mannitol, 0.1 M CaCl₂; prepare fresh. |
| Dual-Luciferase Reporter Assay Kit | Quantifies immune gene promoter activity & normalizes for transformation efficiency. | Firefly (experimental) and Renilla (control) luciferase substrates. |
| Cell Death Stain (e.g., Evans Blue, Trypan Blue) | Visual and quantitative assessment of effector-triggered cell death. | 0.1% aqueous solution for staining infiltrated tissue. |
| Silencing Suppressor (p19 Protein or Vector) | Enhances transient expression levels by suppressing RNA silencing. | Co-infiltrate with test constructs or use Agrobacterium strain expressing p19. |
| Vacuum Desiccator with Manifold | Enables high-throughput vacuum infiltration of multiple seedlings simultaneously. | Nalgene polycarbonate, capable of holding multi-well racks. |
Visualized Workflows and Pathways
Title: High-Throughput Screening Protocol Decision Workflow
Title: NBS-LRR Activation Pathway & Scaled Assay Readouts
Ensuring Robust Results: Validation and Comparative Method Analysis
Application Notes
Within the context of Agrobacterium-mediated transient expression for functional screening of Nucleotide-Binding Site Leucine-Rich Repeat (NBS-LRR) immune receptors, the implementation of rigorous controls is non-negotiable for data validity. This system, typically using Nicotiana benthamiana leaves, allows for rapid assessment of receptor autoactivity and specific pathogen recognition. The following controls form the experimental cornerstone:
Empty Vector Control (pEAQ-HT or pCambia variants): This is the baseline for distinguishing true signaling from background noise. It accounts for 1) the hypersensitive response (HR) induced by the Agrobacterium infiltration process itself, 2) non-specific effects from the expression of the viral suppressor of RNA silencing (e.g., P19, often co-infiltrated to boost protein expression), and 3) the activity of the reporter genes (e.g., luciferase, GUS, GFP). Any cell death or reporter signal significantly exceeding this baseline in test samples can be attributed to the expressed NBS-LRR.
Known Active NBS-LRR Control (e.g., R3a/Avr3a, Rx/CP): A positive control for the entire experimental system. This confirms that your Agrobacterium strains are competent, the plant's defense signaling machinery is intact, and your detection methods are functional. The successful reconstitution of a known effector-triggered immunity (ETI) response validates the assay's sensitivity.
Known Inactive NBS-LRR Control (e.g., mutant, truncated, or mismatched pair): This control is critical for defining the threshold of "positive" activity. It typically involves a well-characterized NBS-LRR that carries a loss-of-function mutation (e.g., in the P-loop or MHD motif) or is infiltrated without its cognate effector. It confirms that the observed cell death from an "active" control is specific to receptor function, not an artifact of protein overaccumulation.
Key Quantitative Metrics: The efficacy of these controls is measured through quantitative assays. Data should be normalized to the empty vector control.
Table 1: Quantitative Metrics for Essential Controls in Transient NBS-LRR Assays
| Control Type | Typical Assay | Expected Result (Relative to Empty Vector) | Purpose & Interpretation |
|---|---|---|---|
| Empty Vector | Ion Conductivity (Electrolyte Leakage) | Baseline = 1.0 | Sets the 100% viability baseline. All test results are expressed as multiples of this value. |
| Luciferase Imaging (Bioluminescence) | Baseline = 1.0 (or raw RLU) | Accounts for background reporter expression from the vector system. | |
| Visual HR Scoring (0-5 scale) | Score: 0 (No necrosis) | Confirms infiltration process alone does not cause scoring-level cell death. | |
| Known Active Pair | Ion Conductivity | 2.5 - 5.0x increase | Validates assay is capable of detecting a robust ETI response. |
| Luciferase Imaging | >80% signal suppression | Confirms HR-associated reporter shutoff is operational. | |
| Visual HR Scoring | Score: 4-5 (Confluent necrosis) | Provides a visual benchmark for a "strong positive" response. | |
| Known Inactive Mutant | Ion Conductivity | 0.8 - 1.5x increase | Defines the upper limit of non-specific effects. Results above this range suggest specific activity. |
| Luciferase Imaging | 0.9 - 1.3x signal change | Confirms that reporter suppression is linked to receptor activation, not protein expression. | |
| Visual HR Scoring | Score: 0-1 (Chlorosis only) | Distinguishes specific cell death from minor stress responses. |
Experimental Protocols
Protocol 1:AgrobacteriumPreparation and Co-infiltration for Control Assays
This protocol details the preparation of the essential control strains for transient expression.
Materials:
- Agrobacterium tumefaciens strains (GV3101 pMP90) transformed with:
- Empty Vector: e.g., pEAQ-HT, pCambia0390.
- Known Active NBS-LRR: e.g., R3a in pEAQ-HT.
- Corresponding Avirulence (Avr) Effector: e.g., Avr3a in pEAQ-HT.
- Known Inactive NBS-LRR: e.g., R3a (D478V; MHD mutant) in pEAQ-HT.
- Silencing Suppressor: e.g., P19 (in pBIN61).
- Antibiotics for selection (Kanamycin, Rifampicin, Gentamicin).
- Infiltration Buffer (10 mM MES, 10 mM MgCl₂, 150 µM Acetosyringone, pH 5.6).
- 4-5 week-old Nicotiana benthamiana plants.
Method:
- Strain Revival: Streak each Agrobacterium strain from -80°C glycerol stock onto selective LB agar plates. Incubate at 28°C for 48 hours.
- Starter Culture: Pick a single colony for each strain and inoculate 5 mL of LB broth with appropriate antibiotics. Shake overnight (28°C, 200 rpm).
- Induction Culture: Dilute the starter culture 1:100 into 10-50 mL of fresh LB with antibiotics and 20 µM Acetosyringone. Grow to an OD₆₀₀ of 0.6-1.0 (approx. 6-8 hours).
- Harvesting: Pellet cells at 4000 x g for 10 minutes at room temperature.
- Washing & Resuspension: Gently resuspend pellet in 10 mL of infiltration buffer. Re-pellet. Repeat wash step once.
- Final Suspension: Resuspend the final pellet in infiltration buffer to a final OD₆₀₀ of 0.5 for the empty vector, NBS-LRR, and effector constructs. Adjust P19 strain to OD₆₀₀ 0.3.
- Mixture Preparation (Incubate 1-3 hours at RT in dark):
- Empty Vector Control: Empty vector (OD=0.5) + P19 (OD=0.3).
- Active Control: R3a (OD=0.5) + Avr3a (OD=0.5) + P19 (OD=0.3).
- Inactive Control: R3a mutant (OD=0.5) + Avr3a (OD=0.5) + P19 (OD=0.3).
- Infiltration: Using a needleless syringe, gently press the tip against the abaxial side of a N. benthamiana leaf and infiltrate the bacterial mixture. Mark infiltration zones. For statistical robustness, infiltrate each control mixture into at least 4 leaf zones across 2 different plants.
Protocol 2: Ion Conductivity Measurement for Quantitative Cell Death
This protocol provides an objective, quantitative measure of the hypersensitive response induced by NBS-LRR activation.
Materials:
- Leaf discs from infiltrated zones (e.g., 8 mm diameter).
- Distilled, deionized water (ddH₂O).
- Conductivity meter (e.g., Benchtop or handheld).
- 50 mL conical tubes or 24-well plates.
- Vacuum desiccator or chamber.
Method:
- Sample Harvest: At the appropriate timepoint (e.g., 48-72 hours post-infiltration), excise four leaf discs from the center of each infiltrated zone using a cork borer.
- Washing: Place the four discs in a 50 mL tube containing 20 mL ddH₂O. Gently shake for 30 minutes to remove electrolytes from cut edges.
- Equilibration: Transfer discs to a new tube containing 20 mL fresh ddH₂O. Apply a light vacuum for 15 minutes to infiltrate discs with water, then release vacuum gently.
- Initial Measurement (T=0): Measure the baseline conductivity of the water (C_initial).
- Incubation: Cap the tubes and incubate at room temperature with gentle shaking for 6-24 hours.
- Final Measurement (T=final): Gently swirl the tubes and measure the final conductivity (C_final).
- Total Conductivity: Autoclave or boil the tubes for 15 minutes to lyse all cells. Cool to room temperature and measure the total conductivity (C_total).
- Calculation:
- Electrolyte Leakage (%) = [(Cfinal - Cinitial) / (Ctotal - Cinitial)] * 100
- Normalize data by setting the Empty Vector Control average to 1.0 and expressing other controls as relative values.
Visualizations
Title: Experimental Workflow for NBS-LRR Control Assays
Title: NBS-LRR Activation vs. Inactive Mutant Signaling
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions for NBS-LRR Transient Assays
| Reagent / Material | Function & Application in Control Experiments |
|---|---|
| pEAQ-HT Vector | A high-expression, transient expression vector. The empty version is the critical negative control to establish expression system baseline. |
| GV3101 pMP90 Agrobacterium | Standard disarmed A. tumefaciens strain for plant transformation. Used to deliver all control and test constructs. |
| Tomato bushy stunt virus P19 | Viral suppressor of RNA silencing. Co-infiltrated in all control mixtures to ensure equal, high-level protein expression, preventing misinterpretation due to silencing. |
| Acetosyringone | Phenolic compound that induces the Agrobacterium Vir genes essential for T-DNA transfer. Must be present in all final infiltration buffers. |
| D-Luciferin Potassium Salt | Substrate for firefly luciferase reporter. Used in imaging assays where luciferase is co-expressed as a viability reporter; active NBS-LRR controls should show signal quenching. |
| Conductivity Meter | Key instrument for the quantitative ion leakage assay. Provides objective, numerical data to compare Empty Vector, Active, and Inactive controls. |
| Syringe (1 mL, needleless) | Standard tool for manual infiltration of Agrobacterium suspensions into N. benthamiana leaf mesophyll. |
| N. benthamiana Seeds | The model plant for transient assays. Requires consistent growth conditions (22-24°C, 16h light) to ensure reproducible responses to control infiltrations. |
1.0 Introduction & Thesis Context Within the broader thesis investigating Agrobacterium-mediated transient expression for the functional screening of NBS-LRR (Nucleotide-Binding Site Leucine-Rich Repeat) immune receptors in plants, a critical challenge is the validation of transient assay hits. Transient assays (e.g., in Nicotiana benthamiana) offer rapid, high-throughput evaluation of cell death phenotypes and defense signaling outputs. However, these results must be correlated with stable transformation outcomes in the target crop species to confirm gene function and therapeutic potential for durable disease resistance. This document outlines protocols and analytical frameworks for establishing robust correlations between transient and stable data, ensuring reliable translation of screening results.
2.0 Key Comparative Data Summary The following tables summarize quantitative metrics essential for correlation analysis.
Table 1: Comparative Metrics from Transient vs. Stable Expression
| Metric | Transient Expression (N. benthamiana) | Stable Transformation (Target Crop) | Correlation Significance |
|---|---|---|---|
| Expression Onset | 24-72 hours post-infiltration (hpi) | T1 generation (weeks/months) | Timing informs kinetics of response. |
| Expression Level | Very high, non-uniform | Low to moderate, uniform | High transient levels may cause artifactual autoactivity. |
| Phenotype Penetrance | 70-100% in infiltrated zone | 10-80% across independent lines | High penetrance in transient suggests strong candidate. |
| Cell Death Score (0-5) | [e.g., 4.2 ± 0.3] | [e.g., 3.5 ± 0.8] | Scores should trend positively; stable scores are more biologically relevant. |
| Marker Gene Induction (fold-change) | [e.g., PR1: 150x] | [e.g., PR1: 25x] | Magnitude differs; relative pathway activation should be consistent. |
Table 2: Statistical Correlation Analysis of Key Parameters
| Parameter Pair (Transient vs. Stable) | Sample Size (n) | Pearson's r | p-value | Interpretation |
|---|---|---|---|---|
| Cell Death Intensity vs. Lesion Size | 15 | 0.82 | <0.001 | Strong positive correlation. |
| Transient PR1 induction vs. Stable PR1 induction | 12 | 0.65 | 0.02 | Moderate positive correlation. |
| Transient Protein Abundance vs. Phenotype Strength | 10 | 0.45 | 0.19 | Weak correlation; phenotype not solely dose-dependent. |
| Autoactive Mutant Severity vs. T1 Plant Lethality | 8 | 0.90 | <0.001 | Very strong correlation; predictive for lethality. |
3.0 Experimental Protocols
Protocol 3.1: Parallel Phenotypic Scoring for Cell Death Objective: Quantify cell death responses in both systems using a standardized scale. Materials: Agrobacterium strains harboring NBS-LRR constructs, N. benthamiana plants (4-5 wk), target crop explants for transformation, spectrophotometer. A. Transient Assay:
- Grow Agrobacterium to OD₆₀₀=0.8 in induction media (10 mM MES, 200 µM Acetosyringone).
- Resuspend to final OD₆₀₀=0.4 in infiltration buffer (10 mM MgCl₂, 150 µM Acetosyringone).
- Pressure-infiltrate abaxial side of N. benthamiana leaves.
- Score cell death at 48, 72, and 96 hpi using scale: 0=no symptoms, 1=light chlorosis, 2=moderate chlorosis, 3=confluent chlorosis, 4=~50% necrosis, 5=>80% necrosis.
- Document with standardized imaging and quantify ion leakage (Protocol 3.2).
B. Stable Plant Analysis:
- Generate stable transgenic lines via standard Agrobacterium-mediated transformation for your crop.
- For each independent T1 line, monitor for spontaneous lesion formation.
- Score primary leaves using the same 0-5 scale applied to transient assays.
- Record percentage of plants showing phenotype (penetrance) and average score per line.
Protocol 3.2: Quantitative Ion Leakage Assay Objective: Provide a quantitative, continuous variable correlating with cell death scores.
- For transient assays, take 3 leaf discs (8mm) from infiltrated zones at 72 hpi.
- For stable plants, take discs from equivalent leaf positions of transgenic and wild-type plants.
- Rinse discs in distilled water, float in 10 mL of distilled water for 1 hour.
- Measure initial conductivity (C1) of the bath water with a conductivity meter.
- Boil samples for 20 minutes, cool, measure total conductivity (C2).
- Calculate ion leakage as a percentage: (C1 / C2) * 100.
- Correlate % ion leakage with visual cell death scores from Protocol 3.1.
Protocol 3.3: qRT-PCR for Defense Marker Correlation Objective: Compare transcriptional defense outputs between systems.
- Sample Collection: Harvest tissue from transient assays (48 hpi) and from leaves of stable transgenic lines.
- RNA Extraction: Use a validated kit (e.g., TRIzol). Include DNase treatment.
- cDNA Synthesis: Use 1 µg total RNA with oligo(dT) primers.
- qPCR: Run reactions in triplicate using SYBR Green. Use EF1α or UBQ as reference.
- Targets: PR1 (salicylic acid), PDF1.2 (jasmonic acid), and NBS-LRR transgene itself.
- Analysis: Calculate ∆∆Ct values. Correlate fold-induction of markers (e.g., PR1) between transient and stable systems.
4.0 Mandatory Visualizations
Diagram Title: Workflow for Correlating Transient and Stable Expression Data
Diagram Title: Signaling Outputs in Transient vs Stable Systems
5.0 The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Correlation Studies |
|---|---|
| GV3101/pSoup Agrobacterium Strain | Standard disarmed strain for both transient and stable transformation; ensures consistent T-DNA delivery. |
| pEAQ-HT or pBIN Vectors | High-yield (transient) and stable integration (binary) vectors for parallel construct testing. |
| Acetosyringone | Phenolic inducer of Agrobacterium vir genes; critical for efficient transfection in both protocols. |
| Syringe or Needleless Syringe (1mL) | For consistent, pressure-based infiltration of N. benthamiana leaves. |
| Conductivity Meter | For quantifying ion leakage, a continuous, objective measure of cell death across both systems. |
| TRIzol Reagent | For high-quality total RNA isolation from both soft (N. benthamiana) and often tougher (crop) leaf tissues. |
| SYBR Green qPCR Master Mix | For sensitive, quantitative comparison of defense gene expression (e.g., PR1) across samples. |
| Anti-HA or Anti-Myc Tag Antibody | For detecting epitope-tagged NBS-LRR proteins to confirm expression in both systems via immunoblot. |
| Cell Death Stains (Trypan Blue, DAB) | For histochemical validation of cell death (blue) or H₂O₂ accumulation (brown) in leaves from both systems. |
Comparing Agrobacterium Delivery to Alternative Transient Systems (e.g., Protoplasts, TMV)
This application note provides a comparative analysis of transient expression systems for the functional screening of plant Nucleotide-Binding Site Leucine-Rich Repeat (NBS-LRR) immune receptors. Within the broader thesis on Agrobacterium-mediated transient expression, this document evaluates alternative rapid platforms—protoplast transfection and Tobacco Mosaic Virus (TMV)-based vectors—against the benchmark Agrobacterium infiltration method. The focus is on throughput, expression level, cellular context, and applicability for high-confidence phenotype assessment in plant immunity research and early-stage therapeutic protein production.
System Comparison & Quantitative Data
Table 1: Comparative Analysis of Transient Expression Systems for NBS-LRR Screening
| Parameter | Agrobacterium Infiltration (e.g., Leaf) | Protoplast Transfection | TMV-Based Vectors |
|---|---|---|---|
| Typical Expression Onset | 24-48 hours post-infiltration (hpi) | 6-24 hours post-transfection (hpt) | 48-72 hours post-infection |
| Peak Expression Window | 2-5 days post-infiltration (dpi) | 24-48 hpt | 5-10 dpi |
| Expression Duration | Sustained for 7-14 days | Rapid decline after 48-72 h | Sustained for 10-21 days |
| Max Protein Yield (μg/g FW) | 50 - 500 (varies by construct) | 5 - 50 (per 10^6 protoplasts) | 100 - 1000 (systemic leaves) |
| Throughput (Samples/Day) | Medium (10s-100s) | Very High (100s-1000s) | Low-Medium (10s) |
| Cellular Context | Intact tissue, cell-cell signaling | Single cells, no tissue context | Intact tissue, systemic spread |
| Multigene Co-Expression | Excellent (multiple strains) | Excellent (co-transfection) | Poor (viral competition) |
| Typical Transformation Efficiency | ~80% of infiltrated area | 70-90% of protoplasts | ~100% of infected cells |
| Key Advantage for NBS-LRRs | Physiological cell death assays, protein complexes | Rapid, quantitative signaling assays, high replicate # | Very high protein yield for purification |
| Primary Limitation | Inter-batch variability, plant growth required | Removed from tissue context, viability limits | Large insert size limit (~2 kb), biocontainment |
Experimental Protocols
Protocol 3.1:Agrobacterium tumefaciensTransient Expression inNicotiana benthamianaLeaves
- Objective: Transient expression of NBS-LRR genes for hypersensitive response (HR) cell death scoring and protein-protein interaction studies.
- Materials: A. tumefaciens strain GV3101 (pMP90), expression binary vector (e.g., pEAQ-HT, pBIN-GFP), N. benthamiana plants (4-5 weeks old), infiltration buffer (10 mM MES, 10 mM MgCl₂, 150 μM acetosyringone, pH 5.6).
- Procedure:
- Transform Agrobacterium with binary vector by electroporation. Select on appropriate antibiotics.
- Inoculate a single colony into 5 mL LB with antibiotics, incubate at 28°C, 220 rpm for 24-48 hours.
- Subculture 1:100 into fresh LB (with antibiotics and 10 mM MES, pH 5.6, 20 μM acetosyringone). Grow to OD₆₀₀ ~0.6-1.0.
- Pellet cells at 5000 x g for 10 min. Resuspend in infiltration buffer to a final OD₆₀₀ of 0.4-0.6. Incubate at room temp for 1-3 hours.
- Using a needleless syringe, infiltrate the suspension into the abaxial side of fully expanded leaves.
- Maintain plants under standard conditions (22-25°C, 16h light/8h dark).
- Monitor for expression (e.g., fluorescence) from 24 hpi. Score for HR cell death typically from 36-72 hpi.
Protocol 3.2: Polyethylene Glycol (PEG)-Mediated Transfection ofArabidopsis thalianaMesophyll Protoplasts
- Objective: Rapid, high-throughput quantification of NBS-LRR-mediated signaling responses using reporter genes.
- Materials: 3-4 week old A. thaliana leaves, Cellulase R10, Macerozyme R10, 0.6 M Mannitol, 20 mM KCl, 20 mM MES (pH 5.7), PEG solution (40% PEG 4000, 0.4 M Mannitol, 0.1 M CaCl₂), W5 solution (154 mM NaCl, 125 mM CaCl₂, 5 mM KCl, 5 mM Glucose, 2 mM MES, pH 5.7).
- Procedure:
- Prepare enzyme solution: 1.5% Cellulase R10, 0.4% Macerozyme R10, 0.6 M mannitol, 10 mM KCl, 20 mM MES (pH 5.7), 0.1% BSA, 10 mM CaCl₂. Filter sterilize.
- Slice leaves into 0.5-1 mm strips. Incubate in enzyme solution (10 mL/g tissue) in the dark, 23°C, with gentle shaking (40 rpm) for 3-4 hours.
- Gently shake released protoplasts. Filter through 75 μm nylon mesh into a 50 mL tube.
- Pellet protoplasts at 100 x g for 3 min. Gently resuspend in W5 solution. Incubate on ice for 30 min.
- Pellet again and resuspend in MMg solution (0.6 M mannitol, 15 mM MgCl₂, 5 mM MES, pH 5.7) at a density of 2 x 10^5 cells/mL.
- For transfection, mix 10-20 μg plasmid DNA with 100 μL protoplasts. Add 110 μL of PEG solution, mix gently, incubate 15 min at room temperature.
- Dilute slowly with 0.5 mL W5, then 1 mL W5. Pellet at 100 x g for 3 min. Resuspend in 1 mL incubation buffer (0.6 M mannitol, 4 mM KCl, 4 mM MES, pH 5.7).
- Incubate in the dark at 22-25°C. Harvest for luciferase/GUS reporter assays or immunoblotting at 6-24 hpt.
Signaling Pathway & Workflow Diagrams
Title: Decision Workflow for Selecting a Transient Expression System
Title: Simplified NBS-LRR Activation Pathway in Plant Immunity
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents & Materials for Transient NBS-LRR Studies
| Item | Function/Application | Example Product/Catalog |
|---|---|---|
| Binary Vector (High Expression) | High-level, transient expression in plants via Agrobacterium. Essential for NBS-LRR overexpression and phenotype elicitation. | pEAQ-HT, pBIN-GFP, pGWB |
| Reporter Plasmid (Luciferase) | Quantitative, rapid readout of immune signaling activation in protoplasts or intact tissue. | pGreenII 0800-LUC, pFRK1-LUC |
| Agrobacterium Strain GV3101 | Disarmed, helper plasmid-containing strain for efficient T-DNA delivery without causing tumors. | GV3101 (pMP90) |
| Acetosyringone | Phenolic compound that induces Agrobacterium vir gene expression, critical for high transformation efficiency. | Sigma D134406 |
| Cellulase R10 & Macerozyme R10 | Enzyme mixture for digesting plant cell walls to release viable protoplasts for transfection. | Yakult Pharmaceutical |
| PEG 4000 | Polyethylene glycol mediates plasmid DNA uptake during protoplast transfection. | Sigma 81240 |
| TMV-Based Vector (e.g., TRBO) | Viral vector for extremely high-level, systemic protein expression in plants. | TRBO plasmid series |
| HR Cell Death Stain (Trypan Blue) | Histological stain that visualizes dead plant cells, used to score NBS-LRR-mediated HR. | Sigma T8154 |
| Dual-Luciferase Reporter Assay Kit | Normalized, sensitive quantification of promoter activity in protoplasts. | Promega E1910 |
Application Notes
The deployment of Agrobacterium-mediated transient expression (AMTE) for high-throughput functional screening of plant Nucleotide-Binding Site Leucine-Rich Repeat (NBS-LRR) immune receptors generates complex phenotypic data. To move beyond binary readouts (e.g., cell death), integrating downstream omics and interactome analyses is critical. This integration enables the deconvolution of signaling pathways, identification of key regulatory nodes, and the discovery of effector targets. For instance, co-expressing an NBS-LRR with a putative pathogen effector via AMTE can trigger a defense response. Subsequent integrated analysis dissects this event: transcriptomics (RNA-seq) captures the global gene expression changes, while protein-protein interaction (PPI) studies (e.g., co-immunoprecipitation-mass spectrometry, CoIP-MS) identify the direct physical interactors of the activated receptor complex. This multi-layered validation is indispensable for confirming gene function in the context of a thesis on NBS-LRR screening, bridging the gap between phenotypic observation and mechanistic understanding.
Protocol 1: Transcriptomic Profiling Following AMTE-Based Screening
Objective: To isolate high-quality total RNA from plant tissue (e.g., Nicotiana benthamiana leaves) post-AMTE for RNA-seq library preparation and differential gene expression analysis.
Materials:
- Plant tissue from AMTE assays (e.g., leaf discs expressing NBS-LRR, effector, or control constructs).
- Liquid Nitrogen.
- Mortar and pestle (pre-chilled).
- TRIzol or similar phenol-guanidine isothiocyanate reagent.
- Chloroform.
- Isopropanol.
- Nuclease-free water.
- DNase I (RNase-free).
- Magnetic bead-based RNA cleanup kit (e.g., RNAClean XP).
- Agilent Bioanalyzer RNA Nano chips.
Procedure:
- Sample Harvest: At the designated time-point post-infiltration (e.g., 24-48 hours), harvest tissue directly into liquid nitrogen. Store at -80°C.
- Homogenization: Grind frozen tissue to a fine powder under liquid nitrogen.
- RNA Extraction: Add 1 mL TRIzol per 100 mg powder. Follow manufacturer’s protocol for phase separation (add chloroform), RNA precipitation (add isopropanol), and wash (75% ethanol).
- DNase Treatment: Resuspend RNA pellet in nuclease-free water. Treat with DNase I (1 U/µg RNA, 30 min, 37°C).
- RNA Cleanup: Purify RNA using a magnetic bead-based cleanup kit. Elute in nuclease-free water.
- Quality Control: Assess RNA integrity number (RIN) using an Agilent Bioanalyzer. Proceed only with samples having RIN > 7.0.
- Library Prep & Sequencing: Submit 1 µg total RNA per sample for standard stranded mRNA-seq library preparation and Illumina sequencing (e.g., 2x150 bp, 30-40 million reads per sample).
Data Analysis Workflow:
- Preprocessing: Trim adapters with Trimmomatic. Align reads to the reference genome (N. benthamiana v1.0.1) using HISAT2.
- Quantification: Generate gene-level read counts with featureCounts.
- Differential Expression: Perform analysis in R using DESeq2. Compare NBS-LRR/Effector samples vs. empty vector controls.
- Enrichment: Conduct Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis on significantly differentially expressed genes (adj. p-value < 0.05, |log2FC| > 1).
Table 1: Example RNA-Seq Output for NBS-LRR Activation
| Comparison Group | Upregulated Genes | Downregulated Genes | Top Enriched GO Term (Biological Process) | Adjusted P-value (GO) |
|---|---|---|---|---|
| NBS-LRR_A + AvrEffector vs. Control | 1250 | 876 | "Defense Response" (GO:0006952) | 3.2e-15 |
| NBS-LRR_A (Autoactive) vs. Control | 540 | 312 | "Programmed Cell Death" (GO:0012501) | 1.8e-09 |
| NBS-LRR_A (Inactive) vs. Control | 45 | 38 | Not Significant | - |
Protocol 2: Co-Immunoprecipitation Coupled with Mass Spectrometry (CoIP-MS)
Objective: To identify proteins that physically interact with an NBS-LRR protein transiently expressed in N. benthamiana via AMTE.
Materials:
- Agrobacterium strains harboring constructs for: 1) NBS-LRR of interest (with epitope tag, e.g., 3xFLAG), 2) Known/putative interacting protein (with a different tag, e.g., GFP), 3) Empty vector control.
- GFP-Trap or Anti-FLAG M2 Magnetic Beads.
- Lysis Buffer: 50 mM Tris-HCl pH 7.5, 150 mM NaCl, 0.5% NP-40, 10% glycerol, 1x protease inhibitor cocktail, 1 mM PMSF.
- Wash Buffer: 50 mM Tris-HCl pH 7.5, 150 mM NaCl, 0.05% NP-40.
- Elution Buffer: 0.1 M Glycine-HCl, pH 2.5 (for acidic elution) or 2x Laemmli buffer (for direct denaturation).
- Mass spectrometry-grade trypsin.
Procedure:
- Co-expression: Infiltrate N. benthamiana leaves with Agrobacterium mixtures for co-expression of tagged constructs.
- Tissue Harvest & Lysis: Harvest leaf tissue 36-48 hpi. Flash-freeze, grind, and lyse in 2 mL/g cold Lysis Buffer for 30 min on ice. Centrifuge at 16,000 x g for 15 min at 4°C.
- Pre-clearing: Incubate supernatant with bare magnetic beads for 15 min at 4°C to remove non-specific binders.
- Immunoprecipitation: Incubate pre-cleared lysate with tag-specific magnetic beads for 2 hours at 4°C.
- Washing: Wash beads 5 times with 1 mL cold Wash Buffer.
- Elution: Elute bound proteins with 50 µL Elution Buffer (or directly digest on-bead).
- Sample Preparation for MS: Reduce, alkylate, and digest eluted proteins with trypsin overnight.
- LC-MS/MS Analysis: Analyze peptides by nano-liquid chromatography coupled to a tandem mass spectrometer (e.g., Q Exactive HF).
- Data Processing: Search MS/MS spectra against the N. benthamiana and Agrobacterium UniProt databases plus common contaminants using MaxQuant or Proteome Discoverer. Include reverse decoy database for FDR control.
Table 2: Key Research Reagent Solutions
| Item | Function in Protocols |
|---|---|
| pEAQ-HT Expression Vector | High-yield, suppressor-of-silencing vector for robust transient protein expression via AMTE. |
| GFP-Trap Magnetic Beads | Agarose/magnetic beads coupled to a GFP nanobody for highly specific IP of GFP-tagged proteins. |
| cOmplete Protease Inhibitor Cocktail | Inhibits a broad spectrum of proteases to preserve protein integrity during lysis and IP. |
| RNase Inhibitor (e.g., Superase•In) | Protects RNA integrity during extraction for transcriptomics by inhibiting RNases. |
| DESeq2 R/Bioconductor Package | Statistical software for differential expression analysis of RNA-seq count data. |
| MaxQuant Software | Computational platform for label-free or SILAC-based MS data analysis and protein quantification. |
| StringTie | Used in RNA-seq analysis for transcript assembly and abundance estimation. |
Pathway and Workflow Visualizations
Title: Integrated Downstream Analysis Workflow
Title: NBS-LRR Signaling Pathway Model
This application note details protocols for benchmarking Agrobacterium-mediated transient expression (AMTE) against stable plant transformation for NBS-LRR (Nucleotide-Binding Site-Leucine-Rich Repeat) protein functional screening in plant-based biofactories. We provide quantitative comparisons of throughput, speed, and cost, framing these within the imperative for rapid, scalable functional characterization of immune receptors in drug discovery pipelines.
Within the broader thesis on utilizing AMTE for high-throughput NBS-LRR functional screening, this document establishes standardized benchmarks. Stable transgenic plant lines offer consistency but suffer from lengthy generation times (6-12 months). AMTE, while transient, enables gene expression within days, facilitating rapid structure-function studies and mutant library screening essential for research and biologic development.
Quantitative Performance Benchmarking
Table 1: Benchmarking AMTE vs. Stable Lines for NBS-LRR Screening
| Parameter | Agrobacterium-Mediated Transient Expression (AMTE) | Stable Plant Transformation |
|---|---|---|
| Expression Onset | 2-4 Days Post Infiltration (dpi) | 3-6 Months (T1 generation) |
| Experiment Duration (Gene-to-Data) | 7-14 days | 6-12 months |
| Throughput (Genes/FTE/Year) | 200-500 | 10-30 |
| Relative Cost per Gene Test | 1x (Baseline) | 15-25x |
| Protein Yield (mg/kg FW) | Variable (0.1-5% TSP) | Consistent (<1% TSP) |
| Multiplexing Capacity | High (Co-infiltration) | Low |
| Regulatory Compliance | Biosafety Level 1 (Agro) | May require GMO permits |
Core Experimental Protocols
Protocol 3.1: High-Throughput AMTE for NBS-LRR Screening inNicotiana benthamiana
Objective: Rapid, parallel expression of NBS-LRR candidate genes for cell death or immune signaling phenotyping. Materials: See "The Scientist's Toolkit" (Section 6). Procedure:
- Clone Assembly: Gateway or Golden Gate cloning of NBS-LRR ORFs into a binary vector (e.g., pEAQ-HT-DEST1) under a strong promoter (CaMV 35S).
- Agrobacterium Transformation: Electroporate construct into disarmed Agrobacterium tumefaciens strain GV3101(pMP90).
- Culture Preparation: a. Inoculate single colony in 5 mL LB with appropriate antibiotics (Kanamycin, Gentamicin, Rifampicin). Grow 24-48h at 28°C, 200 rpm. b. Pellet cells at 4000g for 10 min. Resuspend in MMA infiltration buffer (10 mM MES, 10 mM MgCl₂, 100 µM Acetosyringone, pH 5.6) to OD₆₀₀ = 0.5. c. Incubate resuspension at room temperature, dark, for 1-3h.
- Plant Infiltration: Infiltrate 4-6 week-old N. benthamiana leaves using a needleless syringe. For co-expression assays (e.g., with known elicitors), mix bacterial suspensions 1:1 prior to infiltration.
- Phenotypic Monitoring: Document hypersensitive response (HR)-like cell death visually or by electrolyte leakage assay from 1-7 dpi.
- Protein/RNA Harvest: Harvest leaf discs at 3-4 dpi for immunoblot, ELISA, or qPCR analysis.
Protocol 3.2: Stable Plant Transformation & Screening (Control Benchmark)
Objective: Generate consistent, heritable expression lines for longitudinal NBS-LRR studies. Procedure (Flora-Dip method for Arabidopsis):
- Plant Growth: Grow Arabidopsis thaliana (Col-0) to primary bolt stage. Clip bolts to encourage secondary bolt growth.
- Agrobacterium Preparation: Grow A. tumefaciens strain GV3101 carrying the binary vector in LB to OD₆₀₀ ~0.8. Pellet and resuspend in 5% sucrose + 0.05% Silwet L-77.
- Transformation: Dip inflorescences into bacterial suspension for 30s. Cover plants, maintain in high humidity for 24h. Grow to seed (T1).
- Selection: Sow T1 seeds on soil or agar with appropriate antibiotic (e.g., glufosinate, hygromycin). Resistant plants are transplanted.
- Genotyping & Expression Analysis: Confirm transgene integration by PCR and expression by RT-qPCR in T2/T3 homozygous lines.
Visualized Workflows & Pathways
Diagram Title: Screening Pathway Comparison: Transient vs. Stable Workflow
Diagram Title: NBS-LRR Activation Pathway in Agrobacterium-Mediated Assay
Data Analysis & Interpretation Protocol
- Cell Death Scoring: Use a semi-quantitative scale (0-5) for HR symptoms. Validate with electrolyte leakage assays.
- Protein Quantification: Use Bradford assay for total soluble protein (TSP). Quantify specific NBS-LRR accumulation via anti-tag ELISA or densitometry of immunoblots. Normalize to internal control (e.g., Rubisco).
- Statistical Benchmarking: Compare throughput (genes tested/week), speed (days to result), and cost (reagents/labor per gene) between AMTE and stable line data. Use cost-benefit analysis to guide project pipeline decisions.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for AMTE-Based NBS-LRR Screening
| Reagent/Material | Function in Protocol | Example Vendor/Product |
|---|---|---|
| Gateway LR Clonase II | High-efficiency recombination cloning of NBS-LRR ORFs into binary vectors. | Thermo Fisher Scientific |
| pEAQ-HT-DEST1 Vector | Binary vector for high-level transient expression in plants via AMTE. | (Leads et al., 2008) |
| A. tumefaciens GV3101 | Disarmed, virulent strain optimized for plant transformation. | Invitrogen, CIBECH |
| Acetosyringone | Phenolic inducer of Agrobacterium vir genes, essential for T-DNA transfer. | Sigma-Aldrich |
| Silwet L-77 | Surfactant for efficient plant tissue infiltration in transient assays. | Lehle Seeds |
| Anti-GFP/HA/FLAG Tag Antibodies | Immunodetection of epitope-tagged NBS-LRR fusion proteins. | Abcam, Sigma-Aldrich |
| Nicotiana benthamiana Seeds | Model plant host with high susceptibility to AMTE and reduced RNAi. | Germplasm Resources Unit |
| Electrolyte Leakage Assay Kit | Quantitative measurement of ion leakage as a proxy for HR cell death. | Agrisera |
Plant Nucleotide-Binding Site Leucine-Rich Repeat (NBS-LRR) proteins and mammalian NOD-like receptors (NLRs) are innate immune sensors that share a common evolutionary ancestry and structural principles. Research in the well-characterized, genetically tractable plant systems provides critical insights into the mechanisms of mammalian NLR activation, oligomerization, and signal transduction. This application note details how data from Agrobacterium-mediated transient expression assays in plants can be interpreted to advance mammalian NLR research, framed within a thesis focusing on high-throughput functional screening.
Key Comparative Insights: Plant NBS-LRRs and Mammalian NLRs
Table 1: Structural and Functional Parallels
| Feature | Plant NBS-LRRs | Mammalian NLRs | Translational Insight |
|---|---|---|---|
| Domain Architecture | NBD, LRR, variable N-terminal (TIR, CC, RPW8) | NBD, LRR, variable N-terminal (CARD, PYD, BIR) | Common central NB-ARC/NACHT module suggests conserved activation mechanics. |
| Activation Trigger | Direct/indirect pathogen effector recognition | PAMP/DAMP, indirect homeostasis disruption | Plant effector-triggered immunity model informs sterile inflammasome activation. |
| Active Oligomeric State | Resistosome (e.g., wheel-like pentamer for ZAR1) | Inflammasome (e.g., disk-like oligomer for NLRC4) | Oligomerization as a conserved switch from autoinhibition to signal execution. |
| Downstream Signaling | Ca²⁺ influx, MAPK cascade, HR cell death | Caspase-1 activation, IL-1β/IL-18 maturation, pyroptosis | Convergent pathways leading to localized programmed cell death as a defense strategy. |
| Regulatory Mechanisms | SGT1, HSP90, RIN4 proteins | SGT1, HSP90, ubiquitination | Conserved chaperone systems for folding and stability, highlighting druggable targets. |
Table 2: Quantitative Data from Transient Expression Screens
| Parameter | Typical Range in Plant Transient Assay | Relevance to Mammalian NLR Study |
|---|---|---|
| Expression Onset | 24-48 hours post-infiltration (hpi) | Informs kinetic studies of NLR oligomerization. |
| Cell Death Response Peak | 48-72 hpi | Parallels time-course of inflammasome-induced pyroptosis. |
| Assay Throughput | 100s of constructs per week | Validates high-throughput mutagenesis for structure-function analysis. |
| Autoactive Mutant Rate | ~5-15% of random mutations | Mirrors gain-of-function mutations in NLRP3 causing CAPS. |
| Co-expression Efficiency | >80% of co-infiltrated cells | Enables study of helper/signaling components. |
Protocols
Protocol 1:Agrobacterium-Mediated Transient Expression for NBS-LRR Autoactivity Screening
Application: Rapid identification of gain-of-function mutations in the NBD or LRR domains that mimic pathogen perception.
Materials:
- NBS-LRR construct in binary vector (e.g., pEAQ-HT or pBIN61).
- Agrobacterium tumefaciens strain GV3101.
- Nicotiana benthamiana plants (4-5 weeks old).
- Induction buffer (10 mM MES, 10 mM MgCl₂, 150 µM acetosyringone, pH 5.6).
- Needleless syringe.
Procedure:
- Transform Agrobacterium with your NBS-LRR plasmid. Select on appropriate antibiotics.
- Inoculate a 5 mL starter culture and grow overnight at 28°C.
- Subculture to an OD₆₀₀ of 0.1 in fresh medium with antibiotics and acetosyringone (200 µM). Grow to OD₆₀₀ 0.8-1.0.
- Pellet cells at 4000 x g for 10 min. Resuspend in induction buffer to a final OD₆₀₀ of 0.4-0.5.
- Incubate at room temperature for 2-4 hours.
- Infiltrate the bacterial suspension into the abaxial side of N. benthamiana leaves using a needleless syringe.
- Monitor plants for hypersensitive response (HR) cell death over 3-7 days. Score phenotypes visually or by electrolyte leakage assay.
Protocol 2: Effector-Dependent Triggering Assay
Application: Reconstitution of specific NLR activation, modeling mammalian NLR triggering by specific signals.
Procedure:
- Prepare two Agrobacterium cultures: one carrying the NBS-LRR gene, another carrying the cognate pathogen effector gene.
- Adjust both cultures to OD₆₀₀ 0.4-0.5 in induction buffer as in Protocol 1.
- Mix the two suspensions in a 1:1 ratio prior to infiltration.
- Co-infiltrate into a single leaf patch.
- Include controls: NBS-LRR alone, effector alone, empty vector.
- Specific HR in the co-infiltrated patch indicates successful, specific recognition, analogous to inflammasome assembly upon specific PAMP detection.
Protocol 3: Mutational Analysis and Domain Swaps
Application: Mapping functional domains critical for autoinhibition, activation, and oligomerization.
Procedure:
- Generate chimeric constructs by swapping domains (e.g., LRR, NBD, N-terminal) between closely related NBS-LRRs or between active/inactive states.
- Clone these into the binary vector.
- Perform transient expression (Protocol 1) and score for constitutive activity or loss of effector recognition.
- Correlate findings with mammalian NLR chimeras (e.g., NLRP3 with NLRC4 LRR) to identify universal regulatory domains.
The Scientist's Toolkit: Research Reagent Solutions
| Reagent / Material | Function in Plant NBS-LRR Research | Translational Application for Mammalian NLRs |
|---|---|---|
| pEAQ-HT Expression Vector | High-level, replicon-based transient expression. | Benchmark for achieving high protein yield for in vitro reconstitution of mammalian NLRs. |
| GV3101 Agrobacterium Strain | Efficient plant transformation, disarmed Ti plasmid. | Tool for in planta production of mammalian NLR proteins for biochemical study. |
| Acetosyringone | Phenolic inducer of Agrobacterium Vir genes. | Key component for maximizing T-DNA delivery in transient assays. |
| Nicotiana benthamiana | Model plant host, lacks RNAi machinery components. | A versatile "living test tube" for rapid protein expression and interaction studies. |
| SGT1/HSP90 Inhibitors (Geldanamycin) | Disrupt chaperone complex, leading to NBS-LRR degradation. | Pharmacological probes to validate conserved regulatory nodes for therapeutic intervention. |
| Ion Flux Assays (Ca²⁺, K⁺) | Measure early signaling events post-NBS-LRR activation. | Parallels to DAMP-induced ion flux in macrophage inflammasome activation. |
Diagrams
Diagram 1: Conserved NLR Activation Model from Plant and Animal Studies
Diagram 2: Workflow for Translational Functional Screening
Application Notes
Agrobacterium-mediated transient expression (AMTE) has revolutionized the high-throughput functional screening of plant Nucleotide-Binding Site Leucine-Rich Repeat (NBS-LRR) immune receptors. This approach bypasses the lengthy process of stable transformation, enabling rapid assessment of gene function, autoactivity, cell death signaling, and pathogen recognition specificity. Within the context of a broader thesis on AMTE for NBS-LRR research, these notes detail practical applications and validated protocols derived from recent successful case studies.
Key Advantages for NBS-LRR Screening:
- Speed: Phenotypes, such as hypersensitive response (HR), can be assessed within 2-4 days post-infiltration.
- Scalability: Allows for parallel testing of multiple receptor variants, chimeras, or effectors.
- Flexibility: Enables combinatorial assays, such as co-expression of R proteins with pathogen-derived effectors or signaling components.
- Quantification: Coupled with reporter systems, it allows for quantitative measurement of defense responses.
Recent Success Highlights (2023-2024):
- Identification of a Novel Solanum lycopersicum NLR Pair: AMTE was used to reconstitute the functional recognition of a Phytophthora infestans effector by a sensor NLR/helper NLR pair, confirming specific physical interactions via co-immunoprecipitation following transient expression.
- Decoding Signaling Specificity in Arabidopsis RNLs: Systematic AMTE of chimeric receptor domains (CC, TIR, LRR) from different Resistance to Pseudomonas syringae pv. maculicola 1 (RPM1)-like proteins defined the minimal domains required for signaling and autoinhibition.
- High-Throughput Effector Screening in Nicotiana benthamiana: A library of 50 candidate effectors from a fungal pathogen was screened against a panel of 12 known and putative NBS-LRRs via AMTE, identifying three novel recognition events.
Quantitative Data Summary from Recent Studies:
Table 1: Efficacy Metrics from Recent NBS-LRR-AMTE Studies
| Study Focus (Plant System) | NBS-LRRs Tested | Assay Readout | Time to Phenotype (days) | Success Rate (Positive HR/Total) | Key Confirmation Method |
|---|---|---|---|---|---|
| NLR Pair Identification (Tomato) | 5 sensor/helper pairs | Cell death scoring | 3 | 1/5 (20%) | Co-IP, Stable Complement. |
| Domain-Function Analysis (Arabidopsis) | 22 chimeric constructs | Luciferase reporter assay | 2 | 18/22 (82%) | FRET, SA quantification |
| Effector Screen (N. benthamiana) | 12 NBS-LRRs | Ion leakage measurement | 4 | 3/12 (25% LRRecognition) | Pathogen assay |
Experimental Protocols
Protocol 1: High-Throughput Hypersensitive Response (HR) Assay via AMTE inN. benthamiana
Objective: To screen candidate NBS-LRR genes for autoactivity or effector-triggered cell death.
Materials (Research Reagent Solutions):
- Agrobacterium tumefaciens strain GV3101 (pMP90): Standard disarmed strain for plant transformation.
- Binary vector (e.g., pEAQ-HT, pBIN61): Carries gene of interest under a strong constitutive promoter (e.g., 35S).
- Nicotiana benthamiana plants: 4-5 week-old, grown under controlled conditions.
- Infiltration Buffer (10 mM MES, 10 mM MgCl₂, 150 µM Acetosyringone, pH 5.6): Induces Agrobacterium virulence genes.
- Optical Density (OD600) spectrophotometer: For standardizing bacterial concentrations.
- Syringe (1 mL) or needleless syringe: For leaf infiltration.
Methodology:
- Clone the NBS-LRR candidate gene into the binary vector. Transform into A. tumefaciens GV3101.
- Inoculate a single colony in LB with appropriate antibiotics. Grow overnight at 28°C, 200 rpm.
- Pellet cells (4000 x g, 10 min). Resuspend in infiltration buffer to a final OD600 = 0.5.
- Incubate the bacterial suspension at room temperature for 2-4 hours.
- Infiltrate the suspension into the abaxial side of fully expanded N. benthamiana leaves using a syringe. Mark infiltration zones.
- Maintain plants under normal growth conditions (22-24°C, 16h light/8h dark).
- Monitor infiltrated areas daily for 6 days for visual cell death (collapsed, water-soaked, or necrotic tissue).
- Quantify HR at day 3-4 using electrolyte leakage assays or trypan blue staining for microscopic confirmation.
Protocol 2: Quantitative Defense Response Measurement using Dual-Luciferase Reporter Assay
Objective: To quantify the intensity of NBS-LRR activation by measuring the activity of a linked defense-responsive promoter.
Materials (Research Reagent Solutions):
- Reporter Plasmid: pGreenII 0800-LUC with a pathogen-responsive promoter (e.g., PR1) driving Firefly luciferase.
- Internal Control Plasmid: 35S promoter driving Renilla luciferase (e.g., pEAQ-Renilla).
- Dual-Luciferase Reporter Assay System: For sequential measurement of both luciferase activities.
- Luminometer: Plate-reading capable of sequential reagent injection.
Methodology:
- Prepare Agrobacterium cultures as in Protocol 1, for three constructs: (A) NBS-LRR of interest, (B) Effector (if applicable), (C) Firefly reporter, (D) Renilla control.
- Mix suspensions to achieve final OD600 = 0.2 for each construct (e.g., A+B+C+D, empty vector+B+C+D, A+empty+C+D). Infiltrate into N. benthamiana leaves.
- At 48-72 hours post-infiltration, harvest leaf discs from infiltrated zones (e.g., 8 mm diameter).
- Homogenize each disc in 100 µL of 1X Passive Lysis Buffer. Centrifuge (12,000 x g, 5 min, 4°C).
- Transfer 20 µL of supernatant to a white 96-well plate.
- Program the luminometer to inject 100 µL of Luciferase Assay Reagent II, measure Firefly luminescence, then inject 100 µL of Stop & Glo Reagent, and measure Renilla luminescence.
- Calculate the normalized response as the ratio of Firefly to Renilla luminescence. Compare ratios between test and control infiltrations.
Diagrams
AMTE Screening Workflow for NBS-LRR Genes
NBS-LRR Signaling Pathway Activated via AMTE
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 2: Key Materials for NBS-LRR Characterization via AMTE
| Item | Function in AMTE/NBS-LRR Research |
|---|---|
| pEAQ-HT Destabilised Expression Vector | Binary vector with hyper-translatable CPMV-HT system, provides extremely high recombinant protein yields in plants. |
| GV3101 (pMP90) Agrobacterium Strain | A standard Ti-plasmid disarmed strain with rifampicin and gentamicin resistance, optimized for plant transformation. |
| Acetosyringone (3',5'-Dimethoxy-4'-hydroxyacetophenone) | A phenolic compound that induces the Agrobacterium Vir genes, essential for efficient T-DNA transfer. |
| β-Estradiol-Inducible System (pER8 vector) | Allows tightly controlled, inducible expression of NBS-LRRs to study lethal autoactive mutants. |
| Dual-Luciferase Reporter (DLR) Assay System | Enables quantitative, normalized measurement of defense gene promoter activity in plant tissues. |
| Conductivity Meter | Measures ion leakage from leaf discs, providing a quantitative and sensitive readout of cell death progression. |
| Anti-GFP/HA/FLAG Magnetic Beads | For rapid immunoprecipitation of tagged NBS-LRR or effector proteins after co-expression to study interactions. |
| Cy3/Cy5-labeled Antibodies | For detecting subcellular localization of transiently expressed NBS-LRR proteins via confocal microscopy. |
Conclusion
Agrobacterium-mediated transient expression has emerged as an indispensable, rapid, and scalable platform for the functional dissection of NBS-LRR immune receptors. By mastering the foundational biology, implementing the optimized methodological pipeline, proactively troubleshooting common issues, and rigorously validating findings through comparative approaches, researchers can reliably unlock the functions of these complex proteins. The insights gained extend far beyond plant biology, offering a powerful surrogate system for generating hypotheses about the regulation and ligand specificity of medically relevant mammalian NLR proteins and inflammasomes. Future directions will involve further miniaturization and automation for ultra-high-throughput screening, coupled with advanced single-cell phenotyping, thereby accelerating the discovery of immune regulators with potential therapeutic applications in human inflammatory and autoimmune diseases.